预约演示
更新于:2025-11-29
NCD38
更新于:2025-11-29
概要
基本信息
非在研机构- |
权益机构- |
最高研发阶段临床前 |
首次获批日期- |
最高研发阶段(中国)- |
特殊审评- |
结构/序列
分子式C35H36ClN3O2 |
InChIKeyKYDFMNLSOYZOTR-XFCANUNOSA-N |
CAS号2078047-41-1 |
关联
100 项与 NCD38 相关的临床结果
登录后查看更多信息
100 项与 NCD38 相关的转化医学
登录后查看更多信息
100 项与 NCD38 相关的专利(医药)
登录后查看更多信息
11
项与 NCD38 相关的文献(医药)2024-10-01·MOLECULAR CARCINOGENESIS
KDM1A/LSD1 inhibition enhances chemotherapy response in ovarian cancer
Article
作者: Johnson, Jessica D. ; Venkata, Prabhakar P. ; Sareddy, Gangadhara R. ; He, Yi ; Alejo, Salvador ; Viswanadhapalli, Suryavathi ; Chen, Yihong ; Zou, Yi ; Jayamohan, Sridharan ; Kost, Edward ; Vadlamudi, Ratna K. ; Lai, Zhao ; Jamwal, Diksha
Abstract:
Ovarian cancer (OCa) is the deadliest of all gynecological cancers. The standard treatment for OCa is platinum‐based chemotherapy, such as carboplatin or cisplatin in combination with paclitaxel. Most patients are initially responsive to these treatments; however, nearly 90% will develop recurrence and inevitably succumb to chemotherapy‐resistant disease. Recent studies have revealed that the epigenetic modifier lysine‐specific histone demethylase 1A (KDM1A/LSD1) is highly overexpressed in OCa. However, the role of KDM1A in chemoresistance and whether its inhibition enhances chemotherapy response in OCa remains uncertain. Analysis of TCGA datasets revealed that KDM1A expression is high in patients who poorly respond to chemotherapy. Western blot analysis show that treatment with chemotherapy drugs cisplatin, carboplatin, and paclitaxel increased KDM1A expression in OCa cells. KDM1A knockdown (KD) or treatment with KDM1A inhibitors NCD38 and SP2509 sensitized established and patient‐derived OCa cells to chemotherapy drugs in reducing cell viability and clonogenic survival and inducing apoptosis. Moreover, knockdown of KDM1A sensitized carboplatin‐resistant A2780‐CP70 cells to carboplatin treatment and paclitaxel‐resistant SKOV3‐TR cells to paclitaxel. RNA‐seq analysis revealed that a combination of KDM1A‐KD and cisplatin treatment resulted in the downregulation of genes related to epithelial‐mesenchymal transition (EMT). Interestingly, cisplatin treatment increased a subset of NF‐κB pathway genes, and KDM1A‐KD or KDM1A inhibition reversed this effect. Importantly, KDM1A‐KD, in combination with cisplatin, significantly reduced tumor growth compared to a single treatment in an orthotopic intrabursal OCa xenograft model. Collectively, these findings suggest that combination of KDM1A inhibitors with chemotherapy could be a promising therapeutic approach for the treatment of OCa.
2023-10-01·Cancer letters
Pharmacological inhibition of KDM1A/LSD1 enhances estrogen receptor beta-mediated tumor suppression in ovarian cancer
Article
作者: Alejo, Salvador ; He, Yi ; Lai, Zhao ; Liu, Zexuan ; Vadlamudi, Ratna K ; Weintraub, Susan T ; Chen, Yihong ; Kost, Edward R ; Venkata, Prabhakar Pitta ; Palakurthi, Srinath ; Tekmal, Rajeshwar R ; Zou, Yi ; Viswanadhapalli, Suryavathi ; Johnson, Jessica D ; Jayamohan, Sridharan ; Suzuki, Takayoshi ; Pratap, Uday P ; Palacios, Bridgitte E ; Sareddy, Gangadhara R ; Valente, Philip T
Ovarian cancer (OCa) is the most lethal gynecologic cancer. Emerging data indicates that estrogen receptor beta (ERβ) functions as a tumor suppressor in OCa. Lysine-specific histone demethylase 1A (KDM1A) is an epigenetic modifier that acts as a coregulator for steroid hormone receptors. However, it remain unknown if KDM1A interacts with ERβ and regulates its expression/functions in OCa. Analysis of TCGA data sets indicated KDM1A and ERβ expression showed an inverse relationship in OCa. Knockout (KO), knockdown (KD), or inhibition of KDM1A increased ERβ isoform 1 expression in established and patient-derived OCa cells. Further, KDM1A interacts with and functions as a corepressor of ERβ, and its inhibition enhances ERβ target gene expression via alterations of histone methylation marks at their promoters. Importantly, KDM1A-KO or -KD enhanced the efficacy of ERβ agonist LY500307, and the combination of KDM1A inhibitor (KDM1Ai) NCD38 with ERβ agonist synergistically reduced the cell viability, colony formation, and invasion of OCa cells. RNA-seq and DIA mass spectrometry analyses showed that KDM1A-KO resulted in enhanced ERβ signaling and that genes altered by KDM1A-KO and ERβ agonist were related to apoptosis, cell cycle, and EMT. Moreover, combination treatment significantly reduced the tumor growth in OCa orthotopic, syngeneic, and patient-derived xenograft models and proliferation in patient-derived explant models. Our results demonstrate that KDM1A regulates ERβ expression/functions, and its inhibition improves ERβ mediated tumor suppression. Overall, our findings suggest that KDM1Ai and ERβ agonist combination therapy is a promising strategy for OCa.
