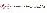预约演示
更新于:2026-02-27
Dimenhydrinate
茶苯海明
更新于:2026-02-27
概要
基本信息
简介苯海拉明 (DPH) 是一种抗组胺剂和镇静剂,主要用于治疗过敏、失眠和普通感冒症状。 它可以口服、静脉注射、肌肉注射或涂在皮肤上。 最大效果通常在服药后两小时左右出现,效果可持续长达七小时。 它是第一代 H1 抗组胺剂和乙醇胺,通过阻断组胺的某些作用发挥作用,从而产生抗组胺剂和镇静作用。 苯海拉明也是一种有效的抗胆碱能药。 常见的副作用包括嗜睡、协调性差和胃部不适。 |
药物类型 小分子化药 |
别名 (O-benzhydryl(dimethylamino)ethanol) 8-chlorotheophyllinate、Benzhydryl-β-dimethylaminoethylether 8-chlorotheophylline、Dimenhydrinate (JP17/USP/INN) + [11] |
作用方式 拮抗剂 |
作用机制 H1 receptor拮抗剂(组胺H1受体拮抗剂) |
非在研适应症- |
非在研机构- |
权益机构- |
最高研发阶段批准上市 |
最高研发阶段(中国)批准上市 |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C24H28ClN5O3 |
InChIKeyNFLLKCVHYJRNRH-UHFFFAOYSA-N |
CAS号523-87-5 |
关联
20
项与 茶苯海明 相关的临床试验CTIS2025-523948-12-00
Bioequivalence of Cinnarizine/Dimenhydrinate 20 mg/40 mg Tablets in Healthy Participants Under Fed Conditions.
开始日期2025-11-27 |
CTIS2024-515170-27-00
Evaluation of bioequivalence of two products containing 50 mg dimenhydrinate: Dimenhydrinate 50 mg (Test) vs. Vomex A 50 mg Lösung zum Einnehmen im Beutel (Comparator). A monocentric, open, randomized, single dose, two-period, crossover trial in healthy volunteers
开始日期2024-11-15 |
申办/合作机构 |
NCT06395064
The Effect of Dimenhydrinate on Postoperative Nausea and Vomiting After Abdominal Hysterectomy: Randomized-controlled Trial
The aim of this clinical trial is to learn if Dimenhydrinate works to prevent postoperative nausea-vomiting in adult women who undergoing abdominal hysterectomy. The primary question is: Does Dimenhydrinate lower the proportion of postoperative nauseavomiting during the day 1 after surgery. Researcher will compare Dimenhydrinate to a placebo (a-look-alike substance that contains no drug) which will give intravenously when the patient come back to the ward, to see if Dimenhydrinate work to prevent nauseavomiting during the postoperative day1. All participants will receive standard treatment during pre-operative, intra-operative and the immediate post-operative periods. Participants will receive Dimenhydrinate or placebo only one dose after discharge from the recovery room. During the day 1 postoperative, participants will report their symptoms and keep a diary of the number of tmes they use additional anti-emetic drugs.
开始日期2024-05-01 |
申办/合作机构 |
100 项与 茶苯海明 相关的临床结果
登录后查看更多信息
100 项与 茶苯海明 相关的转化医学
登录后查看更多信息
100 项与 茶苯海明 相关的专利(医药)
登录后查看更多信息
640
项与 茶苯海明 相关的文献(医药)2025-11-01·CURRENT PHARMACEUTICAL DESIGN
Design, Molecular Docking, In Vitro and In Vivo Evaluation of Dimenhydrinate-Cyclodextrin Complex for Fast Disintegrating Tablet
Article
作者: Abdul Rasool, Bazigha K. ; Seidel, Veronique ; Sammour, Rana M.F. ; Samara, Randa Khalid
Introduction::
This study aimed to formulate and evaluate dimenhydrinate (DMH) as fastdisintegrating
tablets (FDTs) complexed with β-cyclodextrin (β-CD) to enhance its solubility, dissolution profile,
and pharmacological performance.
Methods::
A DMH:β-CD inclusion complex was prepared at a 1:1 molar ratio using the kneading method.
Characterization was performed through phase solubility studies, FTIR analysis, molecular docking, and in
vitro dissolution testing. FDTs were developed using various superdisintegrants and assessed for quality attributes
of a tablet, including hardness, friability, wetting time, water absorption ratio, and drug content.
Results::
Phase solubility and FTIR analyses confirmed the formation of a stable DMH:β-CD complex. Molecular
docking indicated a binding affinity of -4.2 kcal/mol between β-CD and diphenhydramine. Among the
FDT formulations, CP3 containing 9% crospovidone showed the best performance, with a disintegration time
of 4.3 seconds and the highest drug release rate. In vivo pharmacological tests demonstrated enhanced sedative
and antiemetic activities of the optimized FDTs compared to conventional DMH formulations.
Discussion::
The findings suggest that cyclodextrin-based complexation combined with orodispersible tablet
technology can significantly enhance DMH's pharmacological efficacy and patient compliance. However, additional
investigations on long-term stability, pharmacokinetics, and clinical scalability are warranted.
Conclusion::
The DMH:β-CD FDTs developed in this study offer promising improvements in solubility, dissolution,
and therapeutic performance, indicating their potential for better clinical outcomes and patient acceptability.
2025-11-01·Acta otorrinolaringologica espanola
Efficacy and safety of the cinnarizine/dimenhydrinate combination versus betahistine in the treatment of vertigo: A systematic literature review
Review
作者: Gómez Gabaldón, Niceto ; Martín-Enguix, David ; Amaro-Gahete, Francisco J
Vertigo is a frequent reason for medical consultation and may result from a wide range of aetiologies. Betahistine and the fixed low-dose combination of cinnarizine 20 mg and dimenhydrinate 40 mg are commonly used therapeutic options, each with distinct antivertigo profiles. This systematic review, conducted in accordance with the PRISMA guidelines, aims to compare the efficacy and safety of these two treatments in patients with vertigo of various origins. A comprehensive search was conducted in PubMed, Cochrane Library, Google Scholar, and ClinicalTrials.gov, with no restrictions on language or publication date. Eligible studies included clinical trials and meta-analyses comparing the fixed low-dose combination (20 mg/40 mg) versus betahistine (12 or 16 mg), assessing efficacy through the Mean Vertigo Score (MVS) and safety based on the incidence of adverse events (AEs). The RoB 2 and ROBIS tools were used to evaluate the risk of bias. A total of nine studies were identified (six clinical trials and three meta-analyses). In five of the six clinical trials, the fixed low-dose combination significantly reduced MVS compared with betahistine at weeks 1 and/or 4 (p < .05); these findings were corroborated by the three meta-analyses. Regarding safety, both treatments were well tolerated, with no serious AEs reported and a generally lower incidence observed in the fixed low-dose combination group. Overall, the fixed low-dose combination demonstrated superior clinical efficacy from the first week of treatment, along with a more favourable tolerability and safety profile. These results support its preferential use in the management of acute vestibular syndrome.
