预约演示
更新于:2025-08-30
FP-0023
更新于:2025-08-30
概要
基本信息
在研机构- |
权益机构- |
最高研发阶段无进展临床前 |
首次获批日期- |
最高研发阶段(中国)- |
特殊审评- |
关联
100 项与 FP-0023 相关的临床结果
登录后查看更多信息
100 项与 FP-0023 相关的转化医学
登录后查看更多信息
100 项与 FP-0023 相关的专利(医药)
登录后查看更多信息
9
项与 FP-0023 相关的文献(医药)2012-08-01·Biophysical chemistry4区 · 生物学
Molecular dynamics simulation of HIV-1 fusion domain-membrane complexes: Insight into the N-terminal gp41 fusion mechanism
4区 · 生物学
Article
作者: G. Matthias Ullmann ; Siriporn Promsri ; Supot Hannongbua
To understand how viral proteins fuse with the cell membrane, a process that is important for the initial stages of infection, the conformation of the membrane-bound viral fusion peptides (FPs) must be identified. Here, molecular dynamics (MD) simulations were performed to investigate the conformation of the FP-16 and FP-23 of human immunodeficiency virus (HIV) bound to a dimyristoyl phosphatidylcholine (DMPC) bilayer. All FPs were found to penetrate into the bilayer despite the different initial orientations. In addition, the inclusion of residues 17 to 23 was found to play significant role in the fusion ability. Each of the FPs adopts at least two distinct conformations (predominantly α-helical and β-structures) while association with membrane. The peptide seriously affects structural and dynamical parameter of the contacted lipids. The previously experimental data together with our simulation data reveal that fusion ability depends on the membrane-associated conformation and alignment of the peptide.
2011-04-01·European biophysics journal : EBJ4区 · 生物学
Irregular structure of the HIV fusion peptide in membranes demonstrated by solid-state NMR and MD simulations
4区 · 生物学
Article
作者: Erik Strandberg ; Dorit Grasnick ; Parvesh Wadhwani ; Ulrich Sternberg ; Anne S. Ulrich
To better understand peptide-induced membrane fusion at a molecular level, we set out to determine the structure of the fusogenic peptide FP23 from the HIV-1 protein gp41 when bound to a lipid bilayer. An established solid-state (19)F nuclear magnetic resonance (NMR) approach was used to collect local orientational constraints from a series of CF(3)-phenylglycine-labeled peptide analogues in macroscopically aligned membranes. Fusion assays showed that these (19)F-labels did not significantly affect peptide function. The NMR spectra were characteristic of well-behaved samples, without any signs of heterogeneity or peptide aggregation at 1:300 in 1,2-dimyristoyl-sn-glycero-3-phosphatidylcholine (DMPC). We can conclude from these NMR data that FP23 has a well-defined (time-averaged) conformation and undergoes lateral diffusion in the bilayer plane, presumably as a monomer or small oligomer. Attempts to evaluate its conformation in terms of various secondary structures, however, showed that FP23 does not form any type of regular helix or β-strand. Therefore, all-atom molecular dynamics (MD) simulations were carried out using the orientational NMR constraints as pseudo-forces to drive the peptide into a stable alignment and structure. The resulting picture suggests that FP23 can adopt multiple β-turns and insert obliquely into the membrane. Such irregular conformation explains why the structure of the fusion peptide could not be reliably determined by any biophysical method so far.
