预约演示
更新于:2026-03-17
α-Hederin
α-常春藤皂苷
更新于:2026-03-17
概要
基本信息
原研机构- |
非在研机构- |
权益机构- |
最高研发阶段批准上市 |
首次获批日期- |
最高研发阶段(中国)批准上市 |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C41H66O12 |
InChIKeyKEOITPILCOILGM-LLJOFIFVSA-N |
CAS号27013-91-8 |
关联
100 项与 α-常春藤皂苷 相关的临床结果
登录后查看更多信息
100 项与 α-常春藤皂苷 相关的转化医学
登录后查看更多信息
100 项与 α-常春藤皂苷 相关的专利(医药)
登录后查看更多信息
392
项与 α-常春藤皂苷 相关的文献(医药)2026-02-01·JOURNAL OF CHEMICAL ECOLOGY
Structure-activity Relationships of Triterpenoid Saponins Across Phylogenetically Diverse Organisms
Article
作者: Bak, Søren ; Cedergreen, Nina ; Schmitt, Franz Marius ; Günther, Jan ; Dervishi, Malbor
Saponins are structurally diverse bioactive metabolites found in more than 100 plant families and are synthesized by plants as protection against insects, fungi, and other organisms. The mode of action is related to their interference with membranes, and particularly the membrane sterols. As membrane sterol composition varies across kingdoms and species, species-specific differences in toxicity may therefore be expected. The aim of this study was to elucidate the structure-activity relationships of different saponins across four different organisms (Daphnia magna, Enchytraeus crypticus, Saccharomyces cerevisiae, Raphidocelis subcapitata) representing three eukaryotic kingdoms of life using either immobility tests or growth inhibition assays. We hypothesized that monodesmosidic saponins are more bioactive due to higher amphiphilicity/polarity and that species susceptibility depends on sterol composition, with organisms containing plant sterols being less susceptible than those with animal or fungal sterols. The hypothesis was supported for monodesmosidic saponins, as α-hederin and hederacolchiside A1 exhibited significant cytotoxicity (EC50 values ranging from 8.7 to 36.9 and 2.0- 68.9 mg/L, respectively, for the different organisms), whereas bidesmosidic saponins such as hederacoside C and ginsenoside-Ro were inactive at concentrations up to 100 mg/L. The aglycone backbone and sugar moiety composition, however, also play critical roles, with simpler, linear saccharide chains leading to increased toxicity. C-23 hydroxylation has been shown to enhance mortality against insects; however, its absence did not affect the ability of hederacolchiside A1 to exhibit toxic properties. Additionally, species-specific sensitivities varied, with the crustacean D. magna being the most sensitive species, followed by the anelid worm E. crypticus, yeast S. cerevisiae, and the least-sensitive was, as hypothesized, the algae R. subcapitata. These insights contribute to a deeper understanding of saponin structure-activity relationships and open new avenues for the targeted development of saponin-based applications in agriculture, medicine, and biotechnology.
2026-01-01·Journal of Pharmaceutical Analysis
Comment on “α-hederin decreases the glycolysis level in intestinal epithelial cells via SNX10-mediated DEPDC5 degradation”
Article
作者: Hao, Haiping
2025-12-01·Natural Products and Bioprospecting
Therapeutic targeting of ocular diseases with emphasis on PI3K/Akt, and OPRL pathways by Hedera helix L. saponins: a new approach for the treatment of Pseudomonas aeruginosa-induced bacterial keratitis
Article
作者: El-Shiekh, Riham A ; Hatem, Shymaa ; Elshimy, Rana ; Osman, Ahmed H ; Elosaily, Heba ; Hassan, Abrar Gomaa Abd-Elfattah ; Hamdy, Sherif A
Abstract:
Pseudomonas aeruginosa-induced bacterial keratitis is one of the most sight-threatening corneal infections associated with intense ocular inflammatory reactions that may lead to vision loss. Hence, this study investigated the efficacy of three nanocomposite chitosan-coated penetration enhancer vesicles (PEVs) to augment the ocular delivery of saponin(s), α-hederin (PEVI), hederacoside C (PEVII), or both (PEVIII) for treatment of Pseudomonas keratitis and its induced inflammatory response. The three formulations were prepared using the ethanol injection method and comprehensively characterized. In vitro, the antibacterial activity of the three formulations against P. aeruginosa was evaluated using agar well-diffusion method, pyocyanin production inhibition, and swarming and twitching motility inhibition assays. The therapeutic effect of the three formulations has been investigated in P. aeruginosa keratitis by gross lesion monitoring, determination of bacterial bioburden, biochemical markers, histopathological examination, and scoring after 7 days of topical treatment. Data revealed that PEVI, PEVII, and PEVIII nanocomposites showed particle size in the nanometer range, high entrapment efficiency, good stability, and sustained release of the saponins throughout 24 h. Among them, PEVIII exhibited notably strong in vitro antipseudomonal activity. Additionally, animals treated topically with PEVIII showed an appreciable gross lesion reduction, corneal tissue improvement, and formidable bacterial load reduction compared with untreated and gentamicin sulfate eye (GENTAWISE®) ointment-treated groups. Moreover, PEVIII treatment showed the most significant reduction in TNF-α, NF-κB, ROS levels, and OPRL virulence gene expression while enhancing PI3K/Akt activation. Therefore, this study offers PEVIII as a promising treatment for P. aeruginosa keratitis.Graphical Abstract
6
项与 α-常春藤皂苷 相关的新闻(医药)2026-03-02
基于计算机虚拟筛选发现α-常春藤皂苷通过靶向CYP51发挥抗白色念珠菌活性
01
研究背景
由念珠菌属和曲霉属引起的侵袭性真菌感染对人类健康构成严重威胁。在全球范围内,每年约有1.5亿严重真菌感染病例及150万相关死亡病例,其中念珠菌病是主要致病菌之一。其致病性依赖于多种毒性因子,包括黏附宿主细胞、分泌水解酶、从酵母态向菌丝态转换以及形成生物膜。因此,抑制其生长和毒性是关键的治疗策略。
源自植物和草药的天然小分子化合物,提供了结构多样且低毒性的替代方案。传统天然化合物(例如carvacrol 和geraniol)通过破坏细胞膜或抑制生物膜形成等毒性因子有效抑制白色念珠菌。但传统筛选方法效率低且机制不明确。计算机辅助的虚拟筛选技术通过同源建模、分子对接和分子动力学模拟等方法,能系统性地靶向多条通路,高效、机制明确地发现活性化合物。
当前虚拟筛选主要分为基于靶点的分子对接和基于配体的药效团建模两大策略。例如,有研究通过为角鲨烯环氧化酶和CYP51建立药效团模型,成功设计了新型抑制剂。真菌麦角固醇合成途径中的关键酶CYP51,因其与人类同源蛋白差异大,是理想的抗真菌靶点。多项研究通过设计能与CYP51活性位点残基(如T763、Y145等)结合的吲哚类衍生物,有效抑制了麦角固醇合成与白色念珠菌生长。