2025-10-01·ECOTOXICOLOGY AND ENVIRONMENTAL SAFETY
From waterways to the brain: Unraveling the environmental triggers of depression through PPCPs-gene network convergence
Article
作者: Yu, Hao ; Zhang, Guanglei ; Che, Ke ; Wang, Cong
Pharmaceutical and personal care products (PPCPs), as ubiquitous emerging contaminants, present undercharacterized neuropsychiatric hazards through environmental exposure. This investigation employs convergent multi-omics strategies - integrating toxicogenomic discovery, disease-associated genomic mapping, and transcriptomic profiling - to elucidate mechanistic linkages between PPCPs bioactivity and depressive pathogenesis. Through systematic analysis of Nanjing's aquatic chemical burden (prioritizing dimenhydrinate, ibuprofen, padimate-O, caffeine, and roxithromycin), we identified 3073 conserved molecular targets bridging PPCPs toxicity and depression etiology via Comparative Toxicogenomics Database and GeneCards interrogation. Functional ontology revealed dysregulated pathways encompassing lipidomic remodeling, IL-17-mediated neuroinflammation, and synaptic transmission deficits. Ensembled machine learning algorithms (Lasso regression, XGBoost, random forest) converged on seven high-fidelity candidate biomarkers (HSPA8, CBX1, CD59, CHAF1A, CUX1, ID2, RPL3) demonstrating stress-adaptive, chromatin regulatory, and immunomodulatory functions. Molecular docking predicted strong binding affinities between PPCPs and depression-related proteins, notably dimenhydrinate with CHAF1A (- 6.1 kcal/mol) and HSPA8 (- 6.1 kcal/mol), suggesting multi-target modulation. This work proposes a computational framework to map molecular interactions between specific PPCPs and depression-associated pathways. Candidate targets highlight testable hypotheses for future experimental validation. These findings suggest selected PPCPs with neuroactive properties may warrant further investigation as environmental modifiers of depression risk.
14
项与 茶苯海明 相关的新闻(医药)2026-01-02
Gargonia/
iStock
Jefferies analysts envision a steady launch curve that could ultimately drive meaningful sales from people who are dissatisfied with existing treatments.
The FDA has
approved
Vanda Pharmaceuticals’ tradipitant for the prevention of vomiting induced by motion.
The company filed for FDA approval after linking the oral neurokinin-1 receptor antagonist, which Vanda will sell as Nereus, to significant reductions in vomiting in two Phase III real-world provocation studies conducted on boats. In one trial, vomiting incidence was 18.3% to 19.5% with Nereus versus 44.3% with placebo. Vomiting rates were 10.4% to 18.3% with Nereus versus 37.7% on placebo in the second study.
Vanda said the approval marks the first time in over 40 years that a new pharmacologic treatment for motion sickness has come to market. Nereus will compete with dramamine, an over-the-counter antihistamine drug that prevents and treats motion sickness symptoms including nausea, vomiting and dizziness. Jefferies analysts identified a detail of the approval that could differentiate Nereus.
“Encouragingly, Nereus’ label is for the prevention of vomiting caused by motion, rather than the symptoms of motion sickness (e.g. nausea, vomiting, and dizziness) like dramamine,” the analysts said in a note to investors.
The label permits patients to take Nereus around 60 minutes before an event expected to cause vomiting induced by motion. Patients need to take the medicine on an empty stomach, meaning at least one hour before or two hours after a full meal, but otherwise analysts said the “front page label looks clean/simple.”
Vanda’s assessment of the market opportunity for Nereus includes two to three million U.S. patients who take dramamine every month, the analysts said. The lack of innovation in the motion sickness drug field means there may be some pent-up demand for Nereus.
“We envision a steady launch curve as awareness builds over time, with some contribution from patients who have been waiting for an alternative to existing therapies,” the analysts said. “A modest penetration among dissatisfied users or patients who currently avoid travel (due to inadequate symptom control) could drive meaningful sales in theory.”
Vanda is yet to share the list price for Nereus, but Jefferies analysts expect a premium over existing OTC treatments. The company hopes to be well prepared for launch ahead of the 2026 summer season, the analysts said, and plans to focus on digital advertising and direct-to-consumer outreach rather than building a large prescriber‑focused sales force.
The analysts’ overview of the launch plan aligns with comments by Vanda CEO Mihael Polymeropoulos. On an earnings call in October 2025, Polymeropoulos
described
the strategy for both Nereus and jet lag drug Hetlioz.
“We’re developing a quite elaborate strategy that will become very consumer-centric, focusing on concierge service for supplying the drug to both of them,” Polymeropoulos said. “Our recent experiences with direct-to-consumer campaigns, but also the elevation of brand awareness of the company, are going to be very important and have been strategically designed to be in place in advance of those launches.”
The FDA could rule on Hetlioz in jet lag by Jan. 7. The back-to-back regulatory actions follow the creation of a collaborative framework to resolve disputes between the FDA and Vanda, which
sued
the agency in April amid rejections, a partial hold and other setbacks. The FDA
lifted
the partial hold on tradipitant last month.
Vanda’s approval was one of a clutch of authorizations over the holiday period. The FDA also
approved
Agios Pharmaceuticals’ Aqvesme for the treatment of anemia in adults with alpha- or beta-thalassemia, and expanded the labels of Roche’s cancer drug
Lunsumio
and Boehringer Ingelheim’s pulmonary fibrosis medicine
Jascayd
.