2010-11-01·Microbes and infection3区 · 医学
The encapsulated strain TIGR4 of Streptococcus pneumoniae is phagocytosed but is resistant to intracellular killing by mouse microglia
3区 · 医学
Article
作者: Sergio Tripodi ; Bruna Colombari ; Alessio Zanardi ; Marco Rinaldo Oggioni ; Giuliana Fabio ; Elena Righi ; Samuele Peppoloni ; Marcella Cintorino ; Michele Zoli ; Damiana Chiavolini ; Velia Braione ; Carlotta F. Orsi ; Elisabetta Blasi ; Susanna Ricci ; Gianni Pozzi ; Massimino Messinò ; Maria Margherita De Santi
The polysaccharide capsule is a major virulence factor of Streptococcus pneumoniae as it confers resistance to phagocytosis. The encapsulated serotype 4 TIGR4 strain was shown to be efficiently phagocytosed by the mouse microglial cell line BV2, whereas the type 3 HB565 strain resisted phagocytosis. Comparing survival after uptake of TIGR4 or its unencapsulated derivative FP23 in gentamicin protection and phagolysosome maturation assays, it was shown that TIGR4 was protected from intracellular killing. Pneumococcal capsular genes were up-regulated in intracellular TIGR4 bacteria recovered from microglial cells. Actual presence of bacteria inside BV2 cells was confirmed by transmission electron microscopy (TEM) for both TIGR4 and FP23 strains, but typical phagosomes/phagolysosomes were detected only in cells infected with the unencapsulated strain. In a mouse model of meningitis based on intracranic inoculation of pneumococci, TIGR4 caused lethal meningitis with an LD(50) of 2 × 10² CFU, whereas the LD(50) for the unencapsulated FP23 was greater than 10⁷ CFU. Phagocytosis of TIGR4 by microglia was also demonstrated by TEM and immunohistochemistry on brain samples from infected mice. The