近年来,计算机辅助药物发现(CADD)技术显著推动了药物研发的进程。虚拟筛选、深度学习等方法能够快速识别候选分子,为抗菌药物研发带来了革新。然而,现有CADD方法往往受限于分子内能量计算不准、分子对接假阳性率高等问题,影响其实际应用。为克服这些瓶颈,西北工业大学生命科学学院蒋春美教授团队提出一种创新的双靶点筛选策略,通过整合蛋白质结构信息与配体性质参数并对两类筛选结果进行交叉验证,将候选化合物从1000余个缩减至25个高潜力分子,大幅提升了筛选效率与准确性。该策略特别适用于靶点不明确的抗真菌药物发现。研究团队将其应用于靶向CYP51(氟康唑的作用靶点)的天然小分子筛选,成功鉴定出α-常春藤皂苷与榄香素。两者作用机制与氟康唑类似,但抗真菌活性更优,展现出良好的开发前景。
02
靶向CYP51的天然抗真菌
小分子虚拟筛选
利用AutoDock工具对RCSB数据库中下载的白色念珠菌CYP51蛋白的晶体结构数据进行结构优化和能量最小化处理。根据文献数据及活性位点分析,选取Tyr118、His310和Ser378这三个关键残基在来界定配体口袋。最终在位于凸起上方的β5区域进行构建。(图1)
图1 CYP51晶体结构(A);5TZ1的最低能量构象(B);白色念珠菌与人类CYP51 β5序列的比较(C)
首先,基于口服生物利用度(OB>30%)和类药性(DL>18%)指标对TCMSP数据库进行初步筛选;随后,利用Discovery Studio中的LibDock评分函数预测配体构象,并通过cdocker模块进行结合优化与能量最小化。最后,采用AutoDock Vina计算结合自由能,以评估蛋白-配体相互作用的强度。通过将分子对接筛选获得的25个化合物与基于Discovery Studio药效团模型筛选的57个化合物进行交叉比对,优先保留了那些可以与CYP51活性位点形成强氢键作用的分子。两种方法共同筛选出17个候选化合物(图2),并进一步通过实验评估了它们对白色念珠菌的抑菌活性。
图2 CYP51虚拟筛选的工作流程
最终筛选出两种天然小分子—α-常春藤皂苷(α-hederin)和榄香素(elemicin),两类小分子在浓度低于128μg/ml时即表现出显著的抗真菌活性(表1)。
表1 抗真菌候选分子
03
α-常春藤皂苷对白色念珠菌
具有良好的抗菌活性
该团队对α-常春藤皂苷、榄香素和氟康唑对白色念珠菌的最低抑菌浓度(MIC)进行测试(图3A-C)。结果显示α-常春藤皂苷、榄香素均表现出强效抗真菌活性,其MIC值分别为32 μg/mL和16 μg/mL。尤其当浓度超过0.5 μg/mL时,榄香素可抑制50%-70%的白色念珠菌生长;α-常春藤皂苷在8 μg/mL浓度下24h内表现出对白色念珠菌50%的生长抑制。相比之下氟康唑的抗真菌活性明显低于α-常春藤皂苷和榄香素,即便是在24h后128μg/mL浓度下白色念珠菌仍持续性的生长。
为评估α-常春藤皂苷、榄香素和氟康唑的细胞毒性,将293T、Raw264.7和KB细胞分别与不同浓度的这三种化合物共培养24小时。细胞存活率结果如图3D-F所示。α-常春藤皂苷和氟康唑均表现出较低的细胞毒性,即使在最高测试浓度(128 μg/mL)下,三种细胞系的存活率仍保持在80%以上。相比之下,榄香素在浓度高于8 μg/mL时对293T、Raw264.7和KB细胞显示出显著毒性(图3E)。测定结果显示,α-常春藤皂苷的CC90为93.15 μg/mL,氟康唑为60.65 μg/mL,艾米霉素为0.72 μg/mL。基于其良好的用药安全性,α-常春藤皂苷被选中用于后续抗菌机制的研究。
图3 α-常春藤皂苷(A)、榄香素(B)和氟康唑(C)对白色念珠菌的最小抑制浓度,以及它们对Raw264.7(D)、KB(E)和293T细胞(F)的细胞毒性。
04
α-常春藤皂苷对白色念珠菌
细胞微结构的影响
经扫描电子显微镜(SEM)与透射电子显微镜(TEM)观察,α-常春藤皂苷处理后的白色念珠菌细胞表面出现明显变形、凹陷及粗糙(图4A)。相比之下,氟康唑处理组细胞表面则保持完整光滑,内部无明显异常。TEM进一步显示,对照组细胞具有结构清晰的细胞器、完整的核膜、致密的细胞质以及光滑的细胞壁/膜。而经32 μg/mL α-常春藤皂苷处理后,细胞内部发生剧烈改变,表现为胞质内容物严重丢失、大量空泡形成、细胞壁/膜变薄并伴有膜结构断裂、表面凸起以及出现近似中空的细胞区域。以上结果表明,α-常春藤皂苷可通过破坏细胞膜的完整性,引起内容物泄漏,从而发挥抗真菌作用,且其对细胞结构的破坏程度显著高于氟康唑。
图4 α-常春藤皂苷对白色念珠菌微形态结构、生物膜形成及成熟生物膜的影响
05
α-常春藤皂苷对白色念珠菌
生物膜形成及成熟生物膜的影响
生物膜的形成是白色念珠菌的重要致病机制。本文采用 XTT 法检测细胞代谢活性,通过结晶紫染色法测定生物膜总生物量,进而探究α-常春藤皂苷对生物膜形成及成熟生物膜的抑制作用。α-常春藤皂苷以浓度依赖性的方式抑制白色念珠菌早期生物膜的形成,并在32μg/mL和256μg/mL浓度下分别具有55%和75%的抑制率。相比氟康唑的抑制效果显著较弱,即便是在256μg/mL的浓度下也仅显示了75%的抑制率。以此表明,α-常春藤皂苷通过抑制真菌黏附阻止生物膜形成的效果更加明显(图4B-F)。然而,以上两种化合物均未能有效清除成熟生物膜,在128μg/mL的浓度下α-常春藤皂苷和氟康唑的抑制率分别仅为20%和17%(图4G-H)。综合以上数据表明,α-常春藤皂苷在生物膜形成的早期阶段发挥了较强的抑制生长的作用。
06
α-常春藤皂苷对白色念珠菌
菌丝形成、细胞表面疏水性
及麦角固醇的含量影响
菌丝形成通过增强黏附和组织侵袭能力进而影响白色念珠菌发挥毒性。当α-常春藤皂苷浓度为256μg/mL时,其可以完全抑制白色念珠菌菌丝形成,此时细胞呈现表面黏附、溶解和聚集等现象;128μg/mL浓度下则出现部分抑制,导致细胞表面褶皱并残留少部分生长中的菌丝。而氟康唑仅在浓度>32μg/mL时可以减少菌丝形成,在256μg/mL时未能实现完全抑制(图5A)。细胞表面疏水性与白色念珠菌耐药性呈现较强相关性。α-常春藤皂苷在128μg/mL的浓度下以浓度依赖性的方式使细胞表面疏水性降低85%,而氟康唑在此浓度下仅降低了20%(图5B-C)。
除此之外,鉴于α-常春藤皂苷被虚拟筛选为CYP51(麦角固醇生物合成关键酶)的抑制剂,作者猜测α-常春藤皂苷可能影响白色念珠菌中麦角固醇含量(图5D)。在16μg/mL的浓度下,α-常春藤皂苷使麦角固醇的含量相比于未处理的对照组降低90%,而氟康唑仅降低了45%。这些结果证实,与氟康唑相比,α-常春藤皂苷对麦角固醇合成抑制活性显著增强,这与其筛选条件中直接靶向CYP51发挥作用的机制相符。
图5 α-常春藤皂苷和氟康唑对白色念珠菌致病因子的作用及联合影响。
07
α-常春藤皂苷与氟康唑对
白色念珠菌的协同抗真菌作用
联合使用抗真菌药物可降低单一用药引发的耐药风险。本文采用部分抑菌浓度指数(FICI)评估了α-常春藤皂苷(a-hederin)与氟康唑的协同抗真菌作用。如图5E所示,FICI平均值为0.5078,观测到的最低FICI值为0.2578,该结果提示两药联用具有叠加抗真菌效应。单独使用氟康唑时,即使在最高测试浓度(128 μg/mL)下仍未能完全抑制白色念珠菌生长,表明其实际最低抑菌浓度高于该阈值。尽管将128 μg/mL保守地视作氟康唑的MIC参考值,联合用药在显著更低的浓度下(氟康唑1 μg/mL + α-常春藤皂苷16 μg/mL)仍表现出叠加效果。这些结果表现了α-常春藤皂苷在增强氟康唑抗真菌活性的同时降低治疗剂量的潜力。
08
α-常春藤皂苷对白色念珠菌
口腔感染小鼠的治疗作用
以BALB/c小鼠建立白色念珠菌口腔感染小鼠为模型,分别给予不同浓度的α-常春藤皂苷进行治疗(以氟康唑作为阳性对照)。治疗期间小鼠体重变化如图6C所示。从建模时直至安乐死,正常对照组体重稳步上升,而感染模型组在感染后体重持续下降。然而,从建模后第二天开始,使用128 μg/mL α-常春藤皂苷、32 μg/mL α-常春藤皂苷和128 μg/mL氟康唑治疗的小鼠体重均出现回升。值得注意的是,128 Hed组小鼠初始体重下降最缓,并在治疗期间持续增重,表明所有治疗干预均能显著缓解白色念珠菌感染引起的体重减轻。
通过过碘酸希夫(PAS)染色(图6H)和琼脂平板培养定量(图6B)检测感染小鼠舌部真菌定植情况。PAS染色显示,空白对照组未见白色念珠菌附着,舌面呈正常粉红色,表面光滑完整;相比之下,感染模型组表现出广泛的真菌黏附和组织损伤。α-常春藤皂苷治疗显著减少了真菌生长,并减轻了舌表损伤。菌落计数结果进一步证实,经α-常春藤皂苷治疗的小鼠真菌生长数量显著降低(图6D)。值得注意的是,128 Hed组未检测到任何菌落,表明该浓度可完全抑制口腔白色念珠菌感染。
关键血液学指标如图6E-G所示。与空白对照组相比,感染模型组白细胞计数(WBC)和血红蛋白(HGB)水平降低,血小板计数(PLT)升高,提示感染后出现免疫抑制状态。相比之下,所有治疗组(128 Hed、32 Hed和128 Flu)相对于感染模型组均表现出WBC和HGB水平的恢复以及PLT计数的正常化。这些结果表明,α-常春藤皂苷与氟康唑均能有效对抗白色念珠菌诱导的毒性作用,并在一定程度上恢复小鼠的免疫功能。
图6 α-常春藤皂苷对小鼠口腔感染的治疗效果
09
α-常春藤皂苷与CYP51
分子动力学机制
对α-常春藤皂苷-CYP51和氟康唑-CYP51复合物两个体系进行了分子动力学模拟。RMSD分析(图7A)显示,α-常春藤皂苷-CYP51和氟康唑-CYP51系统分别稳定在0.18nm和0.23nm,表明α-常春藤皂苷复合物具有更高的结构稳定性。RMSF值(图7B)表明,当α-常春藤皂苷结合后,CYP51的70-100和180-200残基区域的柔性增加。结合自由能计算(图7C)进一步表明,与氟康唑相比,α-常春藤皂苷复合物中50-150、170-200和400-450残基区域的能量值更低,说明这些区域在α-常春藤皂苷-CYP51体系中稳定性更强。与此一致的是,α-常春藤皂苷复合物的整体结合自由能显著低于氟康唑复合物(图7D-E)。
图7 α-hederin与氟康唑与CYP51相互作用的分子动力学模拟。
通过回转半径(Rg)分析评估了结构结合紧密性(图7F)。两个体系在40ns后均稳定在约2.3nm,反映出结构模拟过程中二者具有相似的结构紧密度和稳定性。溶剂可及表面积(SASA)分析显示(图7G),两种复合物在100 ns内的SASA值均于220–230 nm²区间波动,进一步说明CYP51在两个体系中的结构紧实程度相当。尤其是α-常春藤皂苷复合物与CYP51平均形成4-5个氢键,而氟康唑复合物仅维持1-2个氢键(图7H和7I)。这种更强的氢键网络,结合烷基相互作用、碳氢键及其他类型相互作用(图8B)等额外稳定作用,解释了α-常春藤皂苷-CYP51复合物相较于氟康唑(图8A)具有更高稳定性及更低能量状态的原因。
10
结论
α-常春藤皂苷是来源于常春藤叶的五环三萜皂苷,此前以其抗肿瘤活性闻名,可通过诱导凋亡与铁死亡抑制多种癌细胞。近期研究提示其亦具备抗真菌潜力,能破坏白念珠菌细胞膜并干扰代谢通路。本研究中,α-常春藤皂苷在32 μg/mL浓度下可完全抑制白念珠菌生长,且作用36小时后未见恢复,效果优于氟康唑。此外,该化合物还能抑制菌丝与生物膜形成。在小鼠口腔感染模型中,α-常春藤皂苷同样表现出优于氟康唑的疗效。另一候选化合物榄香素虽对白念珠菌有抑制作用,但细胞毒性较大(>8 μL/mL),且文献提示其可能存在遗传毒性,需通过结构优化改善安全性。
机制研究表明,α-常春藤皂苷通过靶向CYP51抑制麦角固醇合成,作用强于氟康唑。分子动力学模拟显示,α-常春藤皂苷与CYP51的结合更稳定,复合物结构更紧密,氢键网络更牢固,从分子层面解释了其高效抗真菌活性的原因。此外,α-常春藤皂苷与氟康唑联用表现出协同效应,可降低氟康唑用量,这得益于其既能稳定抑制CYP51,又可干扰菌丝与生物膜形成的双重机制。
11
讨论
本研究采用分子对接与药效团模型相结合的双虚拟筛选策略,以CYP51为靶点,对TCMSP数据库中的天然小分子进行了系统筛选。通过对筛选方案的迭代优化,从大量天然分子中初步鉴定出17个候选化合物。实验验证进一步将范围缩小至两种具有抗真菌活性的天然产物:α-常春藤皂苷与榄香素。尤其是α-常春藤皂苷对白念珠菌表现出强效抗真菌活性,且细胞毒性较低,其主要通过抑制菌丝形成和生物膜发育等关键毒力因子发挥作用。在小鼠口腔念珠菌病模型中的体内实验进一步证实了其治疗效果,凸显了其临床转化潜力。原子水平的分子动力学模拟阐明了α-常春藤皂苷与氟康唑和CYP51的相互作用机制,揭示出α-常春藤皂苷具有更稳定的结合特性。该工作不仅确立了α-常春藤皂苷作为抗真菌感染的潜力新型治疗候选物,也为加速抗真菌药物发现与机制研究提供了一套方法学框架。
指导教师:黄肖霄
微生物疗法
2026-01-13
点击蓝字,关注我们
南京中医药大学沈卫星科研团队成果:α-常春藤皂苷通过SNX10介导的DEPDC5降解降低肠上皮细胞的糖酵解水平
Cite: H. Feng, J. Wang, L. Tao, et al., α-hederin decreases the glycolysis level in intestinal epithelial cells via SNX10-mediated DEPDC5 degradation, J. Pharm. Anal. 15 (2025), 101301.