临床3期上市批准临床结果
2025-12-17
·摩熵医药
注:本文不构成任何投资意见和建议,以官方/公司公告为准;本文仅作医疗健康相关药物介绍,非治疗方案推荐(若涉及),不代表平台立场。任何文章转载需要得到授权。
PART 01
周报概述
随着全球医药行业的快速发展,新药研发与创新已成为推动行业进步的重要动力。近期,根据摩熵医药数据统计,新药申请与审批获批频繁,显示出医药创新领域的活跃态势。
本文将深入分析2025年12月8日至2025年12月14日期间,国内外新药申请、临床试验批准、仿制药一致性评价等多个方面的最新进展,为用户提供全面的行业资讯。
PART 02
国内65款新药IND获批
根据摩熵医药数据库统计,2025年12月8日至12月14日期间,共有83个创新药/改良型新药临床申请/上市申请获国家药品监督管理局药品审评中心(CDE)承办(按受理号统计,不含补充申请)。其中国产药品受理号77个,进口药品受理号6个。
本周共计65款创新药/改良型新药临床试验申请获得“默示许可”,包括化学药20款,生物药41款,中药4款。
本周获批临床创新药/改良型新药部分信息速览(不含补充申请)
注:完整数据可识别“文末”二维码下载查看
PART 03
本周全球TOP10创新药研发进展
12月9日,CDE官网显示,强生的1类新药JNJ-78278343注射液拟纳入突破性治疗品种,适应症为用于接受过雄激素受体(AR)通路抑制剂和紫杉烷类化疗治疗的转移性去势抵抗性前列腺癌(mCRPC)成人患者。
截图来源:摩熵医药全球药物研发数据库
公开数据显示,该药是全球首个且目前唯一进入Ⅲ期临床阶段的KLK2靶向药物。JNJ-78278343(Pasritamig)是强生开发的一款潜在first-in-class皮下注射TCE(T 细胞桥接器)双抗,可同时靶向CD3和KLK2。其中,KLK2中文全称为人激肽释放酶2,是一种在前列腺癌细胞表面表达的蛋白,但在正常组织中表达非常有限。目前,强生正在中国、美国、日本、法国等多个国家和地区开展一项国际多中心Ⅲ期临床(CTR20254394),以评估JNJ-78278343联合最佳支持治疗用于转移性去势抵抗性前列腺癌患者的有效性和安全性,研究预计将于2028年5月完成。
同日,国家药品监督管理局(NMPA)官网显示,齐鲁制药的帕尼单抗生物类似药(QL1203)获批上市,用于联合mFOLFOX一线治疗RAS野生型转移性结直肠癌(mCRC)患者。
截图来源:摩熵医药全球药物研发数据库
原研帕尼单抗是武田与安进合作开发的一款靶向表皮生长因子受体(EGFR)的单克隆抗体,暂时未在中国获批上市,齐鲁制药的QL1203的获批因此具有重要的临床意义。EGFR是重要的原癌基因,帕尼单抗可以针对EGFR异常细胞信号通路,抑制肿瘤生长。齐鲁开展了一项III期注册性临床研究,旨在评估QL1203联合mFOLFOX6对比安慰剂联合mFOLFOX6一线治疗RAS野生型的mCRC患者的有效性和安全性。研究结果显示,在既往未接受过抗EGFR疗法的RAS/BRAF野生型mCRC患者中,QL1203联合mFOLFOX6与安慰剂联合mFOLFOX6相比,显著改善了PFS并获得更高的ORR。QL1203联合mFOLFOX6耐受性与安全性良好。QL1203在与化疗联合一线治疗结直肠癌患者的研究中展现出良好的安全性和可耐受性,安全谱与原研药物相似。
本周全球 TOP10 创新药研发进展
截图来源:摩熵医药周报
PART 04
本周全球TOP10临床试验结果
本周全球多项临床试验结果亮点纷呈。12月8日,信达生物在2025年美国血液学会年会(ASH)上以口头报告形式首次公布其自主开发的抗GPRC5D/BCMA/CD3三特异性抗体IBI3003用于复发或难治性多发性骨髓瘤(R/R MM)患者的首次人体试验的初步数据。
截图来源:摩熵医药全球药物研发数据库
IBI3003展示出良好的耐受性和可控的安全性特征,尽管当前随访时间尚短,IBI3003已经显现出令人鼓舞的初步疗效信号,特别在伴有髓外病变或既往接受抗BCMA和GPRC5D单一或双重靶向治疗的高危患者中展现了积极的疗效。具体而言,在中位随访3.25个月期间,接受≥120μg/kg治疗的患者(n=24)ORR为83.3%,其中包括4例sCR、7例VGPR和9例PR。10例伴有EMD的患者ORR达80%,9例既往接受过抗BCMA和/或抗GPRC5D治疗的患者ORR达77.8%。
本周全球 TOP10 积极/失败临床结果
截图来源:摩熵医药周报
PART 05
109款品种过评,石家庄四药领跑
根据摩熵医药数据库统计,2025年12月8日至2025年12月14日期间,共有91项仿制药申报上市/申报临床获CDE承办,其中新注册分类上市申请受理号73项(包括化药3类,4类,5.2类),新注册分类临床申请受理号4项(包括化药3类,4类),一致性评价申请14项。本周11个品种通过一致性评价(按受理号计17项),98个品种视同通过一致性评价(按受理号计152项)。本周3项生物类似物注册申报动态,包括:连云港润众制药的司美格鲁肽注射液;华兰基因工程的地舒单抗注射液;上海复宏瑞霖生物的达雷妥尤单抗注射液。
本周过评/视同过评品种主要为心血管系统药物;过评/视同过评产品剂型主要为注射剂。
截图来源:摩熵医药周报
本周盐酸尼卡地平注射液过评/视同过评受理号最多,为7个,同时该品种过评/视同过评企业最多,为4家。本周石家庄四药过评/视同过评品种数最多,为4种,本周过评/视同过评企业包括石家庄四药、珠海联邦制药和浙江仙琚制药等115家企业。
截图来源:摩熵医药周报
本周,布瑞哌唑片、利伐沙班干混悬剂、茶苯海明片、甲磺酸溴隐亭片4款品种首次过评/视同过评。
截图来源:摩熵医药周报
本周,别嘌醇片、吡格列酮二甲双胍片这两款品种,过评或视同过评的企业数量均达到7家。
截图来源:摩熵医药周报
摩熵咨询本期完整周报
识别二维码领取下载
END
本文为原创文章,转载请留言获取授权
近期热门资源获取
中国临床试验趋势与国际多中心临床展望-202505
2024年医药企业综合实力排行榜-202505
中国带状疱疹疫苗行业分析报告-202505
2023H2-2024H1中国药品分析报告-202504
数据透视:中药创新药、经典验方、改良型新药、同名同方的申报、获批、销售情况-202503
2024年中国1类新药靶点白皮书-202503
中国AI医疗健康企业创新发展百强榜单-202502
解码护肤抗衰:消费偏好洞察与市场格局分析-202502
2024年FDA批准上市的新药分析报告-202501
小分子化药白皮书(上)-202501
2024年中国医疗健康投融资全景洞察报告-202501
2024年医保谈判及市场分析报告-202501
近期更多摩熵咨询热门报告,识别下方二维码领取
联系我们,体验摩熵医药更多专业服务
会议
合作
园区
服务
数据库
咨询
定制
服务
媒体
合作
点击上方图片,即可开启摩熵化学数据查询
点击阅读原文,申请摩熵医药企业版免费试用!
