预约演示
更新于:2025-05-07

Bombay Hospital, Indore
更新于:2025-05-07
概览
关联
100 项与 Bombay Hospital, Indore 相关的临床结果
登录后查看更多信息
0 项与 Bombay Hospital, Indore 相关的专利(医药)
登录后查看更多信息
807
项与 Bombay Hospital, Indore 相关的文献(医药)2024-11-01·Autoimmunity Reviews
Indian Rheumatology Association guidelines for the management of ANCA associated vasculitis
Review
作者: Dharmanand, B.G. ; Samanta, Joydeep ; Chattopadhyay, Arghya ; Jha, Saket ; Acharya, Nupoor ; Kumar, Uma ; Adarsh, M B ; Sharma, Aman ; Pinto, Benzeeta ; Balakrishnan, Canchi ; Jain, Siddharth ; Mishra, Debashish ; Dhooria, Sahajal ; Naidu, G S R S N K ; Dhooria, Aadhaar ; Kumar, Rajiv Ranjan ; Handa, Rohini ; Mittal, Sakshi ; Naidu, G.S.R.S.N.K. ; Sharma, Vikas ; Adarsh, M.B. ; Misra, Durga Prasanna ; Dharmanand, B G ; Shobha, Vineeta ; Kavadichanda, Chengappa ; Jois, Ramesh ; Sharma, Banwari ; Ramachandran, Raja ; Agarwal, Vikas
2024-05-01·The Lancet Regional Health - Southeast Asia
Treatment pattern and outcomes of leptomeningeal carcinomatosis in India – a retrospective study
Article
作者: Kothari, Rushabh Kiran ; Raut, Nirmal Vivek ; Parekh, Deevyashali ; Singh, Ashish ; Chandrasekharan, Arun ; Goyal, Gautam ; Talreja, Vikas ; Shrirangwar, Sameer ; Talele, Avinash ; Gupta, Kushal ; Bhosale, Bharat ; Avaronnan, Manuprasad ; Ghosh, Joydeep ; Poladia, Bhavesh Pradip ; Karpe, Akshay
2024-04-02·Health Systems
Long term simulation analysis of deceased donor initiated chains in kidney exchange programs
Article
作者: Rangaraj, Nayaran ; Verma, Utkarsh ; Billa, Viswanath ; Usulumarty, Deepa
1
项与 Bombay Hospital, Indore 相关的新闻(医药)2023-11-30
SAN DIEGO, Nov. 30, 2023 (GLOBE NEWSWIRE) -- Bionano Laboratories today announced the publication by scientists at Revvity Omics (formerly Perkin Elmer Genomics), Leiden University Medical Centre, Bombay Hospital, and UT Dallas, of the largest peer-reviewed study to date on the use of optical genome mapping (OGM) to diagnose facioscapulohumeral muscular dystrophy (FSHD). The publication describes the evaluation of an OGM-based laboratory developed test (LDT) including the assessment of the accuracy and precision of the OGM test against the prior standard, Southern blot analysis