2021-01-01·Breast cancer research and treatment2区 · 医学
KDM1A inhibition is effective in reducing stemness and treating triple negative breast cancer
2区 · 医学
Article
作者: Sareddy, Gangadhara R ; He, Yi ; Lai, Zhao ; Alejo, Salvador ; Chen, Yihong ; Palacios, Bridgitte ; Liu, Junhao ; Tekmal, Rajeshwar R ; Vadlamudi, Ratna K ; Zou, Yi ; Venkata, Prabhakar Pitta ; Suzuki, Takayoshi ; Viswanadhapalli, Suryavathi ; Pratap, Uday P ; Brenner, Andrew J ; Zhou, Mei
PURPOSE:
Cancer stem cells (CSCs) are highly tumorigenic, spared by chemotherapy, sustain tumor growth, and are implicated in tumor recurrence after conventional therapies in triple negative breast cancer (TNBC). Lysine-specific histone demethylase 1A (KDM1A) is highly expressed in several human malignancies and CSCs including TNBC. However, the precise mechanistic role of KDM1A in CSC functions and therapeutic utility of KDM1A inhibitor for treating TNBC is poorly understood.
METHODS:
The effect of KDM1A inhibition on cell viability, apoptosis, and invasion were examined by Cell Titer Glo, Caspase 3/7 Glo, and matrigel invasion assays, respectively. Stemness and self-renewal of CSCs were examined using mammosphere formation and extreme limiting dilution assays. Mechanistic studies were conducted using RNA-sequencing, RT-qPCR, Western blotting and reporter gene assays. Mouse xenograft and patient derived xenograft models were used for preclinical evaluation of KDM1A inhibitor.
RESULTS:
TCGA data sets indicated that KDM1A is highly expressed in TNBC. CSCs express high levels of KDM1A and inhibition of KDM1A reduced the CSCs enrichment in TNBC cells. KDM1A inhibition reduced cell viability, mammosphere formation, self-renewal and promoted apoptosis of CSCs. Mechanistic studies suggested that IL6-JAK-STAT3 and EMT pathways were downregulated in KDM1A knockdown and KDM1A inhibitor treated cells. Importantly, doxycycline inducible knockout of KDM1A reduced tumor progression in orthotopic xenograft models and KDM1A inhibitor NCD38 treatment significantly reduced tumor growth in patient derived xenograft (PDX) models.
CONCLUSIONS:
Our results establish that KDM1A inhibition mitigates CSCs functions via inhibition of STAT3 and EMT signaling, and KDM1A inhibitor NCD38 may represent a novel class of drug for treating TNBC.
