预约演示
更新于:2025-05-07
MDL-63042
更新于:2025-05-07
概要
基本信息
在研机构- |
非在研机构 |
权益机构- |
最高研发阶段终止临床前 |
首次获批日期- |
最高研发阶段(中国)- |
特殊审评- |
关联
100 项与 MDL-63042 相关的临床结果
登录后查看更多信息
100 项与 MDL-63042 相关的转化医学
登录后查看更多信息
100 项与 MDL-63042 相关的专利(医药)
登录后查看更多信息
60
项与 MDL-63042 相关的文献(医药)2022-02-01·Biotechnology Letters4区 · 工程技术
Production enhancement of the glycopeptide antibiotic A40926 by an engineered Nonomuraea gerenzanensis strain
4区 · 工程技术
Article
作者: Gao, Wen ; Dong, Huijun ; Tian, Li ; Yan, Bingyu ; Wang, Shuai
2021-05-21·ACS Chemical Biology2区 · 生物学
Genomic-Led Discovery of a Novel Glycopeptide Antibiotic by Nonomuraea coxensis DSM 45129
2区 · 生物学
Article
作者: Kalinowski, Jörn ; Vior, Natalia M. ; Marinelli, Flavia ; Berini, Francesca ; Busche, Tobias ; Rückert, Christian ; Yushchuk, Oleksandr ; Truman, Andrew W. ; Binda, Elisa ; Andreo-Vidal, Andres
2021-01-01·The FEBS Journal3区 · 生物学
Understanding the early stages of peptide formation during the biosynthesis of teicoplanin and related glycopeptide antibiotics
3区 · 生物学
Article
作者: Kittilä, Tiia ; Cryle, Max J. ; Kaniusaite, Milda ; Tailhades, Julien ; Goode, Robert J.A. ; Fage, Christopher D. ; Schittenhelm, Ralf B.
100 项与 MDL-63042 相关的药物交易
登录后查看更多信息
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 细菌感染 | 临床前 | 意大利 | - |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