2025-12-09
一、背景与流行病学概述
1. ICI 与神经免疫相关不良反应(n-irAEs)
全身irAEs发生率可达40–80%,其中皮肤、胃肠、内分泌最常见。
神经系统irAEs(n-irAEs)整体发生率约1–7%,重度(≥3 级)<2%,但在所有ICI相关致死事件中占比可达20%左右,与心肌炎并列“最危险的 irAE类型”。
大型系统综述显示:抗CTLA-4的n-irAE发生率约3.8%,抗PD-1/PD-L1 为6.1%,PD-1+CTLA-4联合可达12%。
神经毒性又可分为:中枢型:脑炎、脑膜炎、脱髓鞘病、垂体炎等(约17–25%);周围/神经肌接头/肌肉型:多发神经病、吉兰巴雷(GBS谱系)、重症肌无力样综合征、肌炎/肌病等(约 75–83%)。
颅神经毒性属于周围神经系统受累的一部分,是其中最“隐蔽但高致残”的亚型。
2. ICI 相关颅神经病变的流行病学
目前最系统的资料主要来自三类研究:
(1)Vogrig等2021年Neurology 全国队列+文献系统综述(39 例):
受累颅神经:面神经 (33%)、前庭蜗神经 (21%)、视神经 (18%)、展神经 (10%),三叉/动眼/舌咽等较少见。
约1/3患者治疗后仍有持久性神经功能缺损(多为听力或视力障碍)
(2)Pichon等2024年系统综述+个体病例数据分析(136 例):
面神经受累约38%,视神经35%,前庭蜗神经17%,动眼/展神经/三叉等占少数。
起病中位时间约10周(多数在 ICI 起始后数周–数月)。
MRI约40–50%可见颅神经或脑膜强化,CSF约60%有淋巴细胞轻中度增多/蛋白升高。
80%接受糖皮质激素治疗,约1/5叠加IVIG或血浆置换。完全恢复约1/3,部分恢复约1/2,约15–20%后遗,个别死亡。
(3)de Deus Vieira等2025年 J Neuroimmunol综述:
再次提示面神经与视神经是最常见受累颅神经,主要与nivolumab、pembrolizumab相关。
多数病例在启动ICI数周–数月内起病,常见于黑色素瘤和NSCLC。
联合或序贯多种ICI(如PD-1+CTLA-4)是明确的危险因素。
发生率:在所有ICI使用者中,颅神经毒性发生率大约为0.1–0.5%级别,其中眼科并发症(视神经炎、眼外肌肌炎等)约0.46%。
二、发病机制:从免疫检查点到颅神经炎
目前尚无统一机制,但可以归纳为几条主线:
1.T 细胞介导的自身免疫炎症
ICI阻断CTLA-4、PD-1/PD-L1后,激活CD8+ T细胞和NK细胞杀伤肿瘤,但也削弱了Treg抑制作用和外周耐受。
这类“去刹车”的免疫激活可对髓鞘、神经轴突、血管内皮等结构产生交叉攻击,形成颅神经炎、脱髓鞘性神经病或血管炎。
2.分子模拟与“肿瘤–神经”共享抗原
黑色素瘤与视网膜色素上皮等存在共表达抗原,激活针对肿瘤的免疫反应时,可能同时攻击这些组织(类似经典副肿瘤综合征)。
在ICI诱发的脱髓鞘多发性神经病中,已提示“黑色素–雪旺细胞”之间的抗原交叉;颅神经炎可能同源机制。
3.新生或加重的自身抗体介导过程
部分病例可检测到抗神经节苷脂抗体(如抗GQ1b)、抗AQP4/MOG、抗神经元或抗突触抗体(如NMDAR、LGI1)、甚至抗视神经抗体,提示B细胞及体液免疫参与。
ICI可诱发或加重原有自身免疫病,如重症肌无力、多发性硬化(MS)、视神经脊髓炎谱系疾病(NMOSD)、既往GBS等,颅神经毒性有时只是其表现之一。
4.靶分子在神经系统的本身表达
有研究显示,PD-1/PD-L1、CTLA-4 在CNS中也有RNA或蛋白表达,如脑皮层、基底节和垂体等;抗CTLA-4相关垂体炎被认为与垂体细胞表达CTLA-4 有关。
因此,ICI可能直接干扰这些区域的免疫稳态,引发垂体炎、脑膜炎,继而牵连多对颅神经(尤其 III、IV、VI、VII)。
5.微环境与宿主因素(“易感地基”)
高龄、既往自身免疫疾病、多药联合(特别是PD-1+CTLA-4)、既往神经放疗或神经毒化疗等均与n-irAE风险升高有关。
微生物群、遗传多态性(如CTLA-4 多态)可能影响总体irAE易感性,但在颅神经毒性方面数据尚有限。
简化记忆:
“解除刹车”→T/B 细胞过度活化+交叉抗原+本身PD1/CTLA4 神经表达 → 多灶免疫炎症(神经炎/血管炎/脑膜炎) → 颅神经功能障碍。
三、临床谱系:按颅神经逐条拆解
临床上,ICI相关颅神经毒性往往以“亚急性、单侧或多发颅神经功能障碍”起病,时间多在用药后2–24周,但也可更迟。
1. 视神经(CN II)——视神经炎/缺血性视神经病变
典型表现
双侧或单侧亚急性视力下降,视野缺损,可伴视盘水肿或视盘充血。
多数病例无明显自发眼痛或仅轻度眼球运动痛。
相关ICI:三大类均可见,pembrolizumab、nivolumab、ipilimumab 报道最多。
影像和辅助检查
MRI眶部T2高信号 + 视神经/视神经鞘强化,部分显著累及视神经头。
OCT(光学相干断层扫描)可见视网膜神经纤维层(RNFL)局灶或弥漫增厚或后期变薄。
特殊亚型:类巨细胞动脉炎样动脉炎性前缺血性视神经病变(AAION),可伴ESR/CRP 升高、血管壁炎性改变,需与原发GCA鉴别。
预后:早期停药+大剂量糖皮质激素大多可使视功能稳定,但约1/3留下不同程度视力损害。
2. 动眼/滑车/展神经(CN III/IV/VI)——眼肌麻痹
表现:复视、眼球运动受限、上睑下垂,可单神经受累,也可与视神经/三叉神经/交感纤维共同受累,呈“眶尖综合征”样。
影像:MRI见相应颅神经段或眶尖/海绵窦区域强化,需与转移瘤、侵袭性真菌感染、肉芽肿病等鉴别。
合并症:常伴脑膜炎、垂体炎或其他irAE(皮疹、肠炎等),提示系统性免疫激活。
3. 三叉神经(CN V)——面痛/面部感觉减退
表现:单侧面部麻木、痛觉过敏或神经痛样发作;部分呈“眶尖+三叉”多发颅神经综合征。
影像:Meckel腔、三叉神经根或周围支可见强化或肥厚,但也可完全正常。
鉴别:需排除带状疱疹、肿瘤浸润、颅底转移等。
4. 面神经(CN VII)——“免疫相关Bell麻痹”
是最常见的颅神经irAE之一(在cranial neuropathy队列中约占1/3–2/5)。
