预约演示
更新于:2026-04-25
R-82913
更新于:2026-04-25
概要
基本信息
在研机构- |
非在研机构 |
权益机构- |
最高研发阶段无进展临床2期 |
首次获批日期- |
最高研发阶段(中国)- |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C16H20ClN3S |
InChIKeyRCSLUNOLLUVOOG-NSHDSACASA-N |
CAS号126347-69-1 |
关联
100 项与 R-82913 相关的临床结果
登录后查看更多信息
100 项与 R-82913 相关的转化医学
登录后查看更多信息
100 项与 R-82913 相关的专利(医药)
登录后查看更多信息
113
项与 R-82913 相关的文献(医药)2021-12-01·BMC Pharmacology & Toxicology
Computational determination of toxicity risks associated with a selection of approved drugs having demonstrated activity against COVID-19
Article
作者: Delgado, Williams Ernesto Miranda ; Tuszynski, Jack A ; Houghton, Michael ; Wacker, Soren ; Noskov, Sergey ; Tyrrell, D Lorne J ; Aminpour, Maral
Abstract:
Background:
The emergence and rapid spread of SARS-CoV-2 (severe acute respiratory syndrome coronavirus 2) in thelate 2019 has caused a devastating global pandemic of the severe pneumonia-like disease coronavirus disease 2019 (COVID-19). Although vaccines have been and are being developed, they are not accessible to everyone and not everyone can receive these vaccines. Also, it typically takes more than 10 years until a new therapeutic agent is approved for usage. Therefore, repurposing of known drugs can lend itself well as a key approach for significantly expediting the development of new therapies for COVID-19.
Methods:
We have incorporated machine learning-based computational tools and in silico models into the drug discovery process to predict Adsorption, Distribution, Metabolism, Excretion, and Toxicity (ADMET) profiles of 90 potential drugs for COVID-19 treatment identified from two independent studies mainly with the purpose of mitigating late-phase failures because of inferior pharmacokinetics and toxicity.
Results:
Here, we summarize the cardiotoxicity and general toxicity profiles of 90 potential drugs for COVID-19 treatment and outline the risks of repurposing and propose a stratification of patients accordingly. We shortlist a total of five compounds based on their non-toxic properties.
Conclusion:
In summary, this manuscript aims to provide a potentially useful source of essential knowledge on toxicity assessment of 90 compounds for healthcare practitioners and researchers to find off-label alternatives for the treatment for COVID-19. The majority of the molecules discussed in this manuscript have already moved into clinical trials and thus their known pharmacological and human safety profiles are expected to facilitate a fast track preclinical and clinical assessment for treating COVID-19.
2020-10-01·Nature
Discovery of SARS-CoV-2 antiviral drugs through large-scale compound repurposing
Article
作者: Su, Andrew I ; Dejosez, Marion ; Hull, Mitchell V ; Nguyen, Tu-Trinh H ; Poon, Vincent Kwok-Man ; Mesecar, Andrew D ; Glynne, Richard J ; Chang, Max W ; Choi, Angela ; Herbert, Kristina M ; Pu, Yuan ; Burgstaller-Muehlbacher, Sebastian ; Matsunaga, Naoko ; Martinez-Sobrido, Luis ; Yin, Xin ; Zwaka, Thomas P ; Pache, Lars ; Benner, Christopher ; Chanda, Sumit K ; Lendy, Emma K ; Chatterjee, Arnab K ; Chapman, Mackenzie E ; Miorin, Lisa ; Johnson, Jeffrey R ; Rubanov, Andrey ; White, Kris M ; Rathnasinghe, Raveen ; Cao, Jianli ; Riva, Laura ; Schotsaert, Michael ; Albrecht, Randy ; Cheng, Kuoyuan ; Liu, Wen-Chun ; García-Sastre, Adolfo ; Teriete, Peter ; Nguyen, Courtney ; Chan, Jasper Fuk-Woo ; Yuen, Kwok-Yung ; Yuan, Shuofeng ; Sun, Ren ; De Jesus, Paul D ; Sit, Ko-Yung ; Martin-Sancho, Laura ; Schultz, Peter G ; Ruppin, Eytan
The emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in 2019 has triggered an ongoing global pandemic of the severe pneumonia-like disease coronavirus disease 2019 (COVID-19)1. The development of a vaccine is likely to take at least 12-18 months, and the typical timeline for approval of a new antiviral therapeutic agent can exceed 10 years. Thus, repurposing of known drugs could substantially accelerate the deployment of new therapies for COVID-19. Here we profiled a library of drugs encompassing approximately 12,000 clinical-stage or Food and Drug Administration (FDA)-approved small molecules to identify candidate therapeutic drugs for COVID-19. We report the identification of 100 molecules that inhibit viral replication of SARS-CoV-2, including 21 drugs that exhibit dose-response relationships. Of these, thirteen were found to harbour effective concentrations commensurate with probable achievable therapeutic doses in patients, including the PIKfyve kinase inhibitor apilimod2-4 and the cysteine protease inhibitors MDL-28170, Z LVG CHN2, VBY-825 and ONO 5334. Notably, MDL-28170, ONO 5334 and apilimod were found to antagonize viral replication in human pneumocyte-like cells derived from induced pluripotent stem cells, and apilimod also demonstrated antiviral efficacy in a primary human lung explant model. Since most of the molecules identified in this study have already advanced into the clinic, their known pharmacological and human safety profiles will enable accelerated preclinical and clinical evaluation of these drugs for the treatment of COVID-19.