100 项与 NCD38 相关的药物交易
登录后查看更多信息
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 肿瘤 | 临床前 | 日本 | 2013-07-03 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
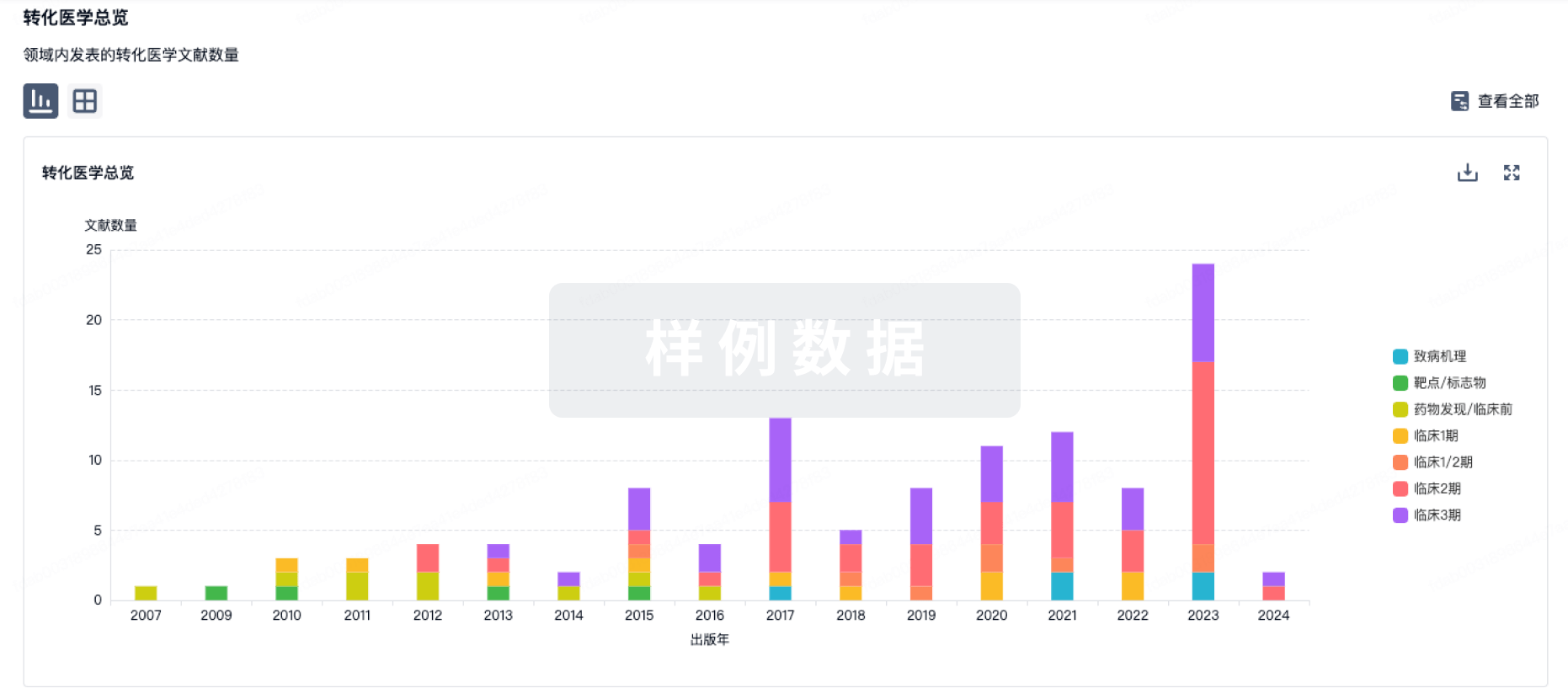
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
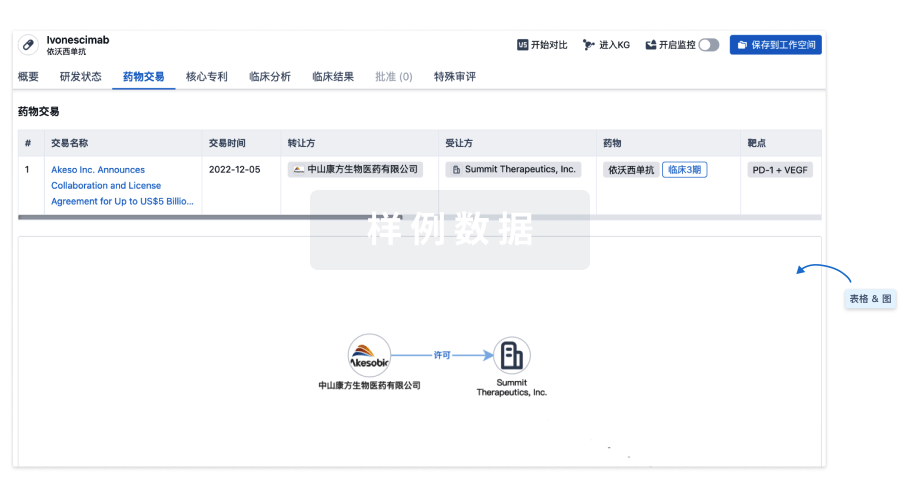
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