DOI: https://doi.org/10.1016/j.jpha.2025.101301
长按扫码
阅读全文
JPA
文章推荐
01
研究背景和研究意义
结直肠癌(CRC)发病机制尚未完全阐明,而正常肠上皮细胞(IECs)是CRC的始发位点,其基因突变、缺失或表达异常是导致CRC发生的始动因素。因此,研究正常IECs中基因的功能对于确定CRC预防靶点具有重要意义。分选连接蛋白10(SNX10)作为CRC中的抑癌基因,参与调节分子伴侣介导的自噬(CMA)活性,该过程与CRC发病机制及糖酵解过程密切相关。哺乳动物雷帕霉素靶蛋白复合物1(mTORC1)是细胞响应外部刺激调节生长与代谢及维持稳态的核心枢纽。DEP结构域蛋白5(DEPDC5)作为mTORC1的上游负调控因子,已被鉴定为胃肠道间质瘤的新型抑癌基因,而α-常春藤皂苷(α-hed)具有抗CRC效应。已有研究发现,在正常人IECs中敲低SNX10可促进糖酵解并降低DEPDC5表达,而α-hed可逆转该效应,但其具体机制尚未阐明。本研究旨在探究SNX10对DEPDC5表达的具体调控机制,以及α-hed在此过程中的作用。
02
图文摘要
03
文章亮点
SNX10敲低可促进正常人IECs中DEPDC5的溶酶体降解。
α-hed与SNX10和DEPDC5具有高亲和力相互作用。
α-hed可降低SNX10与DEPDC5之间的结合。
α-hed抑制DEPDC5的溶酶体降解。
04
文章简介
该研究聚焦CRC发生的早期机制,旨在阐明α-hed通过SNX10调控DEPDC5降解,进而影响IECs糖酵解的分子通路。研究发现,SNX10敲低会通过转录非依赖机制加速DEPDC5蛋白降解,且该降解依赖溶酶体通路而非蛋白酶体通路(图1)。本研究发现DEPDC5具有两个典型的CMA基序,是CMA底物,并与SNX10发生直接相互作用(图2)。同时,SNX10敲低增强了溶酶体功能与CMA活性,进一步推动CMA介导的DEPDC5降解(图3),并促进mTORC1在溶酶体表面定位及信号通路激活(图4)。
进一步研究揭示,α-hed能够抑制由SNX10敲低引起的溶酶体酸化增强,降低CMA活性,减少DEPDC5降解(图5)。分子对接、CETSA和DARTS实验证实,α-hed直接与SNX10和DEPDC5结合,破坏二者间的相互作用,从而抑制CMA介导的DEPDC5降解,进而阻断mTORC1通路异常激活(图6)。最终,α-hed逆转了因SNX10敲低导致的糖酵解关键酶(LDHA、PKM2)表达及活性升高(图7)。
综上,该研究首次揭示SNX10介导的DEPDC5降解是IECs恶性转化的新机制,α-hed通过上调SNX10表达、结合SNX10-DEPDC5复合物抑制其相互作用,进而阻断CMA介导的DEPDC5降解和mTORC1通路激活,最终逆转糖酵解异常升高,阻止IECs恶性转化(图8),为CRC的预防和治疗提供了SNX10、DEPDC5两个潜在靶点及α-hed这一候选药物。
05
图文导读
图1 分选连接蛋白10(SNX10)敲低加速了DEP结构域蛋白5(DEPDC5)在FHC和NCM460细胞中的降解。(A)通过蛋白质印迹法检测shNC和shSNX10细胞中SNX10和DEPDC5的蛋白表达水平。(B)通过实时定量聚合酶链反应(qPCR)检测shNC和shSNX10细胞中SNX10和DEPDC5的基因表达水平。(C、D)采用蛋白质印迹法分析 FHC 细胞中shNC与shSNX10-1(sh1)细胞(C)、NCM460细胞中shNC与shSNX10-2(sh2)细胞(D)在有无环己酰亚胺(CHX;30μM)处理不同时间(0、4、8、12h)后的DEPDC5蛋白水平。(E)通过免疫印迹检测经MG-132(10μM)、巴弗洛霉素A1(BafA1;100nM)和氯喹(CQ;20μM)处理24h的shNC和shSNX10细胞中的DEPDC5蛋白表达。数据以平均值±标准差(SD)表示,*P<0.05,**P<0.01与对照组相比;ns:无显著性差异。
图2 分选连接蛋白10(SNX10)敲低通过分子伴侣介导的自噬(CMA)依赖性自噬途径促进DEP结构域蛋白5(DEPDC5)降解。(A)DEPDC5同源蛋白序列比对。粉色阴影表示保守的CMA基序。(B)重组质粒pcDNA3.1(+)-Flag的双酶切鉴定。bp:碱基对;1:pcDNA3.1(+)-Flag;2:BglII-XhoI限制性酶切片段(理论大小:987/4455 bp);M:DNA分子量标准。(C)重组质粒pcDNA3.1(+)-DEPDC5-Flag的单酶切鉴定。bp:碱基对;1:pcDNA3.1(+)-DEPDC5-Flag;2:EcoRI限制性酶切片段(理论大小:2153/8095 bp);M:DNA分子量标准。(D)通过蛋白质印迹法检测重组蛋白pcDNA3.1(+)-DEPDC5-Flag。(E、F)分别使用Flag抗体(E)和SNX10抗体(F)进行免疫共沉淀(Co-IP)实验,验证DEPDC5和SNX10蛋白的结合作用。(G)使用激光共聚焦显微镜观察DEPDC5与SNX10的共定位,并使用ImageJ软件进行共定位的定性和定量分析。(H)DEPDC5与CMA降解途径关键蛋白的免疫共沉淀实验。(I)在Earle平衡盐溶液(EBSS)诱导CMA激活条件下,DEPDC5与CMA关键蛋白的免疫共沉淀实验。(J)SNX10与CMA相关蛋白的免疫共沉淀实验。(K)对shNC和shSNX10肠上皮细胞(IECs)进行溶酶体分离,随后使用指定抗体进行蛋白质印迹分析。(L)使用激光共聚焦显微镜观察DEPDC5与溶酶体相关膜蛋白2A型(LAMP-2A)的共定位情况,并使用ImageJ软件进行共定位的定性和定量分析。(M、N)在SNX10敲低后,通过实时定量聚合酶链反应(qPCR)(M)和蛋白质印迹法(N)检测CMA相关分子的基因和蛋白表达。数据以平均值±标准差(SD)表示。*P<0.05,**P<0.01与shNC组相比。sh1:shSNX10-1;sh2:shSNX10-2;GAPDH:甘油醛-3-磷酸脱氢酶;LAMP-2A:溶酶体相关膜蛋白2A型;HSC70:热休克同源蛋白70。
图3 分选连接蛋白 10(SNX10)敲低上调溶酶体功能和分子伴侣介导的自噬(CMA)活性。(A) 采用免疫印迹法检测shNC和shSNX10细胞在有无氯喹(CQ)和Earle平衡盐溶液(EBSS)处理下,DEP结构域蛋白5(DEPDC5)和CMA相关蛋白的表达水平。(B) 激光共聚焦显微镜观察shNC和shSNX10细胞中内源性甘油醛 - 3 - 磷酸脱氢酶(GAPDH)(黄色)与溶酶体相关膜蛋白 2A(LAMP-2A)(红色)的共染色情况。(C) 有无CQ和EBSS处理下,采用溶酶体红色荧光探针(Lyso-Tracker Red,50nM,孵育1h)染色后的细胞代表性图像。(D)采用流式细胞术对Lyso-Tracker Red(50nM,孵育1h)染色后的溶酶体活性进行定量分析。数据以平均值±标准差(SD)表示。*P<0.05,**P<0.01与对照组相比;ns:无显著性。sh1:shSNX10-1;sh2:shSNX10-2;GAPDH:甘油醛-3-磷酸脱氢酶;LAMP-2A:溶酶体相关膜蛋白2A型;HSC70:热休克同源蛋白70。
图4 分选连接蛋白10(SNX10)敲低促进了哺乳动物雷帕霉素靶蛋白复合物1(mTORC1)在溶酶体表面的定位,从而激活了mTORC1信号通路。(A)使用共聚焦显微镜观察shNC和shSNX10肠上皮细胞(IECs)中哺乳动物雷帕霉素靶蛋白(mTOR)(黄色)和溶酶体相关膜蛋白2(LAMP2)(红色)的共定位,并使用ImageJ软件进行皮尔逊相关性分析。(B)通过蛋白质印迹法检测SNX10敲低后肠上皮细胞中DEP结构域蛋白5(DEPDC5)是否发生磷酸化修饰。(C)shNC和shSNX10肠上皮细胞中mTORC1通路相关蛋白的表达。(D)雷帕霉素处理后,通过蛋白质印迹法检测与mTORC1信号相关的蛋白。(E)通过溶酶体分离实验检测在有无SNX10敲低的FHC和NCM460细胞系中mTORC1在溶酶体表面的定位情况。数据以平均值±标准差(SD)表示。*P<0.05,**P<0.01与对照组相比。sh1:shSNX10-1;sh2:shSNX10-2;Raptor:mTOR调节相关蛋白;GβL:G蛋白β亚基样蛋白。
图5 α-常春藤皂苷(α-hed)通过抑制分子伴侣介导的自噬(CMA)活性来抑制分选连接蛋白10(SNX10)介导的DEP结构域蛋白5(DEPDC5)的溶酶体降解。(A)FHC shSNX10-1(sh1)和NCM460 shSNX10-2(sh2)细胞与0、5、10、20、40和80μM α-hed孵育24h后,通过3-(4,5-二甲基-2-噻唑)-2,5-二苯基溴化四氮唑噻唑蓝(MTT)法测定细胞活力。计算并标注半数抑制浓度(IC50)。(B、C)通过免疫荧光(B)和流式细胞术(C)检测和分析α-hed对溶酶体活性的影响。(D、E)通过蛋白质印迹法(D)和免疫荧光(E)检测α-hed对shSNX10细胞CMA活性的影响。(F)通过免疫印迹法检测在无处理或Earle平衡盐溶液(EBSS)处理和/或α-hed干预后,shNC和shSNX10细胞内的CMA活性。数据以平均值±标准差(SD)表示。*P<0.05,**P<0.01与对照组相比,#P<0.05,##P<0.01与shSNX10相比。α-hed L:低浓度α-常春藤皂苷;α-hed M:中等浓度α-常春藤皂苷;α-hed H:高浓度α-常春藤皂苷;GAPDH:甘油醛-3-磷酸脱氢酶;LAMP-2A:溶酶体相关膜蛋白2A型;HSC70:热休克同源蛋白70。