2018-09-01·JOURNAL OF MOLECULAR STRUCTURE
QSAR studies of TIBO derivatives as HIV-1 reverse transcriptase inhibitors using HQSAR, CoMFA and CoMSIA
作者: Shan Lei ; Yang Wang ; Jianbo Tong ; Shangshang Qin
The study deals with CoMFA, CoMSIA and HQSAR to explore the important features of tetrahydroimidazo[4,5,1-jk][1,4]benzodiazepinone (TIBO) derivatives for exerting potent HIV-1 reverse transcriptase (HIV-1 RT) inhibitors activity.The cross-validated q2 value of CoMFA model is 0.641 and the non-cross-validated r2 value is 0.847.The best cross-validated q2 value of CoMSIA Model is 0.706 and the non-cross-vaildated r2 value is 0.939.The most effective HQSAR model was obtained that the cross-validation q2 value of 0.839, the non-cross-validated r2 value of 0.942, the standard error of prediction SDCV value of 0.604, and the best hologram length value of 307 using atoms and bonds as fragment distinctions.The statistical parameters from models indicate that the data are well fitted and have high predictive ability.Furthermore, Mol. docking was employed to explore the binding requirements between the ligands and the receptor protein which included several hydrogen bonds between the TIBO inhibitors and active site residues.Observations derived from these QSAR modeling study may be utilized further in designing promising HIV-1 reverse transcriptase inhibitors.
100 项与 R-82913 相关的药物交易
登录后查看更多信息
外链
| KEGG | Wiki | ATC | Drug Bank |
|---|---|---|---|
| - | - | - |