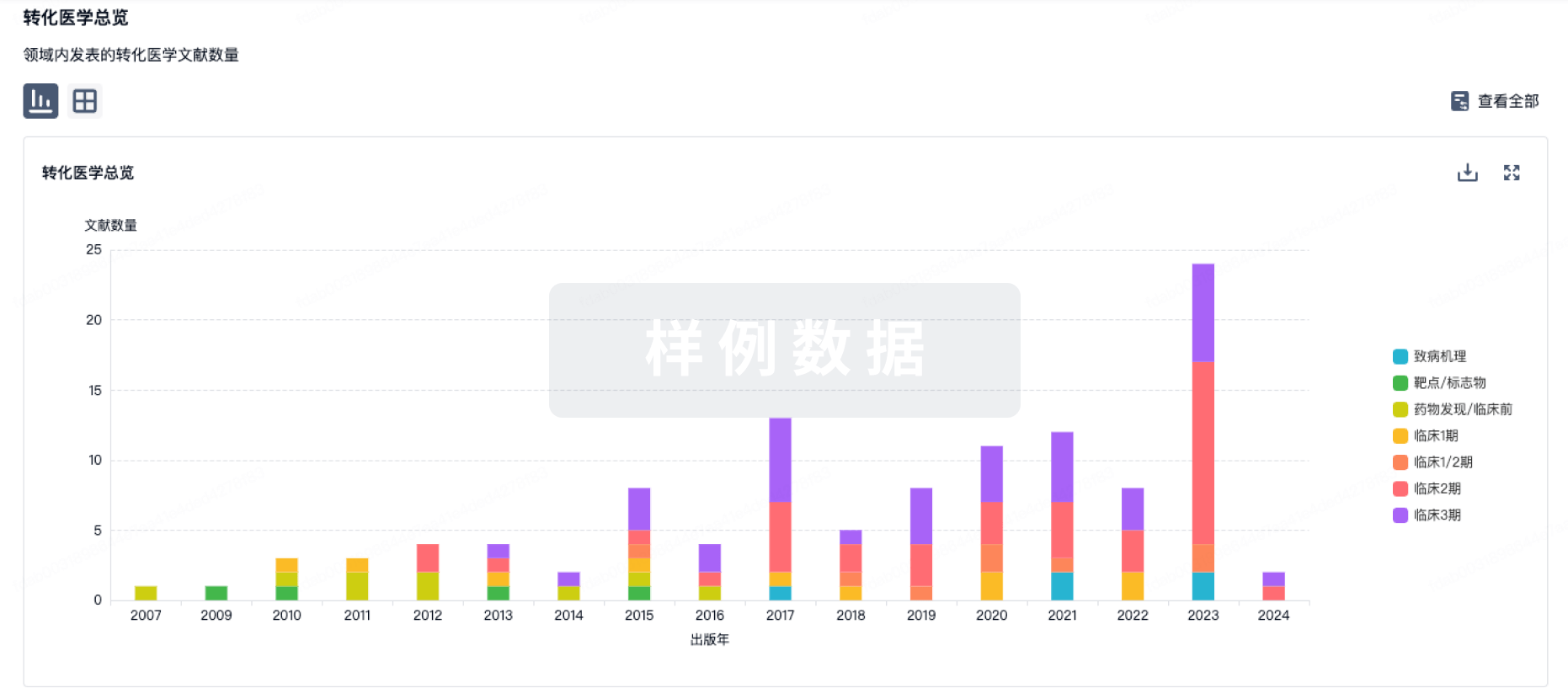
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
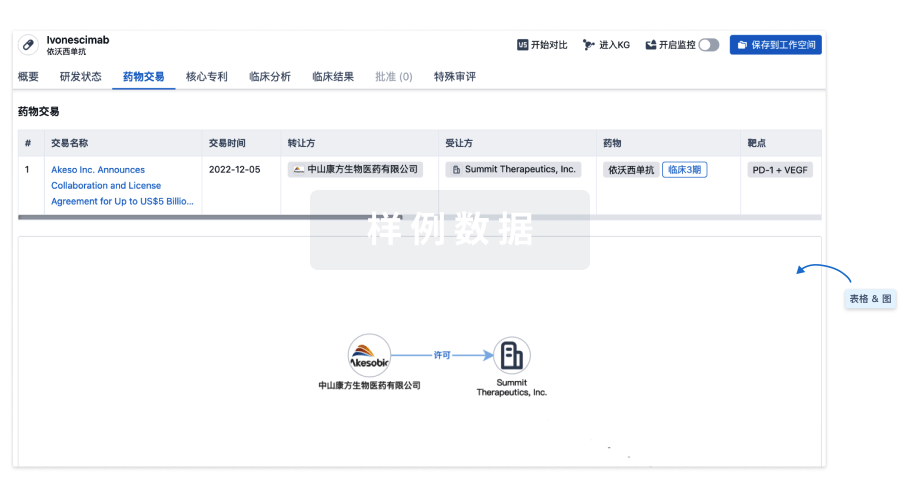
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
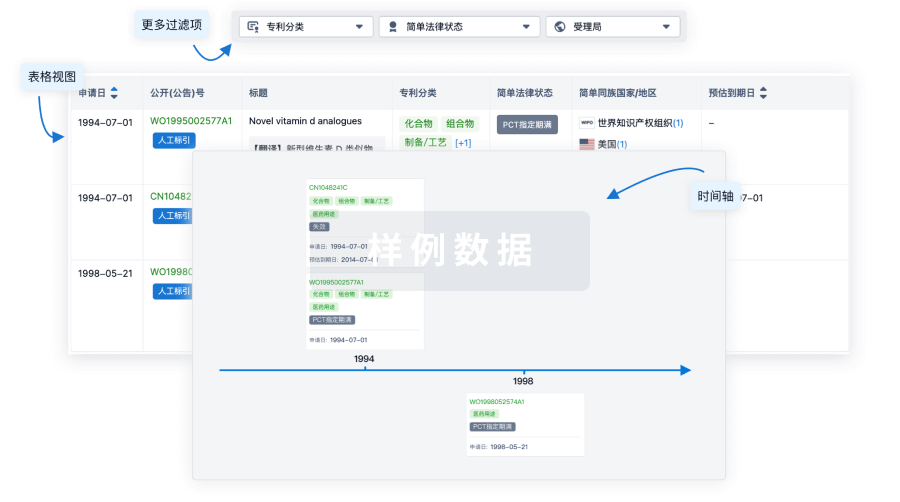
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
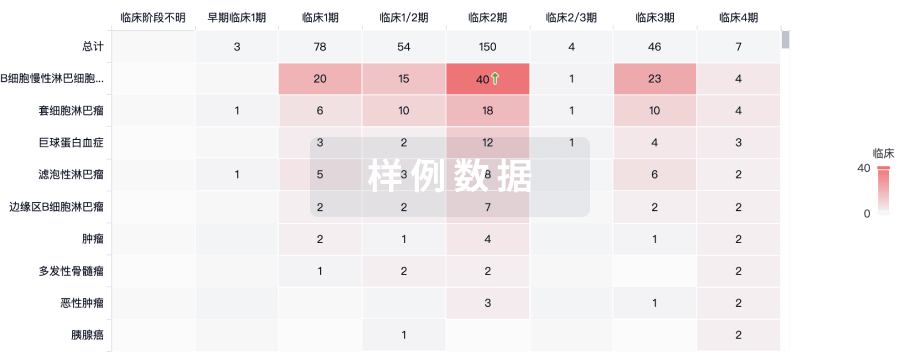
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
批准
利用最新的监管批准信息加速您的研究。
登录
或
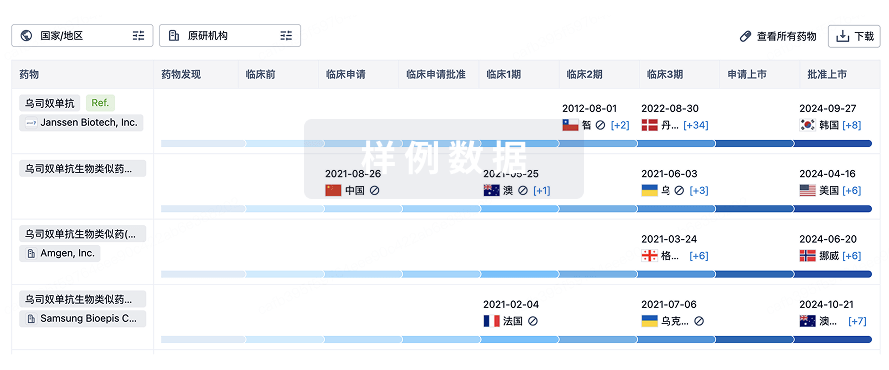
生物类似药
生物类似药在不同国家/地区的竞争态势。请注意临床1/2期并入临床2期,临床2/3期并入临床3期
登录
或
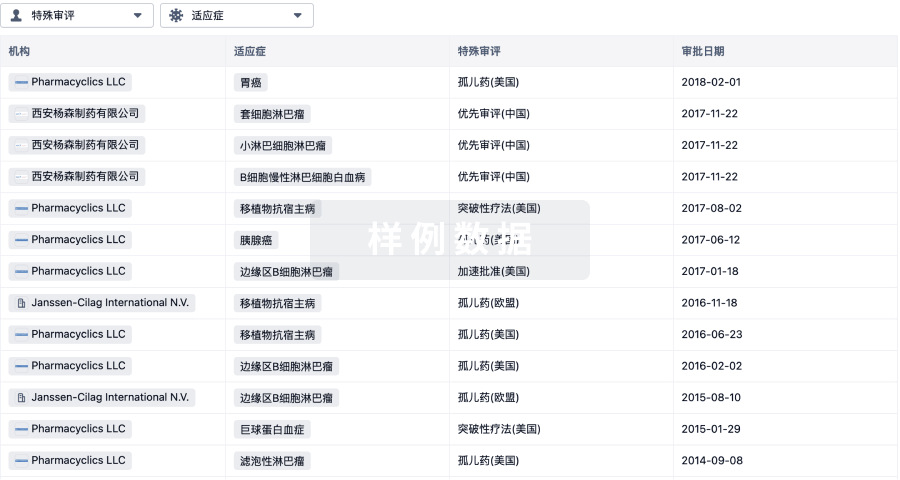
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用