图6 α-常春藤皂苷(α-hed)通过阻断分子伴侣介导的自噬(CMA)介导的DEP结构域蛋白5(DEPDC5)降解来抑制哺乳动物雷帕霉素靶蛋白复合物1(mTORC1)通路活性。(A)使用TIMER数据库分析分选连接蛋白10(SNX10)和DEPDC5蛋白之间的相关性。(B、C)通过实时定量聚合酶链反应(qPCR)(B)和蛋白质印迹法(C)检测正常人肠上皮细胞(IECs)和结直肠癌(CRC)细胞系中SNX10的基因和蛋白表达。(D)SNX10和DEPDC5的皮尔逊相关性散点图。(E、F)α-hed与SNX10(E)和DEPDC5(F)的三维和二维对接。(G)α-hed与SNX10–DEPDC5复合物的三维和二维对接。(H、I)使用细胞热位移分析(CETSA)评估SNX10(H)和DEPDC5(I)蛋白的稳定性。(J)通过药物亲和反应靶点稳定性(DARTS)实验测定α-hed与SNX10和DEPDC5的结合。(K)使用激光共聚焦显微镜观察经α-hed处理的FHC shSNX10-1(sh1)和NCM460 shSNX10-2(sh2)细胞中哺乳动物雷帕霉素靶蛋白(mTOR)(黄色)和溶酶体相关膜蛋白2(LAMP2)(红色)的共定位情况,并使用ImageJ软件进行皮尔逊相关性分析。(L)α-hed对SNX10mRNA表达的影响。(M)α-hed对SNX10和DEPDC5蛋白表达的影响。(N)通过免疫印迹法测定在无处理或Earle平衡盐溶液(EBSS)和/或α-hed干预下,shNC和shSNX10细胞的mTORC1通路活性。(O、 P)通过免疫共沉淀(Co-IP)并使用HSC70抗体(O)和LAMP-2A抗体(P)进行蛋白质印迹分析,检测α-hed给药后shSNX10细胞中DEPDC5与热休克同源蛋白70(HSC70)和溶酶体相关膜蛋白2A型(LAMP-2A)的相互作用。(Q)在指定时间点通过3-(4,5-二甲基-2-噻唑)-2,5-二苯基溴化四氮唑噻唑蓝(MTT)法测定细胞活力。(R、S)分别使用SNX10抗体(R)和Flag抗体(S)进行免疫共沉淀实验,检测FHC sh1和NCM460 sh2细胞在有无α-hed处理下SNX10与DEPDC5的相互作用,并通过蛋白质印迹法检测输入组和免疫沉淀组中SNX10和DEPDC5的水平。数据以平均值±标准差(SD)表示。*P<0.05,**P<0.01与对照组相比,#P<0.05,##P<0.01与shSNX10相比。DMSO:二甲基亚砜;mTOR:哺乳动物雷帕霉素靶蛋白;GβL:G蛋白β亚基样蛋白;p70S6K:p70核糖体S6激酶。
图7 α-常春藤皂苷(α-hed)对分选连接蛋白10(SNX10)敲低细胞系糖酵解水平的影响。(A、B)通过蛋白质印迹法(A)和实时定量聚合酶链反应(qPCR)(B)检测参与糖酵解的分子表达,包括MYC原癌基因(c-Myc)、乳酸脱氢酶A(LDHA)和丙酮酸激酶M2(PKM2)。(C)使用商品化检测试剂盒测定shNC和shSNX10细胞中乳酸脱氢酶(LDH)和丙酮酸激酶(PK)的酶活性。(D、E)通过免疫荧光检测α-hed对shSNX10细胞LDHA(D)和PKM2(E)表达的影响。(F)检测α-hed对shSNX10细胞酶活性的影响。(G、H)通过qPCR(G)和蛋白质印迹法(H)检测不同浓度α-hed处理后shSNX10细胞c-Myc、LDHA和PKM2的表达。数据以平均值±标准差(SD)表示。*P<0.05,**P<0.01与shNC组相比,#P<0.05,##P<0.01与shSNX10相比。α-hed L:低浓度α-常春藤皂苷;α-hed M:中等浓度α-常春藤皂苷;α-hed H:高浓度α-常春藤皂苷。sh1:shSNX10-1;sh2:shSNX10-2。
图8 α-常春藤皂苷恢复正常人肠上皮细胞(IECs)糖酵解水平并阻止其恶性转化的潜在机制图。SNX10:分选连接蛋白10;DEPDC5:DEP结构域蛋白5;HSC70:热休克同源蛋白70;LAMP-2A:溶酶体相关膜蛋白2A型;mTORC1:哺乳动物雷帕霉素靶蛋白复合物1;IECs:肠上皮细胞;c-Myc:MYC原癌基因;LDHA:乳酸脱氢酶A;PKM2:丙酮酸激酶M2。
作者简介
通讯作者
沈卫星,南京中医药大学,教授,主要研究方向:中医药防治肿瘤。
程海波,南京中医药大学,教授,主要研究方向:中医药防治肿瘤。
孙东东,南京中医药大学,教授,主要研究方向:中医药防治肿瘤。
第一作者
冯慧,南京中医药大学,博士研究生,主要研究方向:肿瘤药理。
近期阅读推荐
JPA文章推荐 | 西安交通大学龙建纲教授科研团队研究成果——线粒体膜色谱:发现线粒体靶向调节剂
Journal of Pharmaceutical Analysis 2025年第12期
期刊简介
Journal of Pharmaceutical Analysis(JPA,《药物分析学报(英文)》)创刊于2011年,由教育部主管,西安交通大学主办,是国内第一本有关药物分析的专业英文学术期刊,JPA 始终秉承服务国家重大战略需求、建设世界一流科技期刊的办刊宗旨,重点报道药物发现与药品全生命周期质量控制的新理论、新技术、新方法,临床精准用药,以及药物与生物、人工智能等交叉领域的技术方法方面的最新研究成果,为全球药物研发和药品质量控制提供高水平的国际学术交流平台,持续推动药物分析学科以及药学领域的快速发展。
JPA目前已组建一支以主编贺浪冲教授为核心的国际化的学术团队和专业的编辑出版团队,已实现编委国际化、稿源国际化、同行评议国际化、读者国际化和出版国际化。已被SCIE、PubMed、Scopus、DOAJ、中国科学引文数据库(CSCD)及中国科技论文与引文数据库(中国科技核心期刊)等多种重要国际和国内数据库和评价体系定为刊源。JPA连续8年入选“中国最具国际影响力学术期刊”。2019年入选“中国科技期刊卓越行动计划”重点期刊,2024年入选“中国科技期刊卓越行动计划二期”英文领军期刊。2024年影响因子 8.9,位于全球药理学和药学类学术期刊第14位(14/352),进入药学与药理学前4%,继续稳居于Q1区前列。2025年中科院1区,Top期刊。
收稿范围
药物分析新技术、新方法,分析药理学,药物代谢与递送,中药与天然药物,生物传感,可视化分析,生物功能分析,生物技术药物,药物分析装备,人工智能应用
收稿栏目
原创论文、综述、快报、展望、观点、新闻、社评等
期刊官网
https://www.journals.elsevier.com/journal-of-pharmaceutical-analysis
投稿网址
https://www.editorialmanager.com/jpa/Default.aspx
供稿 | 冯 慧
编辑 | 李 蕾
校对 | 朱丹丹
审核 | 王梦杰、马维娜
阅读原文
了解更多
临床研究
2025-12-16
点击蓝字,关注我们
Volume 15 · Issue 12 · December 2025
长按扫码
直接阅读
Review papers
Recent insights into the roles and therapeutic potentials of GLS1 in inflammatory diseases
Jian-Xiang Sheng, Yan-Jun Liu, Jing Yu, Ran Wang, Ru-Yi Chen, Jin-Jin Shi, Guan-Jun Yang, Jiong Chen
J. Pharm. Anal. 2025. 15(12) 101292
https://doi.org/10.1016/j.jpha.2025.101292
谷氨酰胺酶1(GLS1)是谷氨酰胺代谢的关键限速酶,其催化产物在调控细胞发育、分化及免疫应答中发挥重要作用。本综述系统解析GLS1的结构与功能,重点阐述其通过谷氨酰胺代谢调控炎症信号通路的机制。现有证据表明,GLS1在银屑病、骨关节炎及非酒精性脂肪性肝炎等炎症性疾病中起关键调节作用。BPTES、CB839、DON等GLS1抑制剂在多种炎症模型中展现出显著抗炎活性,其构效关系优化为新药研发提供方向。然而,仍存在组织特异性调控机制不清、抑制剂选择性不足等问题。本文提出结合单细胞代谢组学与AI辅助药物设计的策略,为靶向GLS1的代谢调控疗法提供了理论依据,对炎症性疾病的精准干预具有重要启示。
Highlights
GLS1 is the first rate-limiting enzyme mediating glutamine decomposition.
GLS1 is implicated in various physiological and pathological processes, including inflammation.
GLS1 is a potential therapeutic target for multiple inflammatory diseases.
No GLS1 inhibitor is clinically available up to now.
Advancements in plant-derived exosome-like vesicles: Versatile bioactive carriers for targeted drug delivery systems
Haixia Shen, Shuaiguang Li, Liyuan Lin, Qian Wu, Zhonghua Dong, Wei Xu
J. Pharm. Anal. 2025. 15(12) 101300
https://doi.org/10.1016/j.jpha.2025.101300
外泌体是由各种细胞分泌的纳米级囊泡,广泛存在于动物、植物及微生物中。其凭借优异的生物相容性、高效的递送能力与跨膜运输特性,被视为具有前景的药物递送载体。与动物来源外泌体相比,植物来源外泌体(PDEs)因其低毒性、多来源及更强的靶向性而更具应用优势。随着相关研究与技术的发展,研究者已从多种植物中成功分离外泌体并探索其临床潜力。但PDEs的精准鉴定和药物递送机制仍有待进一步阐明。本综述概述了PDEs的生物发生、提取与分离方法及其鉴定技术,并总结了其工程改造与药物负载策略。同时,讨论了PDEs在疾病治疗中的应用前景及其作为药物递送平台的潜在价值。期望通过对靶向递送机制的认识,为临床药物输送体系的发展提供新的思路。
Highlights
Plant-derived exosomes are less likely to cause immune reactions and are biocompatible.