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| HIV感染 | 临床2期 | 比利时 | - | - |
| HIV感染 | 临床2期 | 法国 | - | - |
| HIV感染 | 临床2期 | 英国 | - | - |
| HIV感染 | 临床2期 | - | - |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
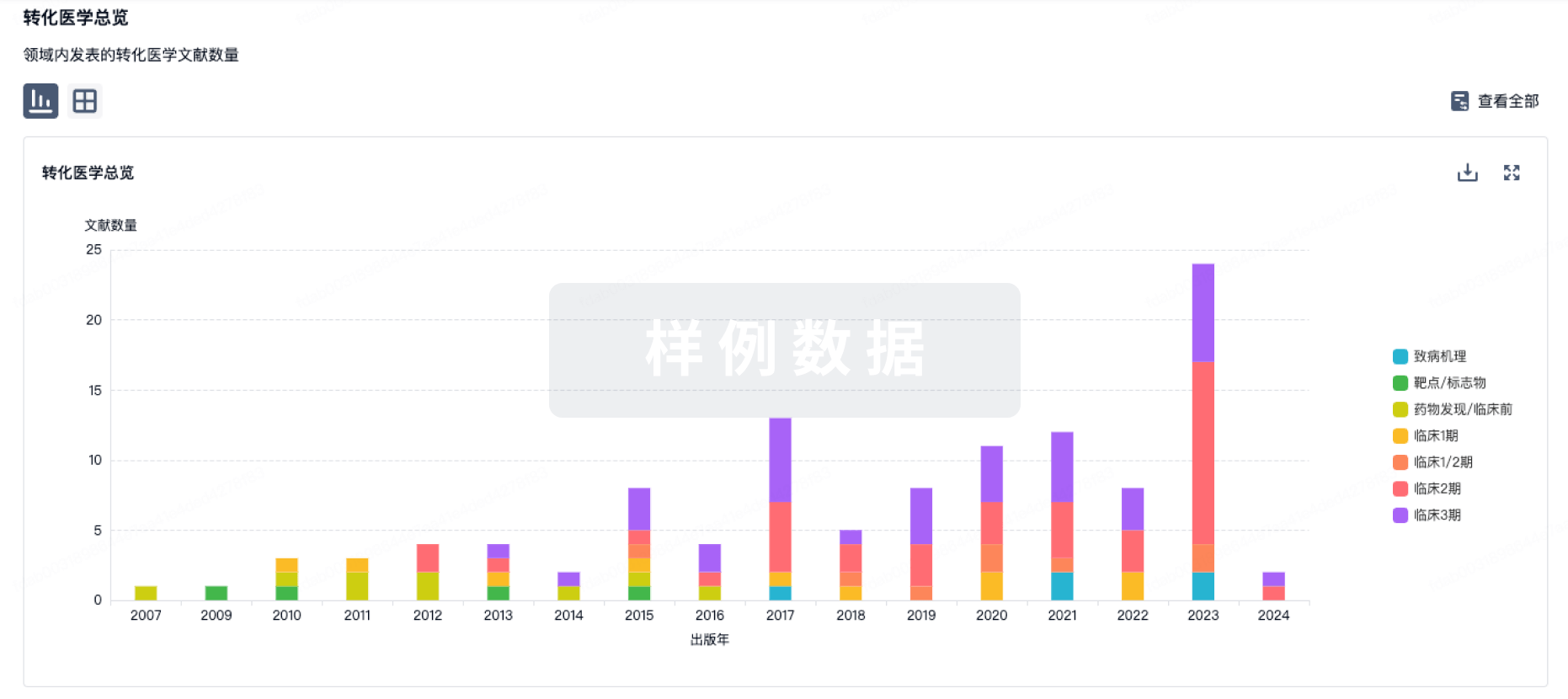
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
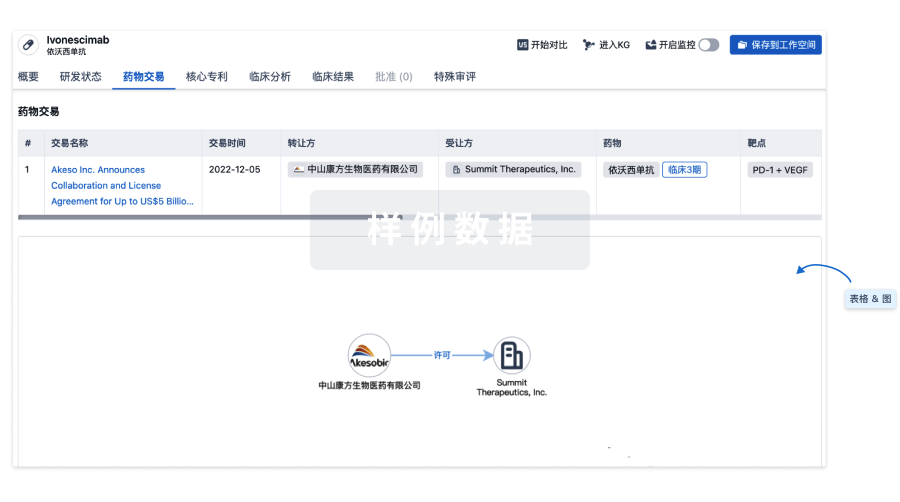
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
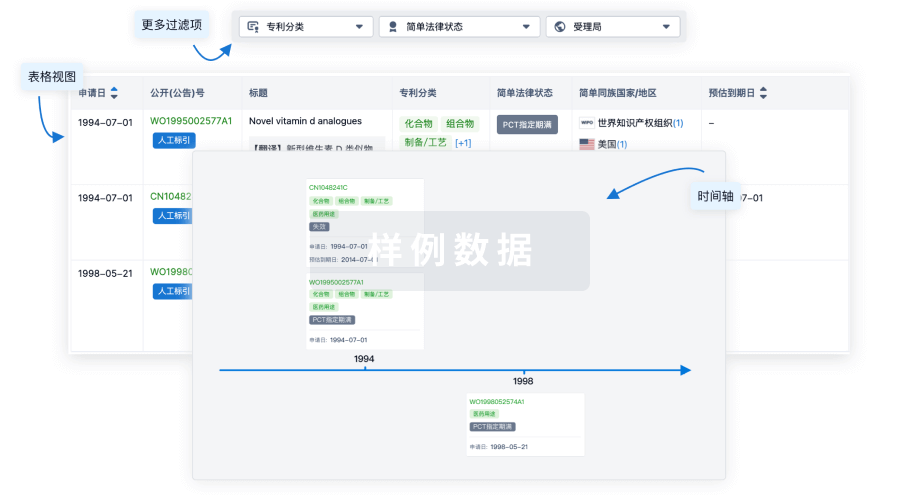
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
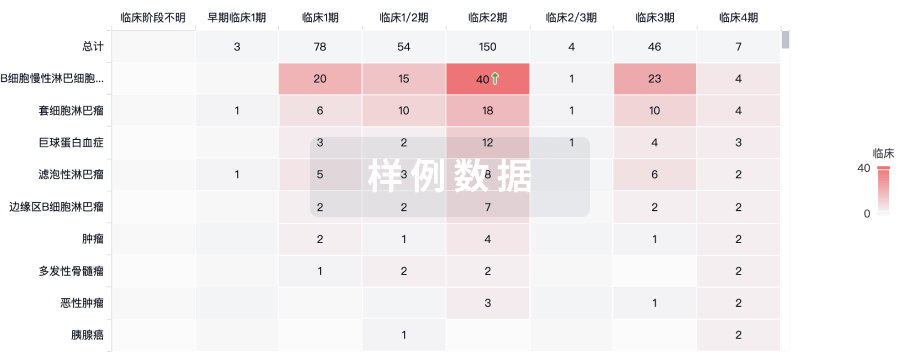
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
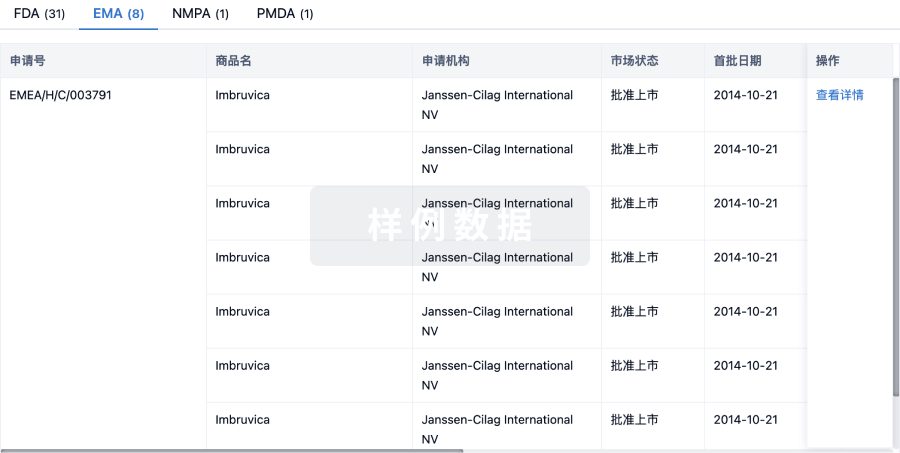
批准
利用最新的监管批准信息加速您的研究。
登录
或
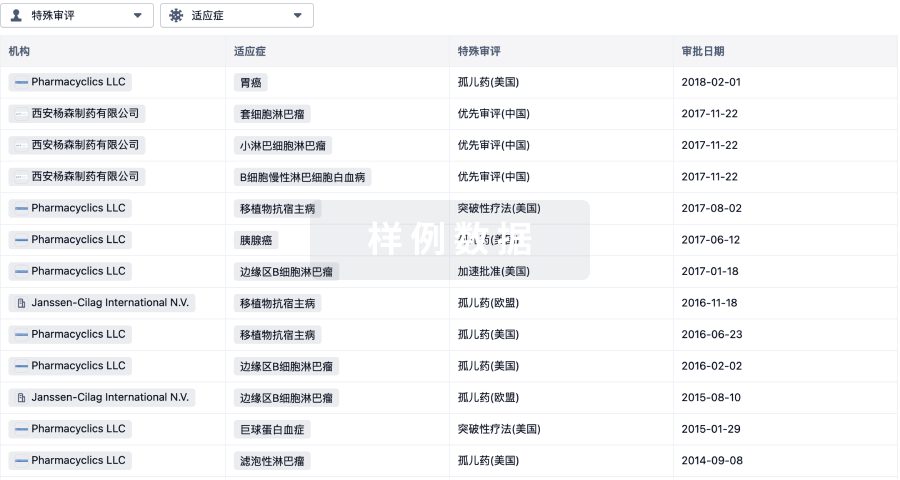
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