by gel electrophoresis. The study authors assessed the yield of OGM for diagnosing FSHD1 and the ability of next generation sequencing (NGS) to be used as a reflex test to identify FSHD2. The study, which included 547 cases with a clinical suspicion of FSHD, used an algorithm that utilized OGM to identify the FSHD haplotype and quantitate the number of D4Z4 repeats found on chromosome 4. Researchers used the NGS LDT to identify sequence variants, using NxClinical software to detect copy number variants in the SMCHD1 gene to diagnose cases with FSHD2. The authors noted that, when compared to other methods, OGM has the advantage of being able to identify mosaicism and to detect repeat sizing on 4q and 10q together with haplotyping in a single run. Key findings: Compared to Southern blot, the OGM LDT was 100% accurate and preciseThe OGM LDT identified 56% of cases positive for FSHD1 (308 out of 547 samples)252/547 cases were referred for concurrent testing for FSHD1 and FSHD2Sequencing identified FSHD2 in 3.6% of cases (9 out of 252 samples)The OGM LDT detected mosaic alleles with at least one contracted 4qA allele in 3% of samples positive for FSHD1 (9 out of 308 samples)The overall diagnostic yield of OGM and NGS combined was 58% (317 out of 547 samples) “Bionano Laboratories has developed a powerful menu of OGM-based LDTs, including one LDT for FSHD1 diagnosis. We are pleased to see this prestigious group of researchers’ findings from the largest FSHD study to date utilizing OGM. Using the study’s algorithm, researchers may be able to diagnose most cases of FSHD, underscoring OGM’s potential to contribute to diagnosis of the disorder, which may lead to better disease management and outcomes,” commented Justin Leighton, vice president of laboratory business at Bionano Laboratories. The publication can be viewed here. About Bionano Laboratories: Bionano Laboratories provides access to genetic answers and support utilizing cutting-edge technologies to advance the way you see the genome. Our clinical services offer a genetic testing experience that combines a comprehensive testing portfolio with thoughtful and accessible support options