Plant-derived exosomes are ideal carriers for drug delivery and gene therapy.
Plant-derived exosomes can precisely regulate their loadings to meet different application requirements.
Therapeutic strategies based on macrophages and their derivatives: Targeted drug delivery platforms and disease treatment
Jiali Fu, Shiyun Huang, Anqi Zhang, Rongying Shi, Yuhao Wei, Shanshan He, Shiqi Huang, Lin Li, Xun Sun, Tao Gong, Ling Zhang, Qing Lin, Zhirong Zhang
J. Pharm. Anal. 2025. 15(12) 101413
https://doi.org/10.1016/j.jpha.2025.101413
传统靶向药物递送系统面临生物相容性差、递送效率低等瓶颈。巨噬细胞因其固有的炎症趋向性、吞噬能力及免疫调节功能,为突破这些障碍提供了极具潜力的天然载体。本文系统综述了基于巨噬细胞的药物递送系统(MDDS) ,重点阐述了其四大设计策略:全细胞载药、细胞膜仿生包被、细胞外囊泡递送及细胞表面“搭便车”,并深入分析了各自的优势、构建方法与技术挑战。文章进一步揭示了MDDS在癌症、动脉粥样硬化及中枢神经系统疾病中的应用机制,例如在肿瘤微环境中主动归巢、在斑块处精准递送抗炎药物以及穿越血脑屏障递送神经治疗药物。最后,文章系统梳理了MDDS面临的关键转化障碍并展望了未来优化方向,为开发新一代细胞靶向递送平台提供了新的思路。
Highlights
Classifying MDDSs by efficiency, safety, and clinical potential.
Highlighting MDDSs in cancer, atherosclerosis, and CNS diseases.
Summarizing challenges and future directions for MDDS development.
Nanometer preparation of natural bioactive compounds for treatment of rheumatoid arthritis
Junping Zhu, Qin Xiang, Liu Li, Jiaming Wei, Rong Yu
J. Pharm. Anal. 2025. 15(12) 101341
https://doi.org/10.1016/j.jpha.2025.101341
类风湿关节炎(RA)是一种以关节滑膜炎为特征的慢性自身免疫性疾病,可致关节破坏、功能障碍甚至残疾。现有疗法效果有限且易出现副作用。天然生物活性化合物(NBCs)因其免疫调节和抗炎特性备受关注,但存在水溶性差、所需剂量高、作用时间短及组织特异性低等问题。纳米颗粒(NP)递送系统,如脂质、聚合物及无机纳米载体,通过被动、主动及刺激响应等靶向策略应对上述挑战。负载NBCs的NP可作用于RA免疫紊乱、滑膜增生、骨破坏、血管生成、炎症及氧化应激等多个病理环节。本文重点介绍了NBCs在RA治疗、NP设计及靶向机制的最新进展,同时探讨了该领域的挑战与未来方向。
Highlights
This review summarizes the therapeutic potential of various NBCs for RA.
Common nanomaterials used for NBCs delivery are discussed.
The pharmacological mechanisms and biomarkers of targeted anti-RA therapies are explored.
Key challenges in the applications of nanomaterials are identified, with innovative solutions proposed.
Virulence arresting drug discovery by strategies targeting bacterial virulence: Mainly focusing on quorum-sensing interference and biofilm inhibition
Lan Lu, Tianyang Yu, Hongping Wang, Xingtong Zhu, Li Liao, Jie Zhu, Xiaobo Wang, Andi Yang, Chen Yang, Yuping Zhang, Yulin Zhang, Kun Zou, Xiaorong Yang, Mingxing Li
J. Pharm. Anal. 2025. 15(12) 101310
https://doi.org/10.1016/j.jpha.2025.101310
多重耐药病原体的日益普遍对全球医疗体系构成重大威胁,亟需紧急治疗干预。微生物对各类抗生素表现出多样化的耐药机制,凸显了发现新型抗菌药物以对抗细菌感染的紧迫性。抗毒力疗法通过靶向病原体的毒力决定因子来中和其毒性,是一种前景广阔的治疗策略。针对细菌毒力抑制药物的筛选策略是一种多维度的方法,包括阐明细菌致病性的分子发病机制、识别不同病原体间进化保守的毒力因子,并采用结合计算机预测与体内外实验验证的综合方法。本综述系统性地探讨了当前的筛选方法,主要聚焦于群体感应破坏和生物被膜抑制策略,包括计算机预测筛选、基于活性的生物测定筛选、相应的体外和体内模型等。此外,研究者强调通过相关动物模型进行标准化临床前验证的必要性,并提出了开发下一代细菌毒力抑制药物筛选平台的构想和建议,为未来新型抗菌药物的发现提供了新方向。
Highlights
Recent advances in strategies were summarized for screening potential virulence arresting drugs (VADs).
The necessity of validation in animal models through the implementation of standardized procedures was underscored.
Suggestions were put forth regarding the development of models and strategies for the screening of VADs.
Sample preparation techniques for quality evaluation and safety control of medicinal and edible plants: Overview, advances, applications, and future perspectives
Lingxuan Ma, Lele Yang, Lijun Tang, Yudi Wang, Hua Luo, Zhangfeng Zhong, Wensheng Zhang, Di Chen, Jinchao Wei, Peng Li, Yitao Wang
J. Pharm. Anal. 2025. 15(12) 101296
https://doi.org/10.1016/j.jpha.2025.101296
药食同源植物(MEPs)兼具营养与药用价值,是中医药大健康产业的重要基础,其质量稳定性与安全性直接关系公众健康。围绕MEPs活性成分精准解析与外源性污染物风险防控这一核心科学问题,本文系统梳理并深入评述了近五年该领域样品前处理技术的最新进展与应用实践。文章首次从“质量评价—安全控制”双重视角,全面整合传统提取方法、绿色微萃取技术及新型纳米材料与功能溶剂策略,突出其在提升灵敏度、实现绿色分析的技术优势,并前瞻性总结了自动化、微型化与原位分析的发展趋势。本文为MEPs质量标准完善与大健康产品安全评价提供了系统、前沿的技术参考与方法学框架。
Highlights
Outline of sample preparation techniques for quality evaluation and safety control of MEPs.
Emphasis on the enrichment and determination of endogenous bioactives and exogenous contaminants.
Profiling the merits and drawbacks of various sample preparation methods.
Sample preparation techniques are trending towards green analytical chemistry.
Decoding protein dynamics with limited proteolysis coupled to mass spectrometry: A comprehensive review
Zilu Zhao, Xue Zhang, Xin Dong, Zhanying Hong
J. Pharm. Anal. 2025. 15(12) 101319
https://doi.org/10.1016/j.jpha.2025.101319
有限蛋白水解联用质谱(LiP-MS)是一种解析近自然条件下蛋白质结构与功能的关键技术,可捕捉蛋白质动态构象变化。本文综述了LiP-MS在药物靶点识别、代谢物机制、蛋白质组动态等领域的应用,指出其在结构功能分析和生物分子互作研究中的独特优势,能助力药物机制解析与靶点开发。尽管LiP-MS存在无法识别功能机制(如磷酸化)、膜蛋白分析受限等挑战,但它仍是蛋白靶点研究的重要工具。未来,结合传统蛋白组学技术,LiP-MS有望推动生物医药及多领域分子生物学研究的发展。
Highlights
This review provides a comprehensive overview of all LiP-MS applications in protein research.
It enhances the understanding of protein dynamics and cellular responses.
It offers new perspectives for advancing drug discovery and biomarker identification.
In vivo analysis techniques for antibody drug: Recent advances and methodological insights
Xiaolu Miao, Beilei Sun, Jian Zhang, Jinge Zhao, Bing Ma, Yongming Li, Weizhi Wang
J. Pharm. Anal. 2025. 15(12) 101314
https://doi.org/10.1016/j.jpha.2025.101314
本文综述了抗体药物在活体分析中的最新技术进展与方法学见解。抗体药物,如单克隆抗体(mAbs)和抗体药物偶联物(ADCs),凭借其高特异性和亲和力,在肿瘤、自身免疫病等多种疾病的治疗中展现出巨大潜力。为深入理解其体内动态行为、药代动力学及疗效,非侵入性、实时的活体分析技术至关重要。文章重点介绍了三种活体成像技术:放射性核素标记成像、近红外荧光成像(NIRFI)以及表面增强拉曼光谱(SERS),系统阐述了各技术的原理、代表性应用、优势与挑战,并展望了多模态融合与结构解析等未来发展方向,为抗体药物的研发与临床转化提供参考。
Highlights
Progress of antibody drugs and the importance of in vivo analysis are introduced.
Non-invasive and real-time methods for in vivo analysis of antibody drugs are summarized.
The application, limitations, and possible solutions of in vivo analysis methods are discussed.
Original articles
Geometry-based BERT: An experimentally validated deep learning model for molecular property prediction in drug discovery
Xiang Zhang, Chenliang Qian, Bochao Yang, Hongwei Jin, Song Wu, Jie Xia, Fan Yang, Liangren Zhang
J. Pharm. Anal. 2025. 15(12) 101465
https://doi.org/10.1016/j.jpha.2025.101465
GEO-BERT模型通过引入分子的三维几何信息(包括原子间距离、键角和原子-键邻接关系),在Transformer架构中构建了多维注意力机制,从而显著提升了分子表征的精度与泛化能力。该模型在多个公开数据集上表现优异,优于现有主流分子表征模型,并在真实药物筛选任务中成功发现了两种高活性DYRK1A抑制剂,验证了其在实际药物研发中的应用价值。此外,研究团队还从模型可解释性与可靠性角度进行了深入分析。这些分析不仅增强了模型的透明度,也为后续药物设计与筛选提供了结构层面的参考依据。未来,GEO-BERT有望在更多靶点筛选、分子生成、个性化药物设计等领域发挥作用,推动人工智能在药物发现中的深度融合与落地应用。
Highlights
A self-supervised representation learning framework named GEO-BERT was developed for molecular property prediction.
GEO-BERT is based on a new molecular representation that incorporates pose information.
With atom-atom, bond-bond and atom-bond relationships, GEO-BERT achieves optimal performance across multiple benchmarks.
