预约演示
更新于:2025-05-07
RTCB
更新于:2025-05-07
基本信息
别名 3'-phosphate/5'-hydroxy nucleic acid ligase、C22orf28、FAAP + [4] |
简介 Catalytic subunit of the tRNA-splicing ligase complex that acts by directly joining spliced tRNA halves to mature-sized tRNAs by incorporating the precursor-derived splice junction phosphate into the mature tRNA as a canonical 3',5'-phosphodiester. May act as an RNA ligase with broad substrate specificity, and may function toward other RNAs. |
关联
100 项与 RTCB 相关的临床结果
登录后查看更多信息
100 项与 RTCB 相关的转化医学
登录后查看更多信息
0 项与 RTCB 相关的专利(医药)
登录后查看更多信息
174
项与 RTCB 相关的文献(医药)2025-04-01·Journal of Biological Chemistry
Structural and biochemical characterization of the 3'-5' tRNA splicing ligases
Article
作者: Sroka, Małgorzata ; Czarnocki-Cieciura, Mariusz ; Jemielity, Jacek ; Śmietański, Mirosław ; Wycisk, Krzysztof ; Chamera, Sebastian ; Koziej, Łukasz ; Nowak, Jakub ; Chramiec-Głąbik, Andrzej ; Nowotny, Marcin ; Jaciuk, Marcin ; Warmiński, Marcin ; Gołębiowski, Filip ; Zajko, Weronika ; Glatt, Sebastian
2025-03-25·mSphere
Characterization of diet-linked amino acid pool influence on
Fusobacterium
spp. growth and metabolism
Article
作者: Allen-Vercoe, Emma ; Vancuren, Sarah J. ; Marcone, Massimo ; Robinson, Avery V.
2024-10-14·Nucleic Acids Research
RTP801 interacts with the tRNA ligase complex and dysregulates its RNA ligase activity in Alzheimer’s disease
Article
作者: Pérez-Navarro, Esther ; Guisado-Corcoll, Anna ; Torres, Adrian Gabriel ; Pérez-Sisqués, Leticia ; Canal, Mercè ; Garcia-Segura, Pol ; Malagelada, Cristina ; Molina-Porcel, Laura ; Campoy-Campos, Genís ; Santamaria, Enrique ; Fernández-Irigoyen, Joaquín ; Martí, Eulàlia ; Giralt, Albert ; de Pouplana, Lluís Ribas ; Chicote-González, Almudena ; Solana-Balaguer, Julia ; Alberch, Jordi
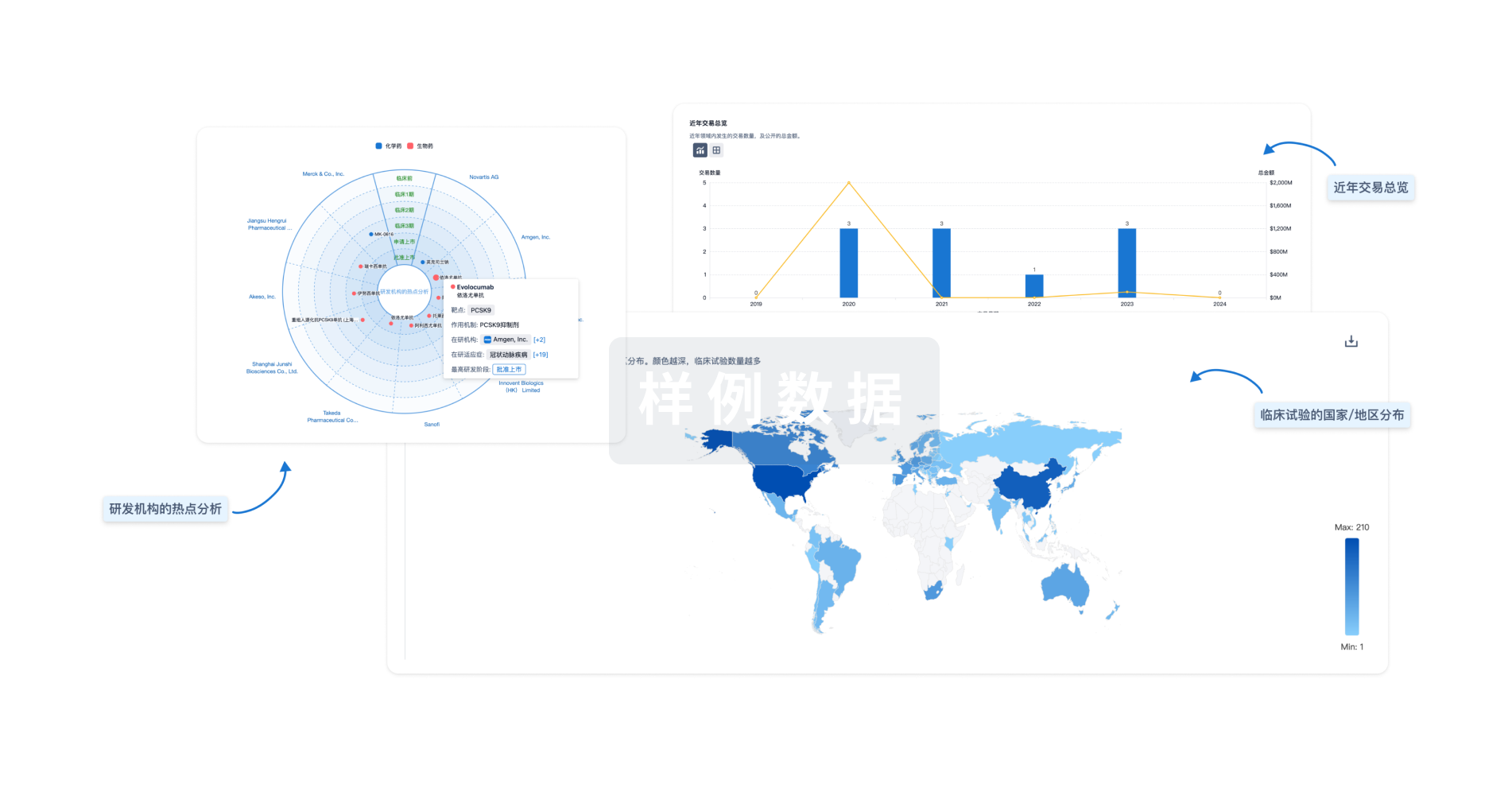
分析
对领域进行一次全面的分析。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用