for the diagnostic journey. Bionano Laboratories also offers direct access to optical genome mapping for applications across basic, translational and clinical research. For more information, visit www.bionanolaboratories.com Forward-Looking Statements of Bionano Genomics This press release contains forward-looking statements within the meaning of the Private Securities Litigation Reform Act of 1995. Words such as “may,” “potential,” “would” and similar expressions (as well as other words or expressions referencing future events, conditions or circumstances) convey uncertainty of future events or outcomes and are intended to identify these forward-looking statements. Forward-looking statements include statements regarding our intentions, beliefs, projections, outlook, analyses or current expectations concerning, among other things, the ability and utility of the OGM for use in the detection or diagnosis of FSHD; and the ability of OGM-based LDTs to remove barriers for OGM adoption in clinical and research settings. Each of these forward-looking statements involves risks and uncertainties. Actual results or developments may differ materially from those projected or implied in these forward-looking statements. Factors that may cause such a difference include the risks and uncertainties associated with: the impact of adverse geopolitical and macroeconomic events, such as recent and potential future bank failures, potential global pandemics and the ongoing conflicts between Ukraine and Russia and Israel and Hamas, on our business and the global economy; general market conditions; the failure of the OGM-LDTs to prove useful for the detection or diagnosis of FSHD; the failure of OGM-based LDTs to remove barriers for OGM adoption in clinical and research settings; the ability of our OGM solutions to offer the anticipated benefits for and contributions to the areas reported in the study results referenced in this press release; future study results contradicting the study results reported in the publication referenced in this press release; general market conditions; changes in the competitive landscape and the introduction of competitive technologies