GEO-BERT facilitated identification of two potent and novel DYRK1A inhibitors, demonstrating its practical utility.
Enhancing polyreactivity prediction of preclinical antibodies through fine-tuned protein language models
Yuwei Zhou, Haoxiang Tang, Changchun Wu, Zixuan Zhang, Jinyi Wei, Rong Gong, Samarappuli Mudiyanselage Savini Gunarathne, Changcheng Xiang, Jian Huang
J. Pharm. Anal. 2025. 15(12) 101448
https://doi.org/10.1016/j.jpha.2025.101448
在抗体药物研发中,多反应性(polyreactivity)易导致抗体与非靶标蛋白发生非特异性结合,进而影响药物的稳定性、安全性及临床成功率。传统实验方法耗时费力,难以在早期高通量且高效筛除高风险候选分子。为此,本研究基于六种蛋白质语言模型构建了多反应性预测工具,其中基于ESM-2微调的PolyXpert表现最优,预测性能显著领先。结果表明,微调后的模型能更精准捕捉抗体序列中的多反应性特征,并具备优异的泛化能力与鲁棒性。PolyXpert为抗体早期开发提供了快速、精准的多反应性AI评估方案,有助于提升筛选效率、降低研发成本,推动高质量抗体进入后续开发。该工具已开源,研究者可在https://github.com/zzyywww/PolyXpert获取使用。
Highlights
We fine-tuned protein language models to develop PolyXpert for assessing antibody polyreactivity.
PolyXpert performs exceptionally well in predicting the polyreactivity of antibody sequences.
PolyXpert is poised to help select therapeutic mAbs candidates with low polyreactivity for clinical development.
We provide user-friendly usage guidance, simplifying the use and exploration of PolyXpert for researchers.
From foe to friend: Rewiring oncogenic pathways through artificial selenoprotein to combat immune-resistant tumor
Weiming You, Zhengjun Zhou, Zhanfeng Li, Jin Yan, Yang Wang
J. Pharm. Anal. 2025. 15(12) 101322
https://doi.org/10.1016/j.jpha.2025.101322
微卫星稳定型结直肠癌(CRC)常因促癌信号通路活跃而产生免疫耐受。本研究针对CRC中高表达的鼠双微体2同源物(MDM2)通路,开发了人工硒蛋白Path-editor。该药物搭载PMI和PPI肽段,进入细胞后释放:PMI抑制MDM2介导的p53降解,恢复抑癌功能并促进热休克同源蛋白70(HSC70)表达;PPI则结合HSC70以选择性降解程序化死亡配体1(PD-L1)。在CT26模型和人源肿瘤异种移植模型中,Path-editor显著降低了PD-L1水平,增强了细胞毒性T淋巴细胞浸润,抑瘤效果优于PD-1抗体。生化指标显示该疗法安全性良好,无明显系统毒性。本研究通过将促癌通路重构为免疫激活轴,为治疗难治性恶性肿瘤提供了新策略。
Highlights
Reprogram MDM2-HSC70 axis triggers antitumor immunity via selenoprotein Path-editor.
Dual-peptide Path-editor degrades PD-L1 and boosts p53 for overcoming immune evasion.
Selenium-driven pathway editing inhibits tumor growth with high biosafety in vivo.
Naringenin boosts Parkin-mediated mitophagy via estrogen receptor alpha to maintain mitochondrial quality control and heal diabetic foot ulcer
Xin-Meng Zhou, Ying Yang, Dao-Jiang Yu, Teng Xie, Xi-Lu Sun, Ying-Xuan Han, Hai-Ying Tian, Qing-Qing Liao, Yu-Jie Zhao, Yih-Cherng Liou, Wei Huang, Yong Xu, Xi Kuang, Xiao-Dong Sun, Yuan-Yuan Zhang
J. Pharm. Anal. 2025. 15(12) 101333
https://doi.org/10.1016/j.jpha.2025.101333
糖尿病足(DFU)是糖尿病高发并发症,致残致死率高,目前缺乏特效治疗药物。柚皮素(NAR)具有雌激素样作用及多效药理活性。本研究通过C57BL/6J野生型及Prkn敲除小鼠DFU模型,以及高糖诱导HaCaT细胞实验,探究NAR的治疗作用及机制。发现NAR局部应用可显著加速DFU创面愈合,减轻氧化应激、炎症及细胞衰老凋亡。机制上,细胞热迁移实验证实NAR特异性可结合ERα,增强ERα与雌激素反应元件(ERE)的结合,上调Parkin转录并促进其线粒体移位,激活Parkin介导的线粒体自噬,维持线粒体质量控制平衡。ER调节剂他莫昔芬或Parkin敲除均可阻断NAR的保护作用。由此证实,NAR通过结合ERα激活Parkin介导的线粒体自噬,为DFU治疗提供了新机制。
Highlights
NAR promotes DFU healing by maintaining MQC.
NAR's beneficial effects are dependent on Parkin.
NAR directly binds to ERα.
NAR enhances the binding of ERα to ERE to upregulate the transcription of Parkin.
Unveiling optimal molecular features for hERG insights with automatic machine learning
Congying Xu, Youjun Xu, Ziang Hu, Xinyi Zhao, Weixin Xie, Weiren Chen, Jianfeng Pei
J. Pharm. Anal. 2025. 15(12) 101411
https://doi.org/10.1016/j.jpha.2025.101411
本研究提出了MaxQsaring——一种融合分子描述符、分子指纹与深度学习预训练表征的通用框架,用于化合物性质的预测。以人ether-à-go-go-related gene(hERG)钾离子通道抑制剂为案例,MaxQsaring通过自动优化特征组合,在两个高挑战性外部数据集上取得了优秀的预测性能,并成功筛选出十大关键可解释特征,可用于构建高精度决策树模型。模型预测结果与经验性hERG抑制剂优化策略高度吻合,彰显其在实际场景中的可解释价值。尽管深度学习预训练表征对提升预测模型性能具有中等程度的影响,但在拓展模型对全新骨架化合物的泛化能力方面作用有限。MaxQsaring在Therapeutics Data Commons(TDC)测试中表现卓越,在22项任务中的19项位列第一,充分验证其作为通用高精度化合物性质预测平台的潜力,有望提升早期药物研发的成功率。
Highlights
A novel universal framework to automatic QSAR model building.
An interpretable hERG prediction model with high accuracy.
Automatic feature screening of molecular descriptors, fingerprints, and deep-learning pretrained representations.
α-hederin decreases the glycolysis level in intestinal epithelial cells via SNX10-mediated DEPDC5 degradation
Hui Feng, Jin Wang, Lihuiping Tao, Liu Li, Minmin Fan, Chengtao Yu, Dongdong Sun, Haibo Cheng, Weixing Shen
J. Pharm. Anal. 2025. 15(12) 101301
https://doi.org/10.1016/j.jpha.2025.101301
结直肠癌(CRC)是全球高发且致命的恶性肿瘤,肠上皮细胞(IECs)作为CRC发生的主要部位,其基因功能异常与癌变密切相关。分选连接蛋白10(SNX10)是CRC易感候选基因及抑癌基因,DEP结构域包含蛋白5(DEPDC5)作为mTORC1通路的上游负调控因子,其在CRC中的调控机制尚未明确。本研究发现SNX10在翻译后水平维持DEPDC5蛋白稳定性,DEPDC5是新型SNX10相互作用蛋白及分子伴侣介导的自噬(CMA)底物;SNX10通过溶酶体途径调控DEPDC5降解,激活mTORC1通路并促进糖酵解,进而推动IECs恶性转化。而α-常春藤皂苷可破坏SNX10与DEPDC5的相互作用,抑制DEPDC5溶酶体降解,最终逆转由SNX10敲低引起的糖酵解水平升高。该研究为CRC预防提供了理论基础,并提示靶向SNX10的基因疗法或α-hederin方案在CRC防治中的潜在价值。
Highlights
SNX10 knockdown promotes DEPDC5 lysosome degradation in normal human IECs.
α-hederin interacts with SNX10 and DEPDC5 with a high affinity.
α-hederin reduces the binding between SNX10 and DEPDC5.
α-hederin inhibits the lysosomal degradation of DEPDC5.
Targeted reduction-responsive nanovehicles for photodynamic therapy-primed immunotherapy in melanoma
Chenqian Feng, Lingfeng Zhou, Bo Chen, Hui Li, Min Mu, Rangrang Fan, Haifeng Chen, Gang Guo
J. Pharm. Anal. 2025. 15(12) 101311
https://doi.org/10.1016/j.jpha.2025.101311
本文围绕黑色素瘤治疗中光动力治疗与免疫治疗协同不足的问题,构建了一种靶向还原响应型纳米递送系统 BM@HSSC。该体系以透明质酸为骨架,通过二硫键(-S-S-)连接光敏剂二氢卟吩e6,并共载小分子免疫检查点抑制剂BMS-1,实现肿瘤微环境高谷胱甘肽触发的精准释药。纳米颗粒可经 CD44受体靶向富集于肿瘤部位,在激光照射下产生活性氧,诱导肿瘤细胞凋亡并触发免疫原性细胞死亡(ICD),促进树突状细胞成熟与T细胞活化。同时,BMS-1阻断程序性死亡受体-1/程序性死亡配体-1(PD-1/PD-L1)通路,逆转免疫抑制性肿瘤微环境。该策略在黑色素瘤模型中展现出显著的抗肿瘤效果与长效免疫记忆,为肿瘤联合治疗提供了新思路。
Highlights
BM@HSSC nanoparticles target tumors through CD44 receptors and release Ce6 and BMS-1 in response to high glutathione levels.
Elevated oxidative stress induces ICD, releasing DAMPs, which promote DCs maturation and T-cell infiltration.
The immunosuppressant BMS-1 blocks the PD-1/PD-L1 pathway, reversing the immunosuppressive tumor microenvironment.
The combination of BM@HSSC NPs-mediated PDT with BMS-1 significantly inhibits distal tumor progression.
A customizable continuous and near real-time TEER platform to study anti-cancer drug toxicity in barrier tissues
Curtis G. Jones, Chengpeng Chen
J. Pharm. Anal. 2025. 15(12) 101266
https://doi.org/10.1016/j.jpha.2025.101266
Highlights
This paper reports an unprecedented TEER sensor system to investigate drug toxicity in barrier tissues.
This system included a customizable and near real-time TEER sensor and a modular microfluidic device to integrate endothelium in relevant extracellular matrices.
We report for the time how doxorubicin, a common anti-cancer drug, could impact endothelium permeability in 1-min resolution.
The system was based on low-cost parts and 3D printing (engineering sketches provided in the SI) with vast customization room.