临床特征:
急性或亚急性单侧周围性面瘫,House-Brackmann II–VI级不等,可伴耳后疼痛、味觉减退、听觉过敏等。
可孤立存在,也可与前庭蜗神经炎、脑膜炎、垂体炎等并存。
影像:脑MRI常提示内听道/面神经管内强化(尤其膝部、迷路段),但亦可阴性。
预后:多数在停用ICI+1–2 mg/kg泼尼松等效剂量治疗后2–8周内明显好转;少数遗留轻度对称不全或联带运动。
5. 前庭蜗神经(CN VIII)——突发性听力下降/眩晕
发生率在颅神经病变中约15–20%。
表现:急性或亚急性耳鸣、感音神经性聋、旋转性眩晕、步态不稳。
辅助检查:
纯音测听、听性脑干反应测试(ABR)可提示高频感音性聋及脑干通路受累。
MRI内听道及迷路可见神经或迷路强化。
预后:相比面神经/眼肌,听力恢复率较差;Vogrig、Pichon 队列中,持续听力障碍是常见残疾类型。
6. 下组颅神经(CN IX/X/XI/XII)
较少见,但一旦发生临床往往凶险:
声嘶、吞咽困难、呛咳、软腭反射减弱、舌肌萎缩/纤颤等。
常合并脑干炎、脑膜炎或GBS谱系疾病。
任何新发的严重吞咽困难或呼吸受累均应视为3/4级毒性,立刻停药并评估ICU监护需求。
7. 多发颅神经病变与重叠综合征
表现为“多对颅神经+脑膜炎/脑神经根炎/GBS/肌炎/重症肌无力”等复杂重叠。
典型表现:
多发颅神经炎伴蛋白细胞分离、影像见脑神经根强化,类似GBS变异型(Miller–Fisher等)。
眼肌麻痹+近端无力+CK升高+呼吸肌受累,提示3M综合征。
四、诊断策略:实用评估路径
1. 何时高度怀疑ICI相关颅神经毒性?
在ICI治疗期间或停药半年内出现下列任何情况,应进入“高度怀疑”通道:
新发单侧或双侧面瘫
新发视力明显下降或复视
突发或亚急性听力下降/持续耳鸣
严重眩晕、步态不稳并伴眼震
吞咽困难、呛咳、鼻音、声嘶
颅神经症状伴其他irAEs(皮疹、肠炎、内分泌异常等)
颅神经症状与肿瘤影像学进展不匹配(如肿瘤稳定甚至部分缓解时出现)
2. 分级(基于CTCAEv5.0+神经专科共识)
可简化为:
1级:轻度症状,不影响日常生活活动(ADL)(如轻度面瘫但能完全闭眼、轻度耳鸣)
2级:中度症状,影响部分ADL(闭眼困难、复视影响阅读、轻–中度吞咽困难)
3级:重度症状,严重影响ADL(不能闭眼、吞咽明显困难、步态不能独行、视力 ≤0.1)
4级:危及生命(进行性呼吸衰竭、严重吸入性肺炎、双眼近盲)
临床要点:多数指南建议对任何新发颅神经功能缺损至少按“≥2 级 nirAE”处理,因为即便症状轻微,也可能是脑干炎或GBS的早期表现。
3. 标准化检查项目
(1)基础实验室
血常规、电解质、肝肾功能、炎症指标(ESR/CRP)
甲状腺功能、糖代谢、维生素B12/叶酸(排除代谢性原因)
(2)自身免疫与感染筛查(按流行病学情况加减)
抗核抗体谱、ENA、ANCA 等
抗AChR/MuSK、抗神经节苷脂抗体(GM1、GQ1b 等)
抗AQP4、抗MOG(视神经炎/脑脊髓病)
经典副肿瘤抗体(Hu、Yo、Ri、Ma2、CRMP5 等)
感染筛查:梅毒、HIV、乙丙肝、根据地区考虑结核、莱姆病等。
(3)影像学(非常关键)
脑及脑干MRI+颅神经序列(推荐含增强、薄层)
关注:视神经、眶尖、海绵窦、内听道、Meckel腔、颅底孔等。
典型表现:受累神经增粗、T2高信号、强化;或弥漫/局灶脑膜强化。
有视症状者:加做眶部 MRI、眼底/OCT
必要时全脊髓MRI(怀疑 GBS 谱系/根炎时看神经根强化)。
(4)脑脊液检查
几乎所有中重度n-irAEs都建议腰穿:
细胞数及分类、蛋白、糖、氯
细菌/真菌/结核培养+必要时病毒学(HSV/VZV/EBV 等核酸)
细胞学+流式细胞术(排除脑膜转移或淋巴瘤)
OCB、IgG 指数(脱髓鞘性疾病)
自身抗体(如 NMDAR、LGI1、CASPR2 等)
(5)电生理
神经传导/肌电(区分轴索vs脱髓鞘、合并周围神经或肌病)
诱发电位:blink reflex(三叉–面)、ABR(听神经)、VEP(视神经)等。
(6)生物标志物(前沿但可考虑)
血/CSF神经丝轻链(NfL)、胶质纤维酸性蛋白(GFAP):
Schmitt 等 2025 年研究指出,nirAE 患者的血NfL明显高于对照,水平与毒性分级、预后相关,且周围型与中枢型模式不同。
若有条件,可用于帮助区分nirAE与功能/心理性症状,评估预后。
4. 鉴别诊断要点
(1)肿瘤进展/转移/脑膜癌病
多灶颅神经受累+脑膜强化时尤需警惕:查CSF细胞学、多次重复。
(2)感染性脑膜炎/脑神经炎
特别是在长期激素或其他免疫抑制治疗后,需排除细菌、结核、隐球菌、VZV 等。
(3)经典自身免疫或副肿瘤综合征
如原发GCA、NMOSD、MOGAD、副肿瘤小脑变性等,可因ICI加重。
(4)放疗/化疗相关神经毒性
如累及颅底骨的放疗后迟发颅神经病、铂类/紫杉类相关多发周围神经病等。
(5)偶发疾病
必须结合时间关系:n-irAE常发生在ICI用药后6个月内,且常伴其它irAEs或肿瘤控制好转。
总体原则:排除其他一切可能原因之后,才能考虑ICI相关——但在危重/高疑似时,不要等所有结果回来才开始免疫治疗。
五、治疗与管理:按严重程度+受累部位分层
综合ASCO、ESMO、SITC指南及最新综述,总结出实用版治疗策略。
1. ICI处理原则
1级、无进展趋势:多数指南允许在密切监测下继续ICI,但对于新发颅神经受累,建议至少暂缓1个周期,排除进展/脑膜炎后再评估。
≥2级:一般建议立即暂停ICI,启动免疫抑制治疗。
3/4级或危及重要功能(例如视神经炎、重度吞咽困难):建议永久停用当前ICI,除非在多学科评估下有非常强的再挑战指征。
2. 糖皮质激素:核心基石
推荐剂量(可按病情微调):
2级颅神经毒性:建议口服泼尼松0.5–1mg/kg/日(或等效),症状稳定3–5天后减量,总疗程≥4–6 周,防止复燃。
3/4级或视神经炎、多发颅神经炎、伴脑膜炎/GBS/MG/肌炎的重叠综合征:甲泼尼龙静脉冲击500mg/日×3–5d,然后改为口服1–2mg/kg/日,至少6–8周减量。
