预约演示
更新于:2025-05-07
Medical University of Ilam
更新于:2025-05-07
概览
关联
97
项与 Medical University of Ilam 相关的临床试验IRCT20211110053030N7
Investigating the effect of marital communication skills education on couple conflicts and marriage performance in infertile couples referring to health centers in Ilam 2025
开始日期2025-01-04 |
申办/合作机构 |
IRCT20241126063862N1
Investigating the effect of education based on the health belief model on attitudes toward childbearing in married couples
开始日期2024-12-21 |
申办/合作机构 |
IRCT20241126063859N1
Comparison of the effects of two herbal mouthwashes, one made from pistachio fruit and the other chlorhexidine, on gum healing after crown lengthening surgery.
开始日期2024-12-18 |
申办/合作机构 |
100 项与 Medical University of Ilam 相关的临床结果
登录后查看更多信息
0 项与 Medical University of Ilam 相关的专利(医药)
登录后查看更多信息
373
项与 Medical University of Ilam 相关的文献(医药)2023-08-01·Virus Genes
Immune response induced by recombinant pres2/S-protein and a pres2-S-protein fused with a core 18-27 antigen fragment of hepatitis B virus compared to conventional HBV vaccine
Article
作者: Khosravi, Afra ; Amani, Jafar ; Behzadi, Elham ; Imani Fooladi, Abbas Ali ; Parizad, Elaheh Gholami ; Sedighian, Hamid