Unlocking the potential of atractylenolide II: Mitigating non-alcoholic fatty liver disease through farnesoid X receptor-endoplasmic reticulum stress interplay
Ming Gu, Zhiwei Chen, Yujun Chen, Yiping Li, Hongqing Wang, Ya-ru Feng, Peiyong Zheng, Cheng Huang
J. Pharm. Anal. 2025. 15(12) 101318
https://doi.org/10.1016/j.jpha.2025.101318
法尼醇X受体(FXR)激活可通过缓解内质网应激(ERS)来减轻非酒精性脂肪性肝病(NAFLD),但二者相互作用的具体机制尚不明确。本研究发现,FXR激活可直接上调肌浆/内质网钙ATP酶2(SERCA2)的转录,进而促进肝细胞eIF2α的去磷酸化,最终缓解ERS。研究者还鉴定出白术内酯II(AT-II)为一种新型天然FXR激动剂,可有效激活SERCA2,并抑制体外肝细胞脂质蓄积与ERS;体内实验表明其可改善饮食或化学诱导的小鼠肝脏脂肪变性,并缓解肥胖、血脂异常、胰岛素抵抗等代谢紊乱。机制上,AT-II通过激活FXR-SERCA2-eIF2α轴,减轻ERS、抑制脂质合成、缓解炎症并改善胰岛素信号。Serca2敲低及FXR沉默实验更进一步验证了这点。本研究阐明了FXR调控ERS对抗NAFLD的新分子机制,并提示AT-II是靶向FXR-SERCA2通路的潜在治疗药物。
Highlights
FXR activation inhibits hepatic ER stress by inducing SERCA2 transcriptionally.
Atractylenolide II (AT-II) is identified as a novel FXR activator.
AT-II potentiates SERCA2 in hepatocytes by activating FXR.
AT-II attenuates ER stress and NAFLD via promoting FXR-SERCA2 axis.
Dephosphorylation of eIF2α by SERCA2 is involved in AT-II's anti-NAFLD.
Reactivating T cell immunity in Wnt-hyperactivated non-small cell lung cancer through a supramolecular droplet of carnosic acid and peptide
Na Liu, Yuzhen Tu, Hanyu Wang, Xiaoqiang Zheng, Fanpu Ji, Mingsha Geng, Xin Wei, Jingman Xin, Wangxiao He, Qian Zhao, Tianya Liu
J. Pharm. Anal. 2025. 15(12) 101309
https://doi.org/10.1016/j.jpha.2025.101309
非小细胞肺癌(NSCLC)中Wnt/β-catenin信号通路的异常激活与肿瘤免疫抑制微环境的形成密切相关,导致T细胞浸润不足与免疫治疗耐受。本研究针对该挑战,设计了一种酸敏感性肽-鼠尾草酸(CA)超分子液滴(Pep1@CA),通过靶向肿瘤酸性微环境实现药物的可控释放与精准递送。本研究首先通过代谢组学分析发现Wnt高活化NSCLC具有显著的肿瘤区域酸积聚特征。基于此,团队构建的Pep1@CA在酸性条件下发生电荷反转,增强肿瘤细胞的内吞作用,并在细胞内释放CA,从而特异性抑制Wnt信号通路。实验结果表明,Pep1@CA在多种NSCLC小鼠模型中均表现出显著的抗肿瘤效果,并能有效恢复CD8⁺ T细胞浸润、降低调节性T细胞(Tregs)比例,重塑肿瘤免疫微环境。此外,该递送系统显示出良好的生物安全性,为Wnt通路靶向治疗与免疫联合治疗提供了具有临床转化潜力的新策略。
Highlights
Targeting Wnt/β-catenin pathway in NSCLC effectively achieved antitumor effects and restored T lymphocyte infiltration in TME.
Design a Supramolecular Droplet of Carnosic Acid and Peptide.
Pep1@CA restored intratumoural CD8+ T cell infiltration and reduced Tregs in the subcutaneous tumor-carrying mouse model.
1,8-Cineole ameliorates vascular endothelial senescence in diabetes mellitus by directly targeting and deubiquitinating PPAR-γ in vivo and in vitro
Lingyun Fu, Shidie Tai, Jiajia Liao, Youqi Du, Guangqiong Zhang, Die Guo, Xingmei Chen, Tian Zheng, Xiaoxia Hu, Wenbing Yao, Ling Tao, Xueting Wang, Yini Xu, Xiangchun Shen
J. Pharm. Anal. 2025. 15(12) 101307
https://doi.org/10.1016/j.jpha.2025.101307
血管内皮细胞衰老是糖尿病心血管并发症的关键病理生理因素。1,8-桉叶油素(1,8-cineole)是具有抗炎、抗菌及抗氧化活性的单萜类化合物。本研究通过生物信息学分析,揭示了1,8-cineole作用于糖尿病及衰老的核心靶点与信号通路;结合分子对接、分子动力学模拟、细胞热位移实验、药物亲和反应靶向稳定性实验及表面等离子共振技术,阐明了其与靶标的相互作用机制;研究证实1,8-cineole通过靶向PPAR-γ抑制其Lys-466位点的泛素化和降解,改善ROS累积、衰老相关分泌表型及DNA损伤,从而缓解糖尿病血管内皮细胞衰老,抑制糖尿病血管并发症。本研究证实抑制PPAR-γ的Lys-466位点泛素化降解是临床防治糖尿病血管并发症的有益的防治策略和靶位,1,8-cineole是防治糖尿病血管并发症的潜在有效候选化合物,为其临床应用转化应用提供了理论指导和实验基础。
Highlights
PPAR-γ was a potential therapeutic target of 1,8-cineole in DM-related senescence.
1,8-Cineole alleviated lipid abnormalities, vascular injury, and vascular senescence in DM.
1,8-Cineole ameliorated vascular endothelial senescence in DM by deubiquitinated PPAR-γ at the Lys-466 site.
Metabolomics-driven elucidation of the synergistic therapeutic mechanism of a novel SGLT-2/PPAR-γ dual receptor supramolecular system for treatment diabetes and obesity
Saisai Ren, Han Hao, Wei Guo, Mo Zhang, Honglin Feng, Jing Wang
J. Pharm. Anal. 2025. 15(12) 101308
https://doi.org/10.1016/j.jpha.2025.101308
含活性药物成分(APIs)的超分子体系可改善药物理化性质,并实现两组分间的协同增效,但其体内作用机制尚不明确。代谢功能障碍可加剧胰岛素抵抗与肥胖,进而导致肝脏脂肪变性与心肌肥厚。为此,研究者构建了一种新型SGLT-2/PPAR-γ双受体(达格列净-吡格列酮,DAP-PIO)超分子体系,该体系在糖尿病与肥胖治疗中展现出协同疗效。代谢组学分析揭示了DAP/PIO简单物理混合物(PM)与DAP-PIO超分子体系吸收血液后的代谢差异,指出其协同作用与PI3K/AKT和AMPK信号通路密切相关。基于上述通路,通过体内外实验进一步验证了DAP-PIO超分子体系可通过改善胰岛素抵抗,协同发挥降血糖、改善肝脏脂肪变性及逆转心肌肥厚的作用。本研究为开发具有协同增效的SGLT-2/PPAR-γ双受体超分子体系提供了创新性策略。
Highlights
A novel SGLT-2/PPAR-γ dual receptor supramolecular system was prepared.
Metabolomics was used to elucidate the synergistic therapeutic mechanism in diabetes and obesity.
The supramolecular activated PI3K/AKT and AMPK pathways featured by sphingolipid metabolism.
The insulin resistance was relieved effectively by supramolecular in the treatment of diabetes and obesity.
Short communications
Intestinal anti-inflammatory drug targets as potential modifiers of cardiovascular disease risk
Shuangshuang Tong, Junjun Ye, Yanlin Lyu, Jiating Su, Baoxin Yan, Xianzhen Cai, Barkat Ali Khan, Muhammad Azhar Ud Din, Kaijian Hou, Jilin Li
J. Pharm. Anal. 2025. 15(12) 101403
https://doi.org/10.1016/j.jpha.2025.101403
心血管疾病(CVD)是全球首要死因,慢性炎症是其核心病理机制之一。观察性研究虽提示肠道抗炎药物(如5-氨基水杨酸类)或具心血管保护作用,但二者间因果关系尚不明。为填补此空白,本研究运用综合性遗传学分析框架,包括孟德尔随机化、基于汇总数据的孟德尔随机化以及共定位分析,系统评估了肠道抗炎药物靶点与九种心血管结局之间的因果关联。结果表明,部分肠道抗炎药物靶点(TPMT、PPARG、NOS2)是CVD风险的潜在调控因子。其中,TPMT被鉴定为心房颤动的关键保护性靶点。基于此,其抑制剂奥沙拉嗪展现出用于心房颤动治疗的药物再利用潜力。本研究为肠道抗炎治疗的心血管效应提供了新的因果证据,并为CVD的药物开发和精准干预提供了新的思路。
Highlights
Inflammatory bowel disease and cardiovascular disorders exhibit significant comorbidities.
Mendelian randomization identifies novel drug targets for cardiovascular disorders through genetic evidence.
Increased expression of TPMT in the atrial appendage is associated with an elevated risk of atrial fibrillation.
Mendelian randomization findings are supported by SMR and colocalization analyses.
Olsalazine emerges as a promising therapeutic candidate for atrial fibrillation through TPMT modulation.
Identification of forsythoside A from Forsythia fruit for alleviating MAFLD via metabolic remodeling and IL-17 pathway regulation
Chenglin Song, Yuxi Huang, Xiaolin Sa, Linlin Wang, Mingju Yao, Zhengtong Jin, Yang Sun, Min Ye, Xue Qiao
J. Pharm. Anal. 2025. 15(12) 101321
https://doi.org/10.1016/j.jpha.2025.101321
代谢功能障碍相关脂肪性肝病(MAFLD)是全球高发的代谢性疾病之一,目前临床治疗手段极为有限。连翘作为治疗MAFLD的经典中药,其抗MAFLD的活性成分及作用机制尚未明确。本研究首先通过细胞水平降脂活性筛选,证实以连翘酯苷A(FA)为代表的连翘苯乙醇苷类成分具有显著降脂效应。进一步采用蛋氨酸-胆碱缺乏饮食构建MAFLD小鼠模型,结合代谢组学、网络药理学深入分析,发现连翘酯苷A可有效逆转MAFLD小鼠的肝脏代谢紊乱。机制研究揭示FA通过靶向抑制IL-17信号通路,降低丝裂原活化蛋白激酶家族蛋白磷酸化水平发挥改善MAFLD作用。此外首次发现FA能显著下调MAFLD病理状态下苯丙氨酸-脯氨酸等四种二肽的异常表达水平,明确了以FA为核心的苯乙醇苷类成分在改善MAFLD中的关键作用。
Highlights
Forsythoside A (FA) was identified from Forsythia fruit to effectively alleviate MAFLD.