注意事项:长疗程需联合PPI、预防骨质疏松及PJP感染(如SMZ-TMP),并监测血糖、血压、电解质等。
临床经验提示:
与经典GBS不同,ICI相关神经根炎/GBS对糖皮质激素通常被纳入一线治疗,因为其机制更偏炎性神经根炎而非纯典型GBS。
3. IVIG / 血浆置换(PLEX)
适应证:
3/4级神经毒性;
合并GBS/MFS谱系、重症肌无力危象、严重肌炎、生命体征不稳;
高剂量激素3–5天内无明显好转。
方案:
IVIG:2g/kg 总量,3–5天内完成;
PLEX:隔日1次,共5–7次,可与激素联用。
4. 二线免疫抑制剂
目前多基于个案或小系列证据,主要针对激素+IVIG/PLEX 仍难控制的重症:
利妥昔单抗(尤其合并B细胞或抗体介导过程)
环磷酰胺
吗替麦考酚酯 / 硫唑嘌呤(维持期)
针对其它irAEs已有较多经验的生物制剂,如:抗TNF-α(infliximab)主要用于结肠炎;抗IL-6(tocilizumab)主要用于关节炎,但也有个案用于顽固n-irAE。
目前尚无针对颅神经毒性的标准二线方案,建议在神经免疫+肿瘤+风湿多学科讨论后个体化选择。
5. 器官/症状针对治疗
(1)视神经炎 / 视神经缺血
视神经炎处理原则基本等同于MS相关视神经炎:
立即停用ICI;
甲泼尼龙1g/日×3–5d静滴;
视功能无改善或进展者考虑PLEX(5 次)。
对动脉炎性前部缺血性视神经病变(AAION)样表现,需按巨细胞动脉炎流程加做颞动脉活检/PET-CT,并尽早大剂量激素甚至加用托珠单抗。
(2)面神经麻痹
绝大多数可按“免疫相关面神经炎”管理:
口服泼尼松1mg/kg/日×7–10 d,然后4–6周缓减。
眼部保护:人工泪液、夜间眼罩/胶布、必要时眼睑缝合术。
面肌康复训练±物理治疗。
一般无需常规抗病毒药物,除非有带状疱疹感染证据。
(3)前庭蜗神经受累
大剂量激素为主,辅以短期前庭抑制剂(如氯丙嗪、茶苯海明等)和康复训练。
早期耳科/听力学介入,有助于明确基线并随访。
(4)下组颅神经/多发颅神经炎
极易导致误吸和窒息,需早期评估吞咽功能:
床旁吞咽试验→视频透视吞咽造影/纤维鼻内镜吞咽检查。
必要时暂时鼻饲或胃造瘘;密切监测肺部感染。
合并GBS/MG/肌炎的重叠综合征,治疗上以激素+IVIG/PLEX早期联合为宜。
六、ICI 再挑战(Rechallenge)
关于n-irAE 后是否再启ICI,目前数据有限、争议较大。
1. 现有证据
Pichon136例病历中,少部分患者在颅神经症状控制后再次接受 ICI,多数未出现同型颅神经复发,但有相当比例出现新的irAEs。
其他n-irAE队列显示:对于1/2级、完全缓解的不良事件,再挑战的irAE复发/新发率约20–40%;对于3/4 级nirAE(尤其脑炎、GBS、重症肌无力、严重视神经炎),绝大多数指南不推荐再挑战,除非无替代治疗且患者充分知情。
2. 实用决策框架
可以考虑再挑战的情形:
颅神经毒性为2级以下;
经治疗后完全临床缓解且停用激素≥4–8 周;
无持久损伤,如视力基本恢复、面瘫完全恢复等;
肿瘤学方面存在强烈用药指征(无其他有效方案或ICI获得显著持久缓解);
多学科(肿瘤、神经、风湿、眼科/耳鼻喉等)共同评估,患者充分知情并签署详细告知同意。
再挑战策略建议:
优先选择毒性较温和的PD-1/PD-L1单药,避免再次给予含CTLA-4 的联合方案。
初始阶段缩短随访间隔,设立明确“停止阈值”(一旦出现任何新的颅神经症状即立刻停药+激素)。
必要时可在再挑战前后监测NfL/GFAP等生物标志物,作为辅助监控工具。
七、随访与预防
1. 基线评估与患者宣教
起始ICI前建议记录:
详细神经系统及颅神经查体(面肌力量、眼外肌、咽反射、舌运动、听力等);
有既往视神经炎/多发性硬化、听力损失或重症肌无力等者需请神经/眼科/耳科联合评估。
把以下症状写入患者的告知单:
一侧脸“歪”、“无法闭眼”;
任何新发视力明显下降、视野缺损或持续复视;
突然听不清、持续耳鸣或旋转性眩晕;
吃饭呛咳、喝水呛、说话含糊或鼻音;
以上任何一条均需要在24小时内联系医生。
2. 随访与功能评估
前几个月(尤其是前3–6 个月)建议每次输注前进行简短的颅神经筛查(面肌、眼外肌运动、视力主观问诊、听力主观问诊、咽反射)。
对已发生n-irAE 的患者:
视神经受累:每1–3个月复查视功能、OCT±视野;
听神经受累:听力学随访;
多发颅神经:神经功能量表(mRS、Barthel)及生活质量评估。
八、前沿进展:从“经验医学”走向“精准神经免疫毒性学”
1. 统一的n-rAE疾病定义与分型
2021年起,国际专家组提出了n-irAE 共识疾病定义,对脑炎、脑膜炎、周围神经病、神经肌接头病、肌炎等进行了标准化分类与诊断标准,为临床试验和队列研究提供了统一框架。
最近对颅神经和眼科irAEs也制定了专门的共识定义,强调眼科及影像学证据的重要性。
2. 生物标志物
可溶性标志物:Farina 2024 mini-review综合了ICI-脑炎中多种可溶性生物标志物(细胞因子、可溶性免疫检查点、神经抗体等),提示其在诊断、预后评估中的潜在价值。
神经丝轻链(NfL)与GFAP:多项研究显示,在n-irAE中血/CSF NfL和GFAP水平升高,与病变部位(中枢vs周围)、严重程度以及预后相关,甚至可能在临床症状出现前就升高。
预测irAE风险的全身性指标:多组学分析(基因、转录组、微生物组)已发现多种与irAE风险相关的候选标志物,如LCP1+ADPGK 模型、外周炎症指数(NLR、SII)、循环肿瘤细胞负荷等;但在颅神经毒性方面尚缺特异数据。
3. 治疗策略的优化
大量回顾性研究正在比较早期高剂量激素vs逐级递进对肿瘤疗效与n-irAE预后的影响,有研究提示“过早、过量的激素可能削弱ICI抗肿瘤效应”,尤其是在作为一线治疗的黑色素瘤中。
新型免疫调节药物(如JAK抑制剂、选择性IL-6/IL-17抑制剂等)在部分顽固irAE中显示出前景,但在颅神经毒性中的经验目前仅限零星病例。
九、临床要点
1.任何ICI用药后6个月内的新发颅神经功能障碍,都必须主动问:会不会是n-irAE?