or improvements to existing technologies; changes in our strategic and commercial plans; our ability to obtain sufficient financing to fund our strategic plans and commercialization efforts and the ability of our parent corporation, Bionano Genomics, Inc., to continue as a “going concern”; the ability of medical and research institutions to obtain funding to support adoption or continued use of our services and technologies; and the risks and uncertainties associated with our business and financial condition in general, including the risks and uncertainties described in the filings of our parent corporation, Bionano Genomics, Inc., with the Securities and Exchange Commission, including, without limitation, its Annual Report on Form 10-K for the year ended December 31, 2022 and in other filings subsequently made by them with the Securities and Exchange Commission. All forward-looking statements contained in this press release speak only as of the date on which they were made and are based on management’s assumptions and estimates as of such date. We do not undertake any obligation to publicly update any forward-looking statements, whether as a result of the receipt of new information, the occurrence of future events or otherwise. CONTACTS Company Contact:Erik Holmlin, CEOBionano Genomics, Inc.+1 (858) 888-7610eholmlin@bionano.com Investor Relations:David HolmesGilmartin Group+1 (858) 888-7625IR@bionano.com
临床结果临床研究
100 项与 Bombay Hospital, Indore 相关的药物交易
登录后查看更多信息
100 项与 Bombay Hospital, Indore 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年02月08日管线快照
无数据报导
登录后保持更新
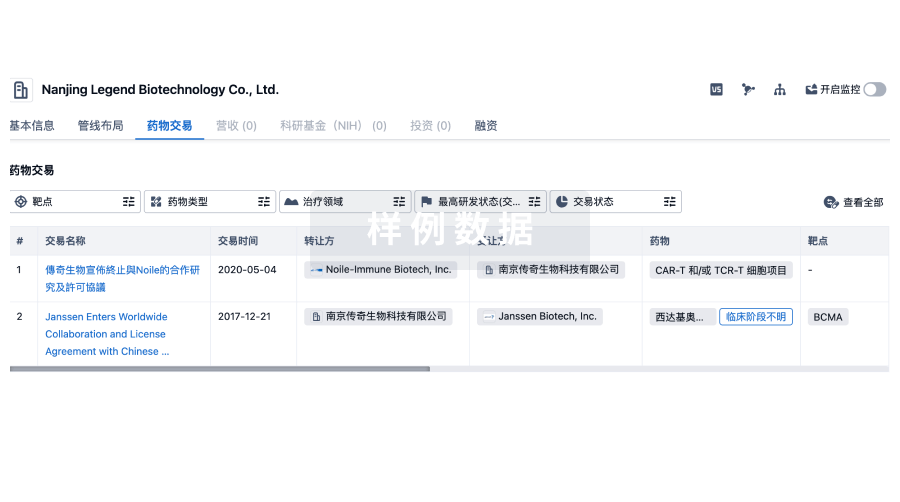
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
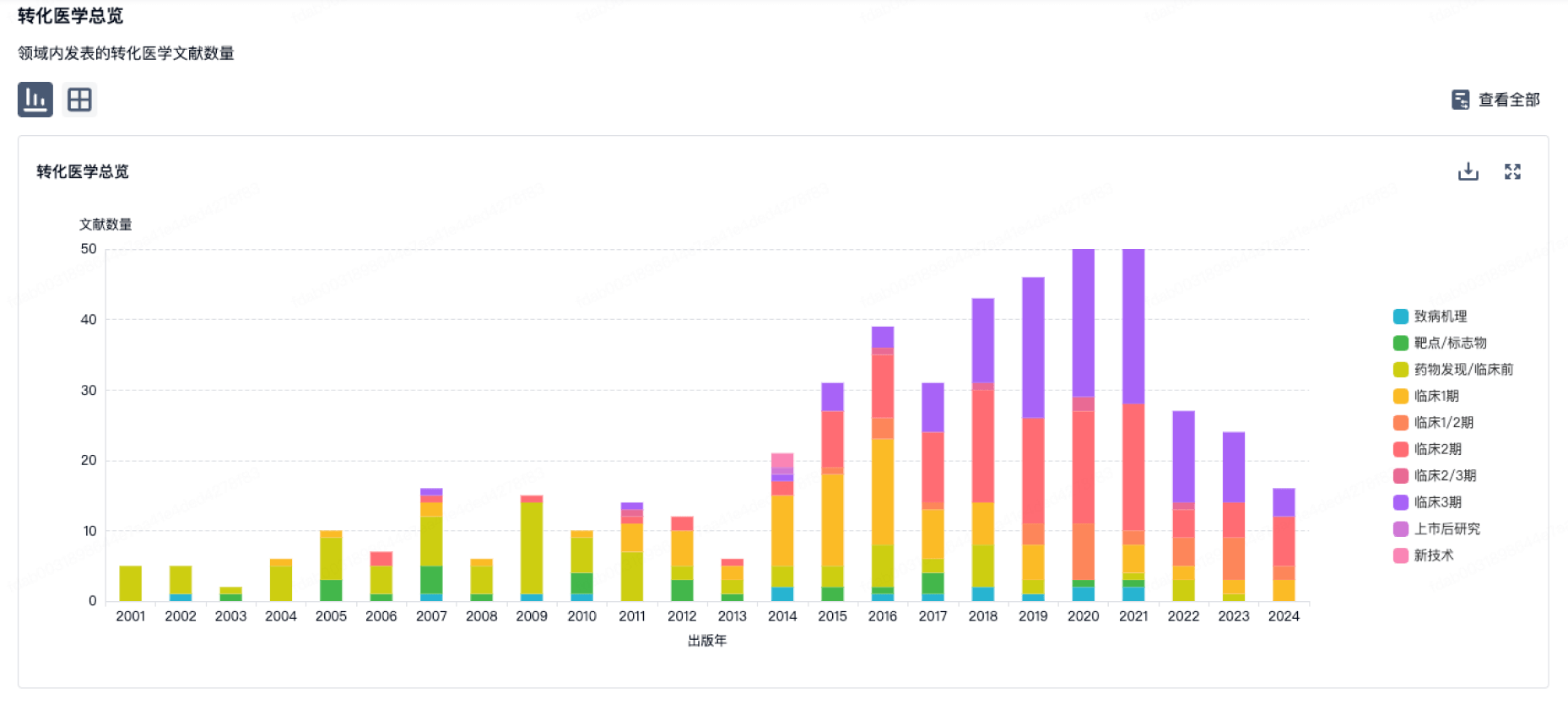
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
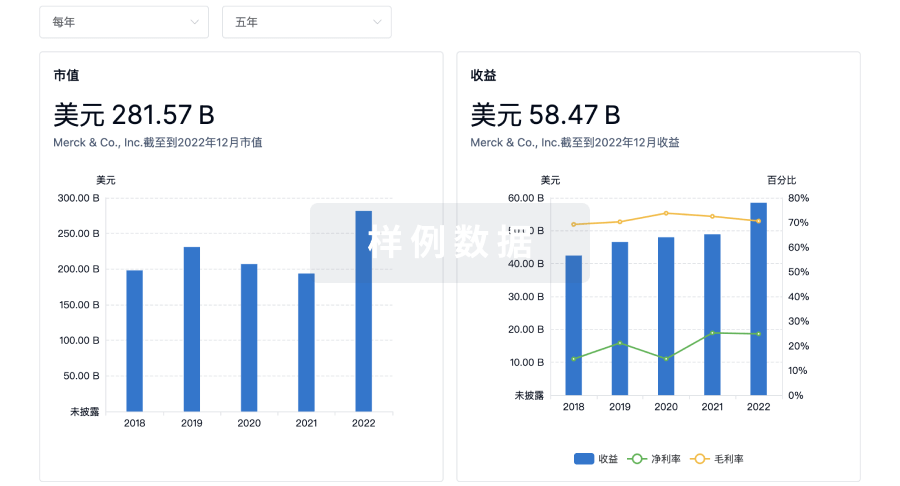
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
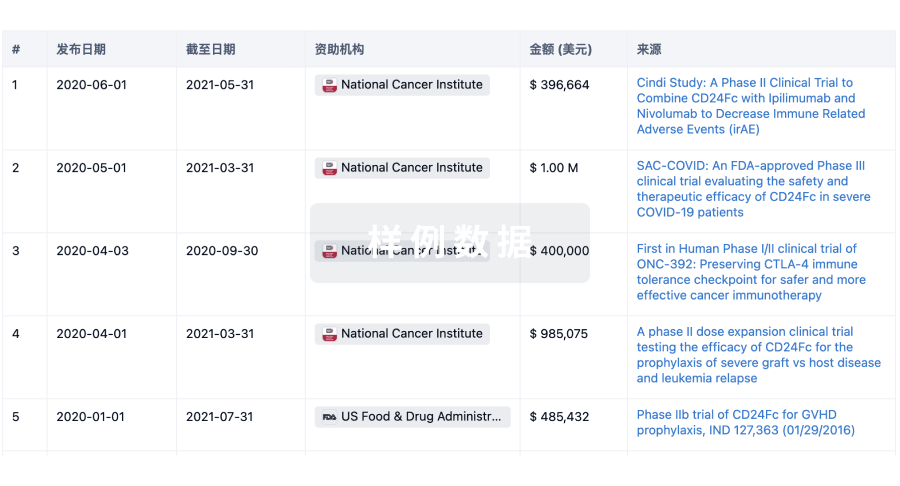
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
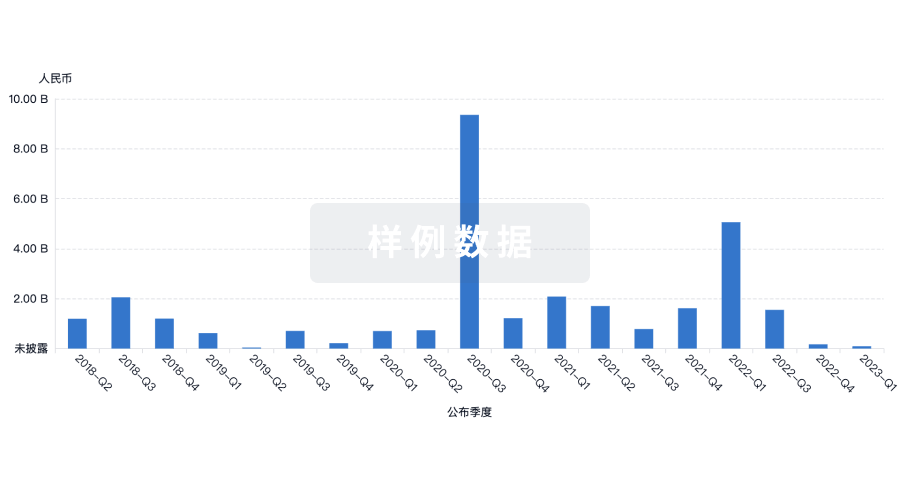
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
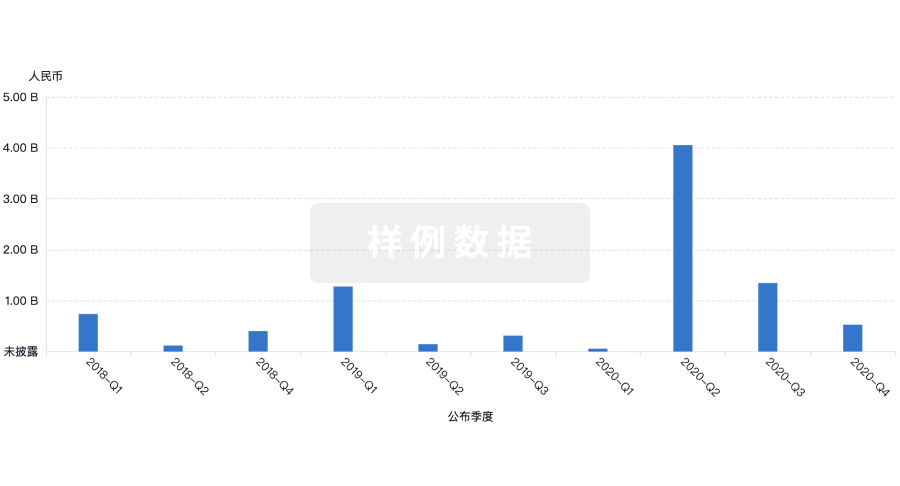
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用