FA alters MAFLD mice liver metabolism, reducing elevated dipeptides such as Phe-Pro.
FA suppresses IL-17 signaling pathway and MAPK family protein phosphorylation.
近期阅读:
JPA文章推荐 | 厦门医学院王琛教授科研团队成果:靶向黑色素瘤的氧化石墨烯掺杂微针用于光热-化疗协同治疗
JPA文章推荐|北京中医药大学高晓燕科研团队成果——基于前药探针的原位生物转化鉴定番泻苷A致泻靶点
期刊简介
Journal of Pharmaceutical Analysis(JPA,《药物分析学报(英文)》)创刊于2011年,由教育部主管,西安交通大学主办,是国内第一本有关药物分析的专业英文学术期刊,JPA 始终秉承服务国家重大战略需求、建设世界一流科技期刊的办刊宗旨,重点报道药物发现与药品全生命周期质量控制的新理论、新技术、新方法,临床精准用药,以及药物与生物、人工智能等交叉领域的技术方法方面的最新研究成果,为全球药物研发和药品质量控制提供高水平的国际学术交流平台,持续推动药物分析学科以及药学领域的快速发展。
JPA目前已组建一支以主编贺浪冲教授为核心的国际化的学术团队和专业的编辑出版团队,已实现编委国际化、稿源国际化、同行评议国际化、读者国际化和出版国际化。已被SCIE、PubMed、Scopus、DOAJ、中国科学引文数据库(CSCD)等多种重要国际和国内数据库和评价体系定为刊源。JPA连续7年入选“中国最具国际影响力学术期刊”。2019年入选“中国科技期刊卓越行动计划”重点期刊,2024年入选“中国科技期刊卓越行动计划二期”英文领军期刊。2024年影响因子 8.9,位于全球药理学和药学类学术期刊第14位(14/352),进入药学与药理学前4%,继续稳居于Q1区前列。2025年中科院1区,Top期刊。
收稿范围
药物分析新技术、新方法,分析药理学,药物代谢与递送,中药与天然药物,生物传感,可视化分析,生物功能分析,生物技术药物,药物分析装备,人工智能应用
收稿栏目
原创论文、综述、快报、展望、观点、新闻、社评等
期刊官网
https://www.journals.elsevier.com/journal-of-pharmaceutical-analysis
投稿网址
https://www.editorialmanager.com/jpa/Default.aspx
编辑 | 李 蕾
校对 | 朱丹丹
审核 | 王梦杰、马维娜
阅读原文
了解更多
100 项与 α-常春藤皂苷 相关的药物交易
登录后查看更多信息
外链
| KEGG | Wiki | ATC | Drug Bank |
|---|---|---|---|
| - | - | - |
研发状态
批准上市
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| 高脂血症 | 中国 | - | - |
未上市
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 三阴性乳腺癌 | 临床前 | 中国 | 2025-03-08 | |
| 乳腺癌 | 临床前 | 斯里兰卡 | 2024-04-12 | |
| 非小细胞肺癌 | 临床前 | 中国 | 2024-02-01 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
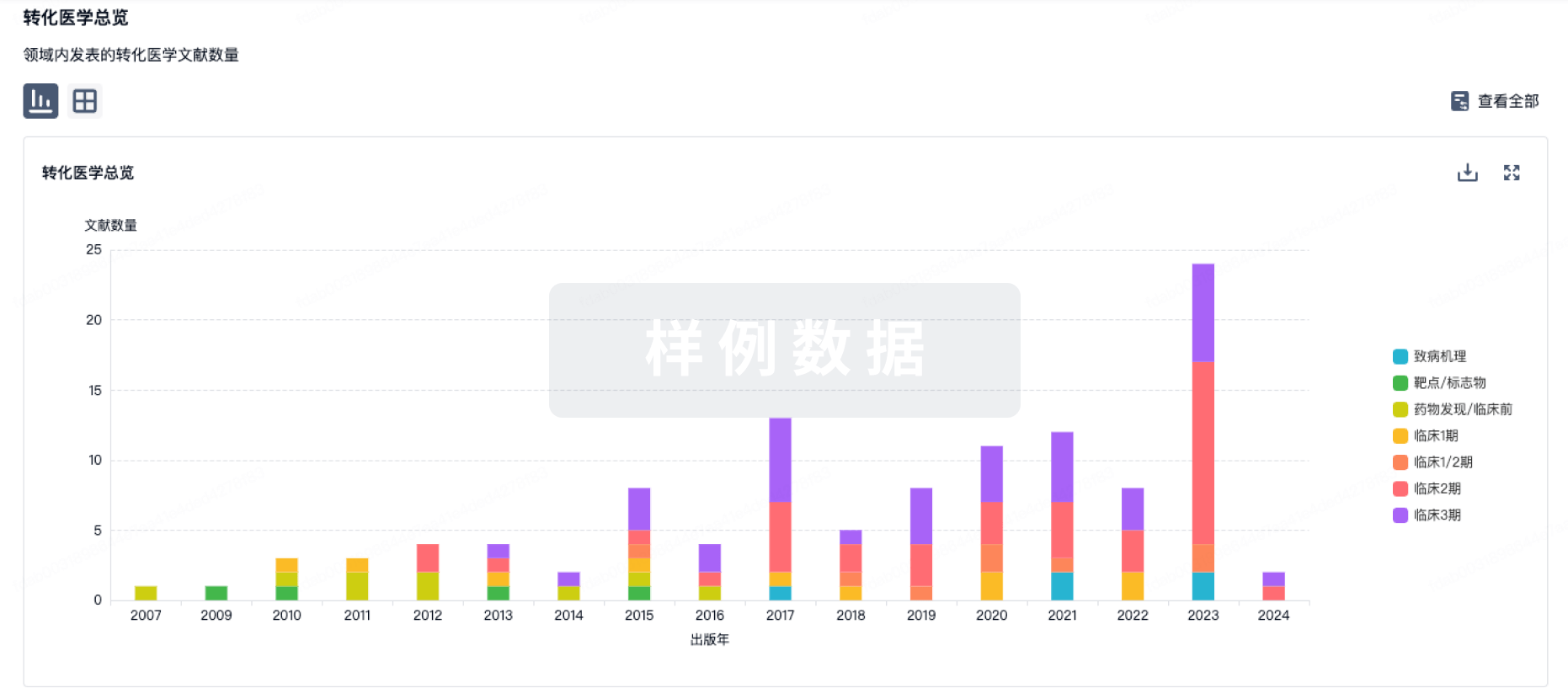
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
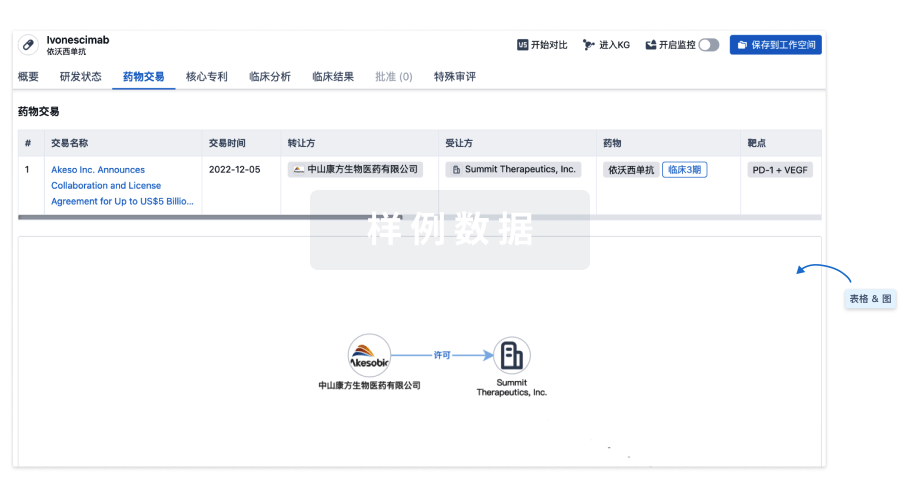
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
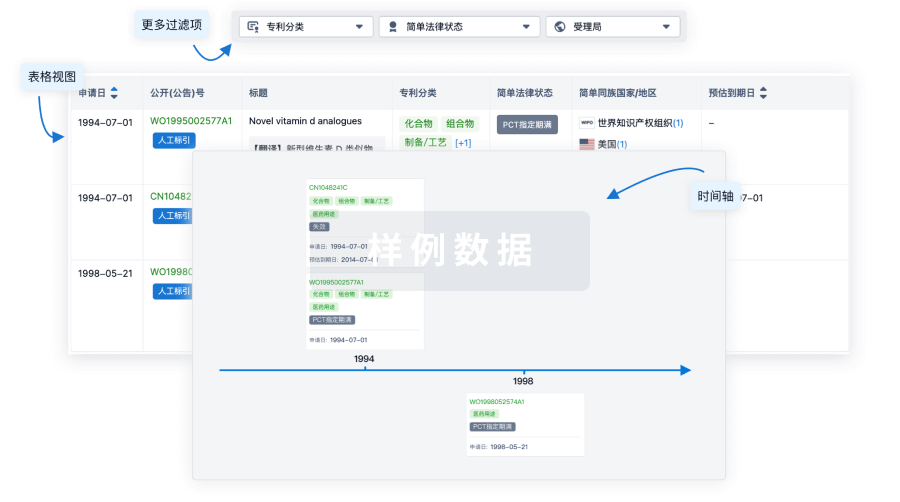
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
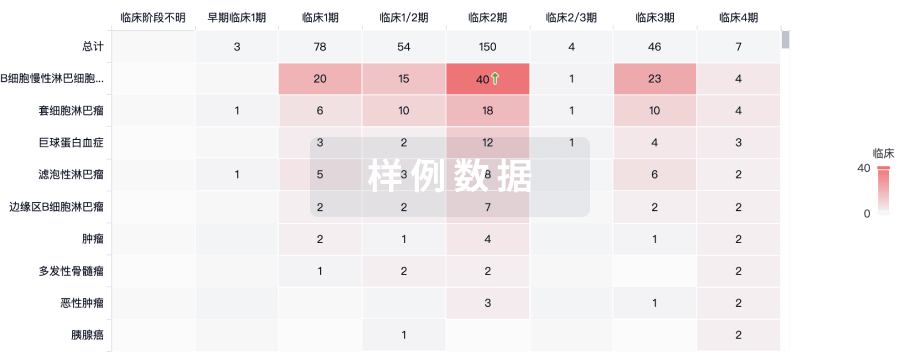
批准
利用最新的监管批准信息加速您的研究。
登录
或
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用


