预约演示
更新于:2025-05-07
GSTA1
更新于:2025-05-07
基本信息
别名 13-hydroperoxyoctadecadienoate peroxidase、Androst-5-ene-3,17-dione isomerase、Glutathione S-transferase A1 + [8] |
简介 Glutathione S-transferase that catalyzes the nucleophilic attack of the sulfur atom of glutathione on the electrophilic groups of a wide range of exogenous and endogenous compounds (Probable). Involved in the formation of glutathione conjugates of both prostaglandin A2 (PGA2) and prostaglandin J2 (PGJ2) (PubMed:9084911). It also catalyzes the isomerization of D5-androstene-3,17-dione (AD) into D4-androstene-3,17-dione and may therefore play an important role in hormone biosynthesis (PubMed:11152686). Through its glutathione-dependent peroxidase activity toward the fatty acid hydroperoxide (13S)-hydroperoxy-(9Z,11E)-octadecadienoate/13-HPODE it is also involved in the metabolism of oxidized linoleic acid (PubMed:16624487). |
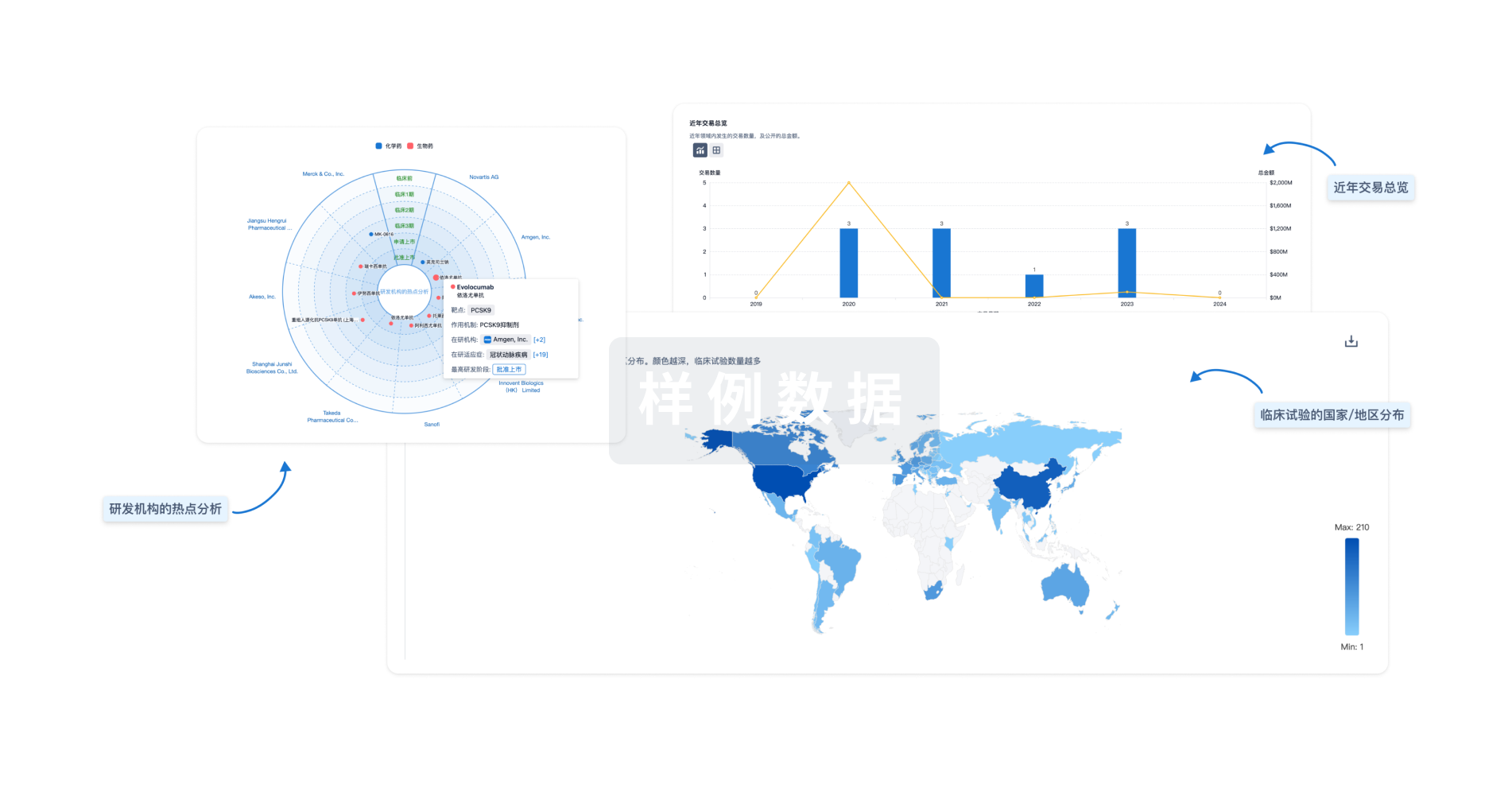
分析
对领域进行一次全面的分析。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用