2.面神经和视神经是最常见受累部位,其次为前庭蜗神经;一旦出现视力或听力损害,预后较差,应当“抢时间”。
3.诊断一定要“先排除肿瘤进展和感染”,但不要为此延误激素和IVIG/PLEX 的启动。
4.一旦判断为≥2级颅神经n-irAE:
立即停用 ICI;
立即开始0.5–2mg/kg 等效泼尼松治疗(视严重程度);
标准化行MRI+CSF+电生理+自身抗体检查。
5. 3/4 级、累及视神经或下组颅神经者,一律按抢救级别处理:IV甲泼尼龙+IVIG/PLEX+ICU/吞咽管理。
6.绝大多数患者在适当免疫治疗后可显著改善,但约1/3将留下持久缺损,要提前与患者沟通预后。
7.再挑战ICI要慎之又慎:轻度完全缓解病例可在多学科评估后考虑换用 PD-1单药,严重类型一般不建议。
临床结果临床研究AHA会议
100 项与 茶苯海明 相关的药物交易
登录后查看更多信息
研发状态
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| 头晕 | 日本 | 1952-01-01 | |
| 恶心和呕吐 | 日本 | 1952-01-01 | |
| 术后恶心呕吐 | 日本 | 1952-01-01 | |
| 湿疹 | 日本 | 1951-09-11 | |
| 昆虫咬伤和螫伤 | 日本 | 1951-09-11 | |