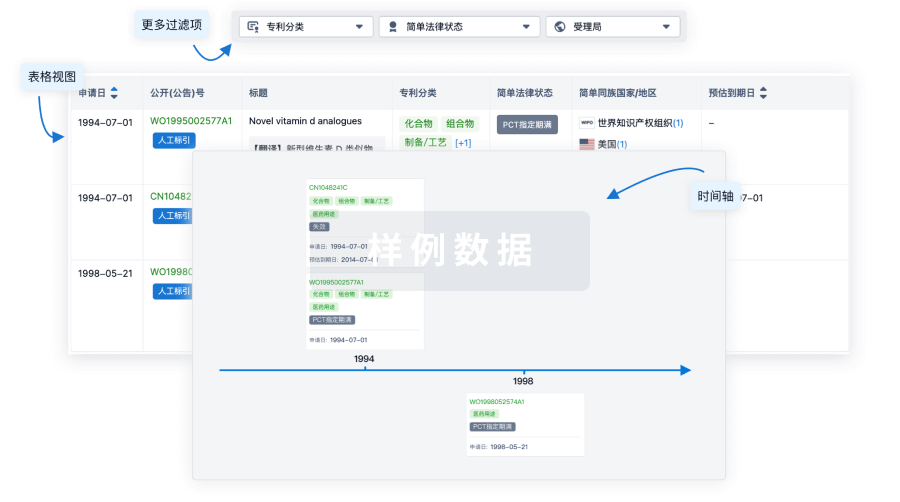
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
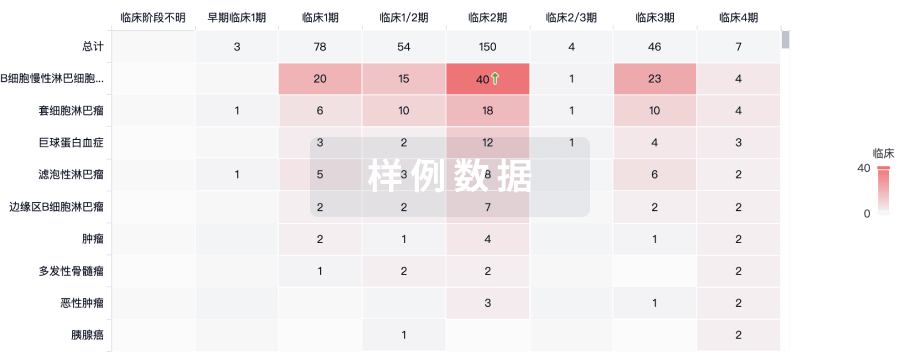
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
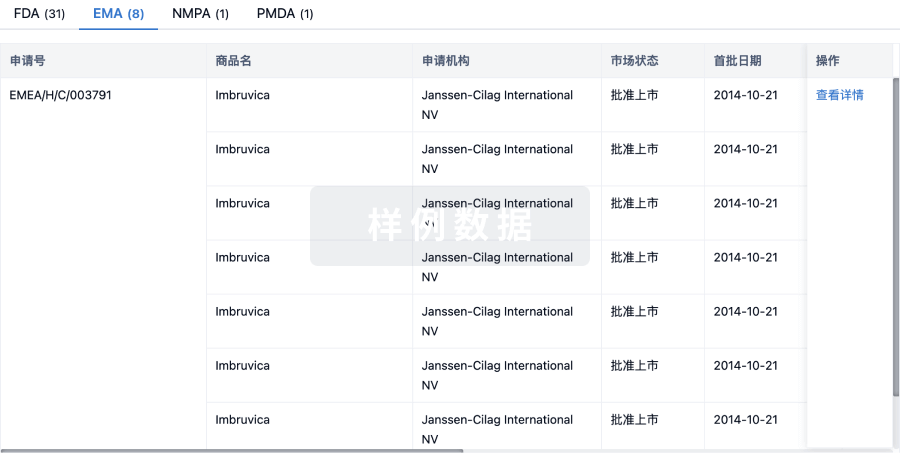
批准
利用最新的监管批准信息加速您的研究。
登录
或
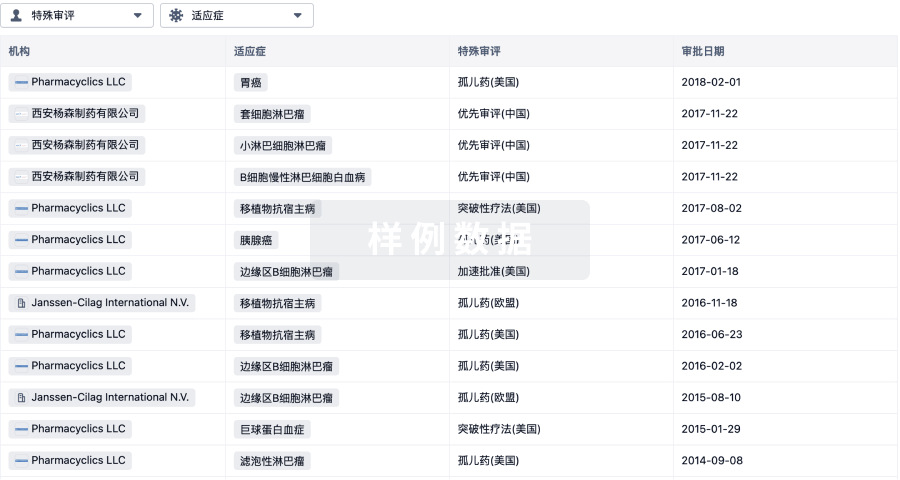
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