results indicate that encapsulation does not protect the TIGR4 strain from phagocytosis by microglia, while it affords resistance to intracellular killing.
100 项与 FP-0023 相关的药物交易
登录后查看更多信息
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 肌营养不良 | 临床前 | 法国 | - |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
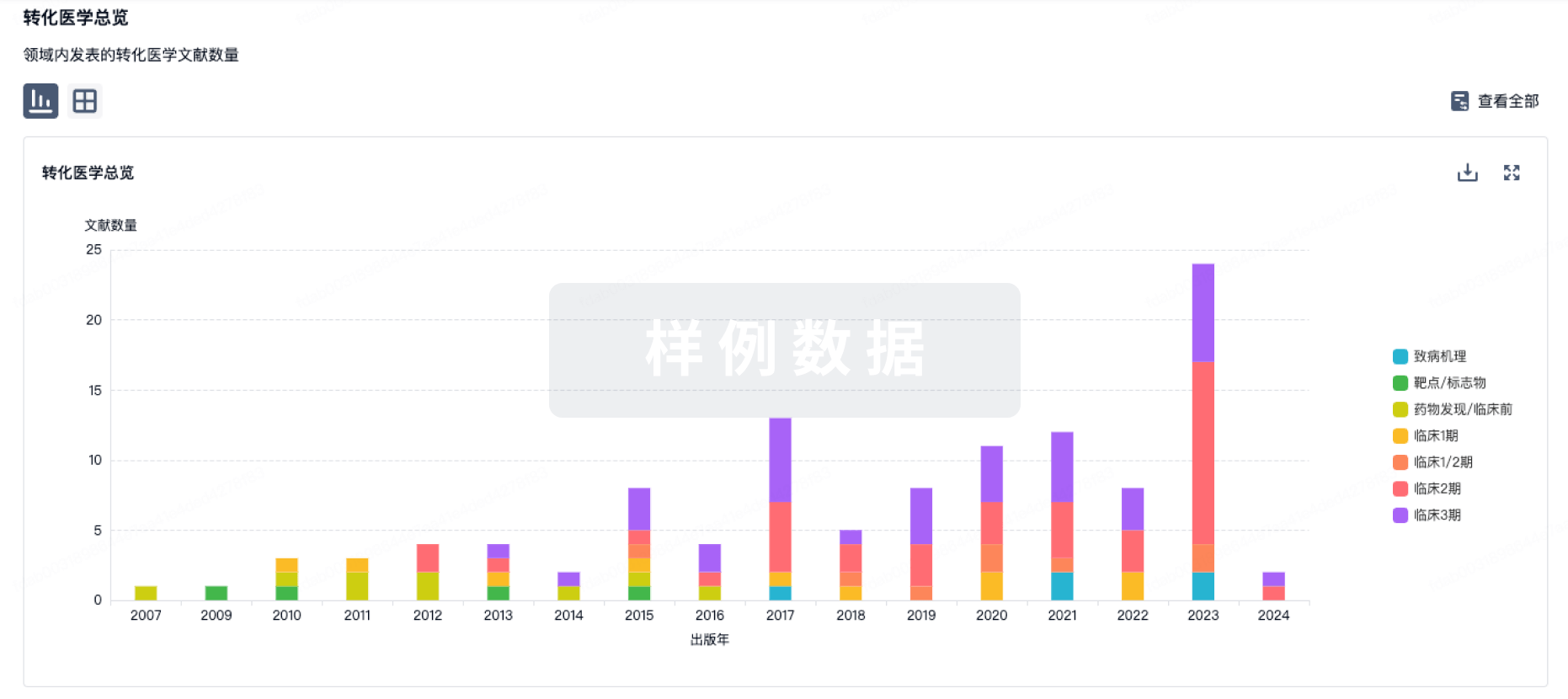
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
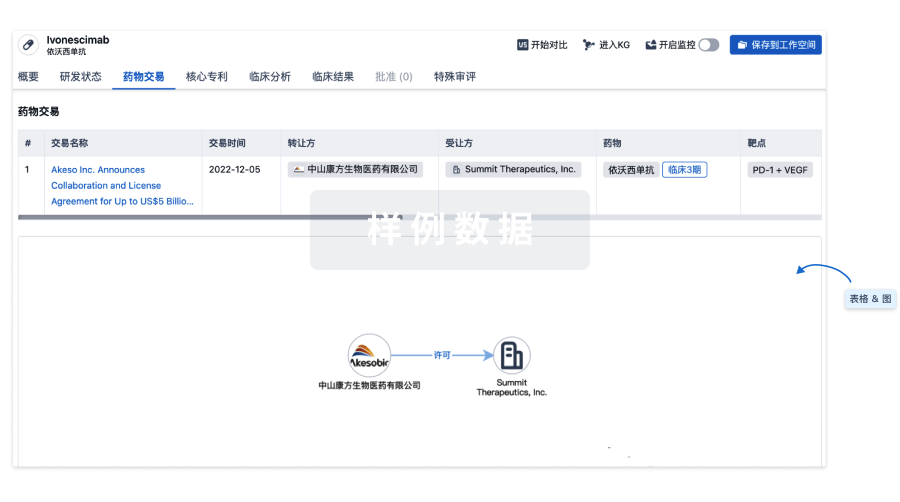
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
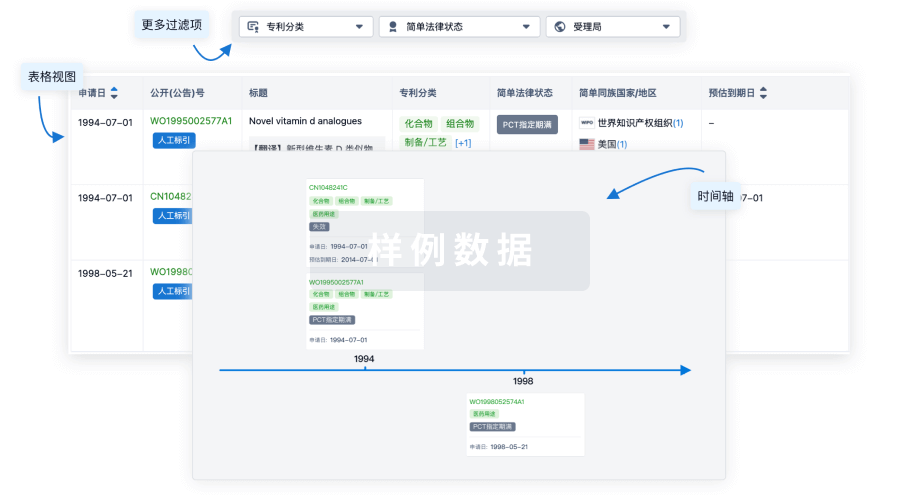
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
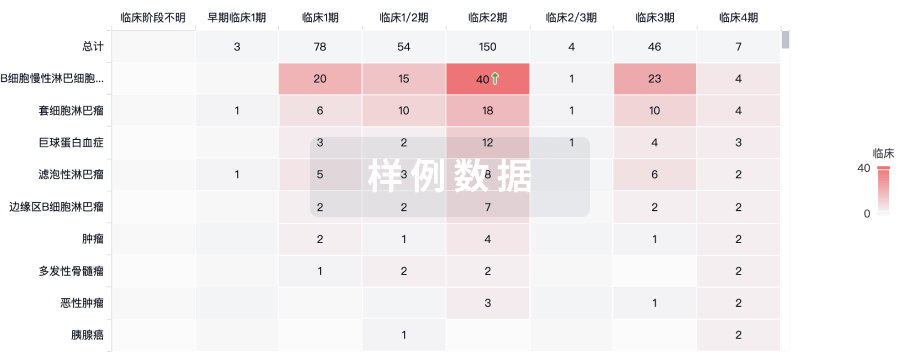
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
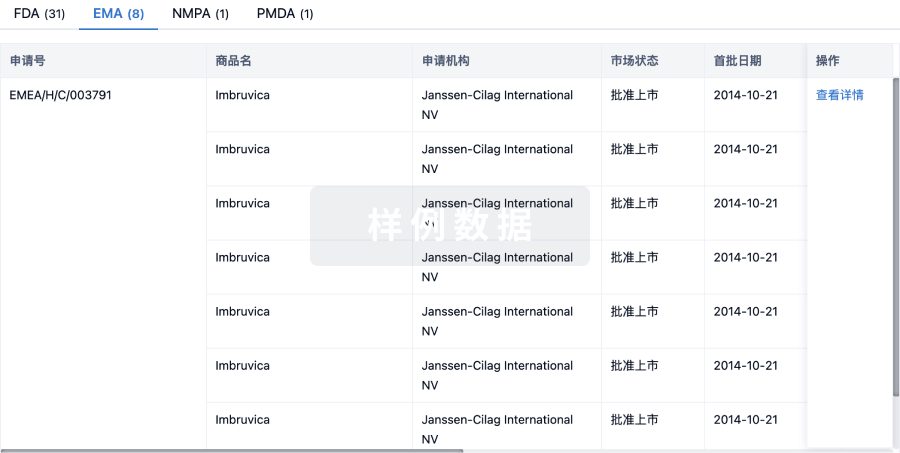
批准
利用最新的监管批准信息加速您的研究。
登录
或
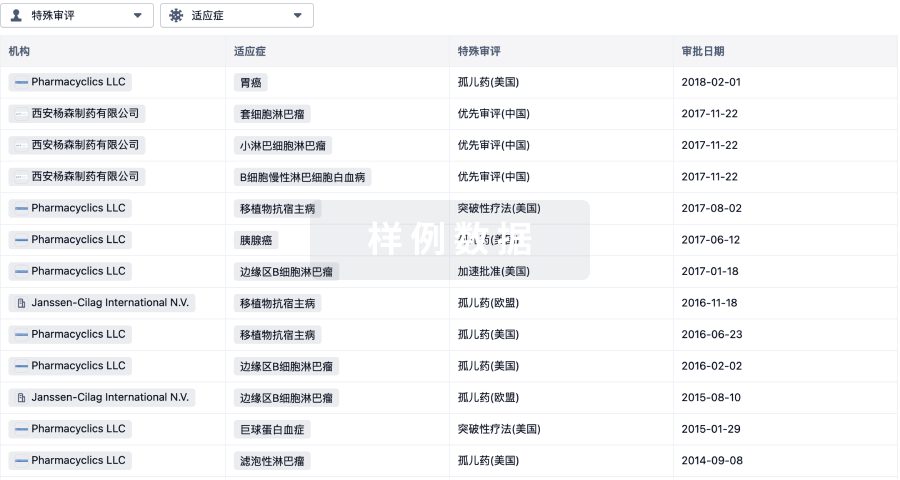
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