| 瘙痒 | 日本 | 1951-09-11 | |
| 荨麻疹 | 日本 | 1951-09-11 | |
| 晕动病 | 中国 | - |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
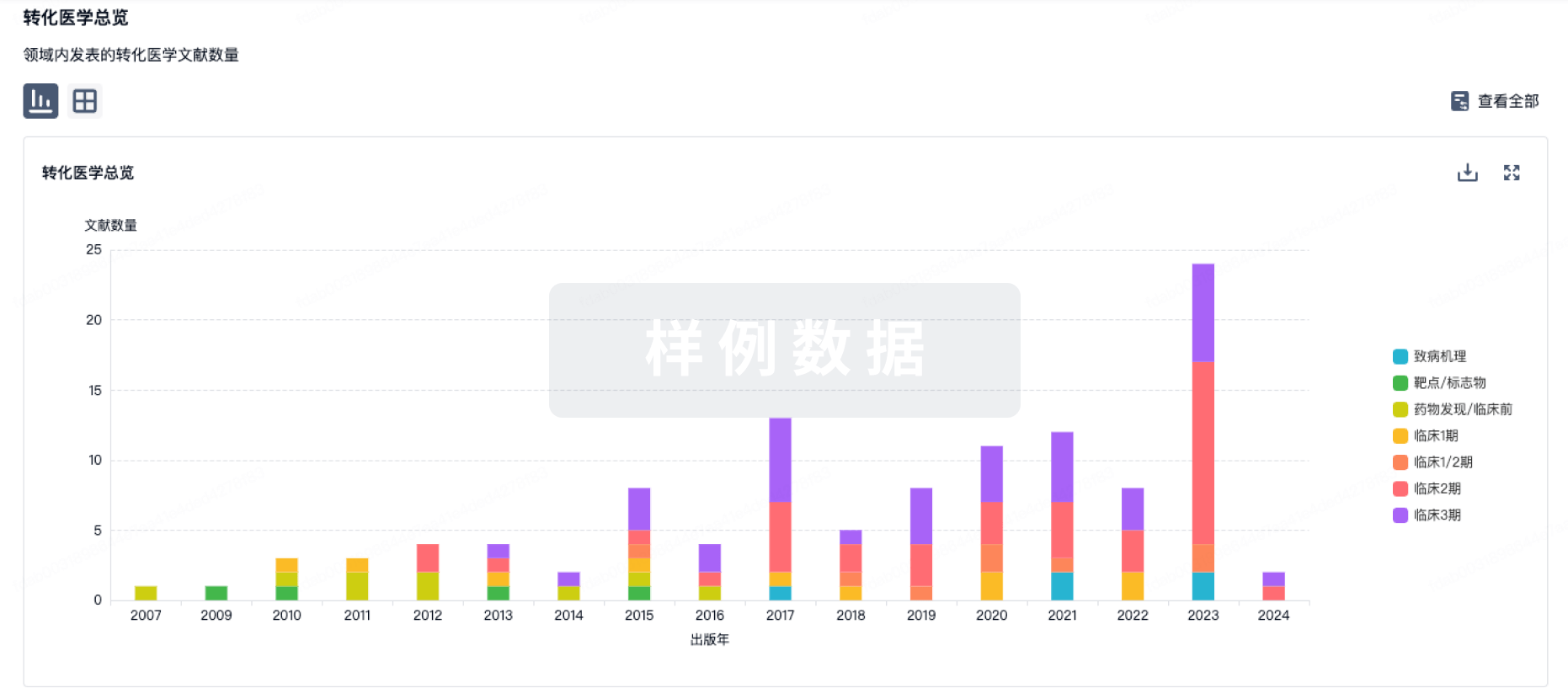
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
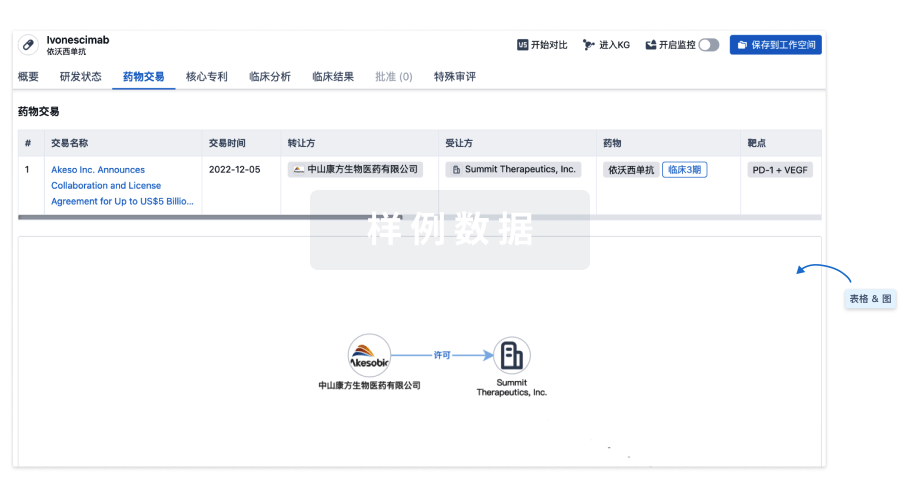
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
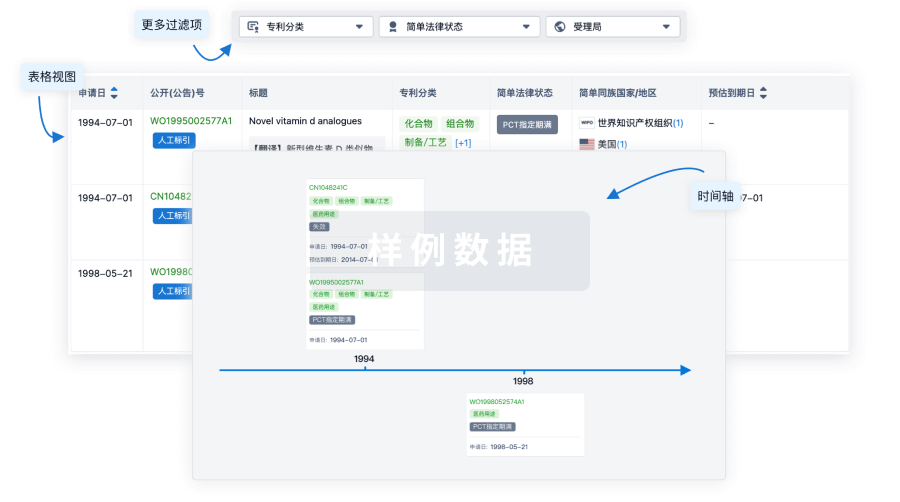
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
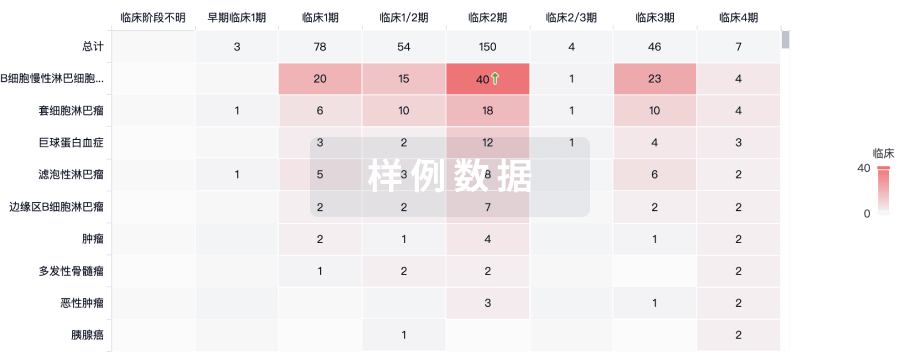
批准
利用最新的监管批准信息加速您的研究。
登录
或
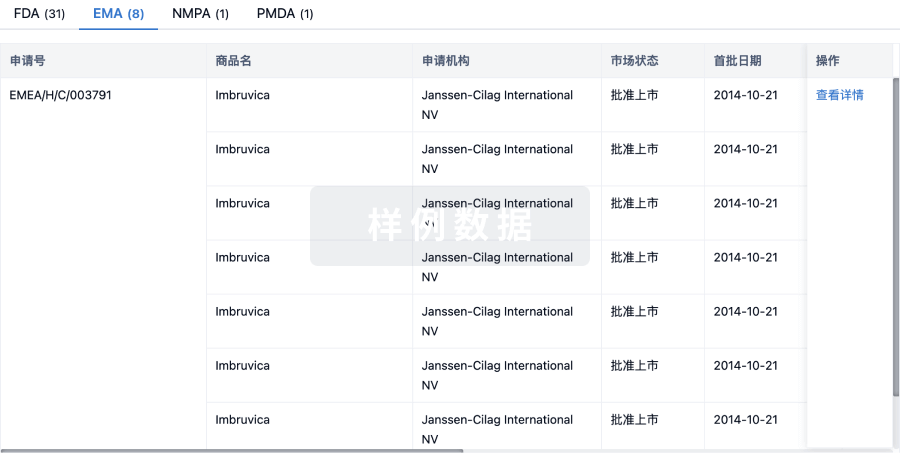
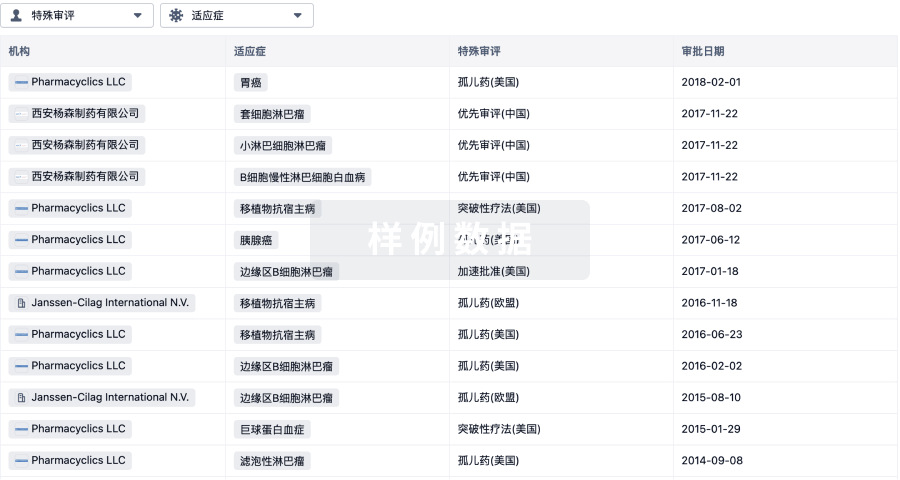
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用