2023-07-01·Journal of the Iranian Chemical Society
Efficient sonocatalytic degradation of orange II dye and real textile wastewater using peroxymonosulfate activated with a novel heterogeneous TiO2–FeZn bimetallic nanocatalyst
作者: Jaafarzadeh, Neemat ; Amarloei, Ali ; Noorimotlagh, Zahra ; Mirzaee, Seyyed Abbas ; Martinez, Susana Silva ; Dehvari, Mahboobeh
2023-07-01·Molecular biology reports
Nosocomial infections and antimicrobial susceptibility patterns among patients admitted to intensive care unit of Imam Khomeini hospital in Ilam, Iran.
Article
作者: Hashemian, Marzieh ; Bagheri, Mohammad Reza ; Kazemian, Hossein ; Kaviar, Vahab Hassan ; Khoshnood, Saeed ; Karamolahi, Somayeh ; Nazari, Ali ; Sadeghifard, Nourkhoda
100 项与 Medical University of Ilam 相关的药物交易
登录后查看更多信息
100 项与 Medical University of Ilam 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年06月10日管线快照
无数据报导
登录后保持更新
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
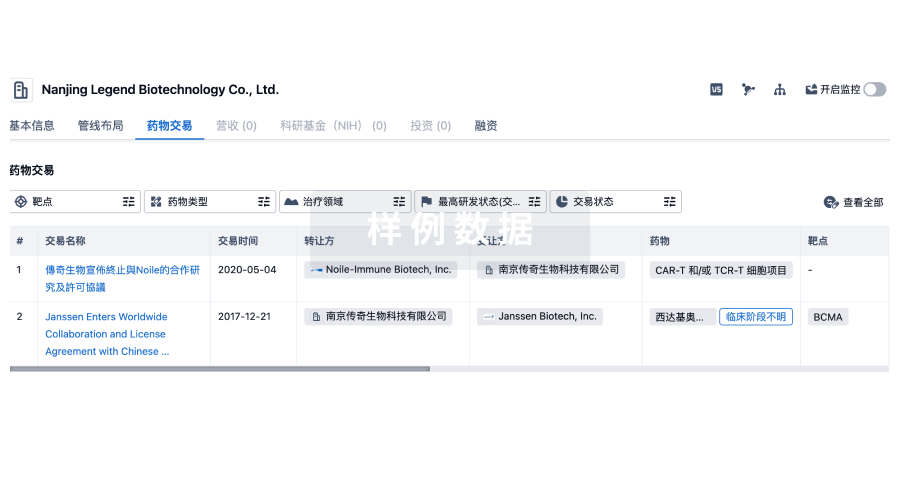
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
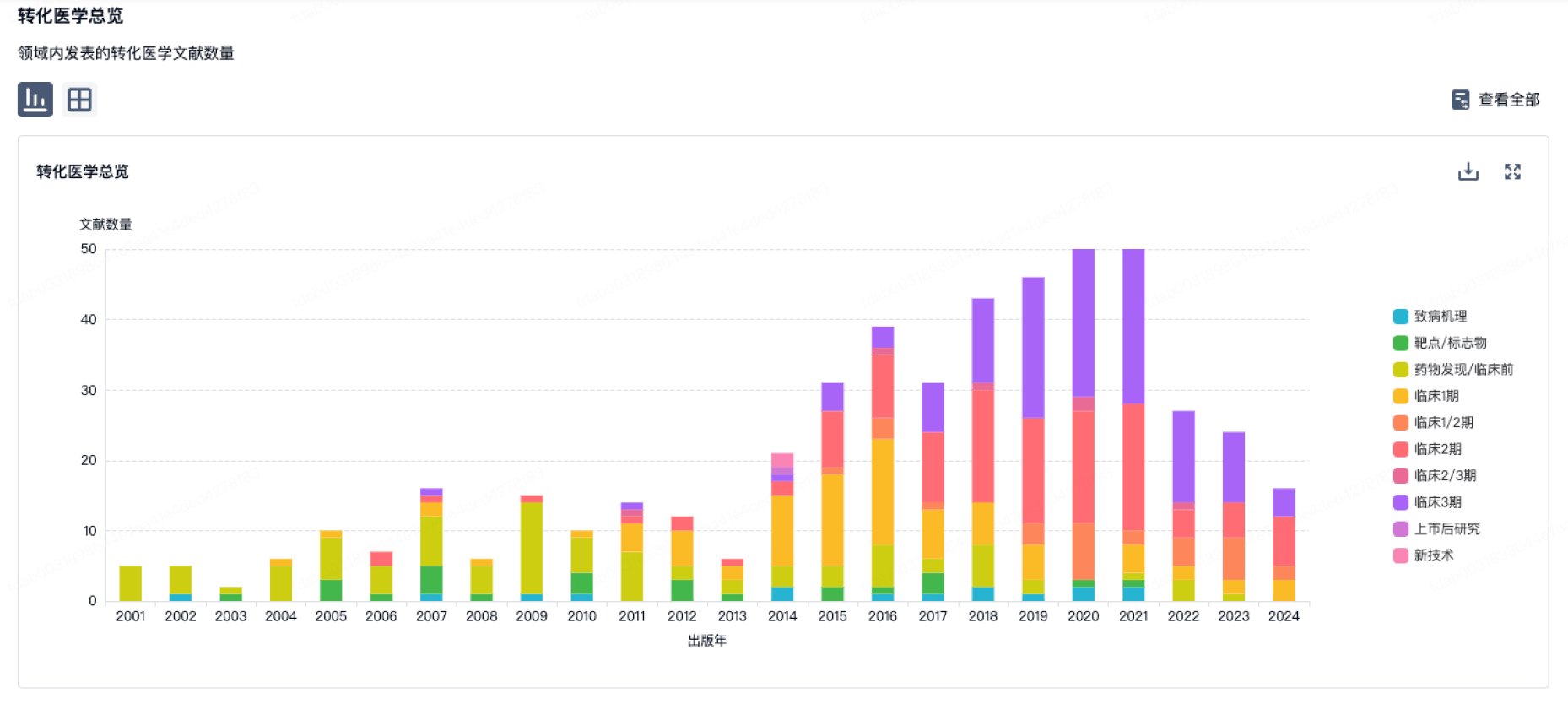
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
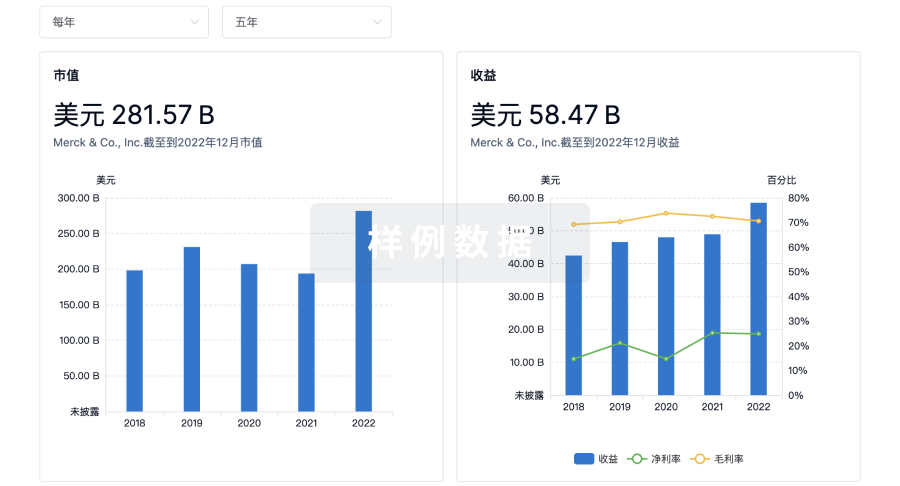
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
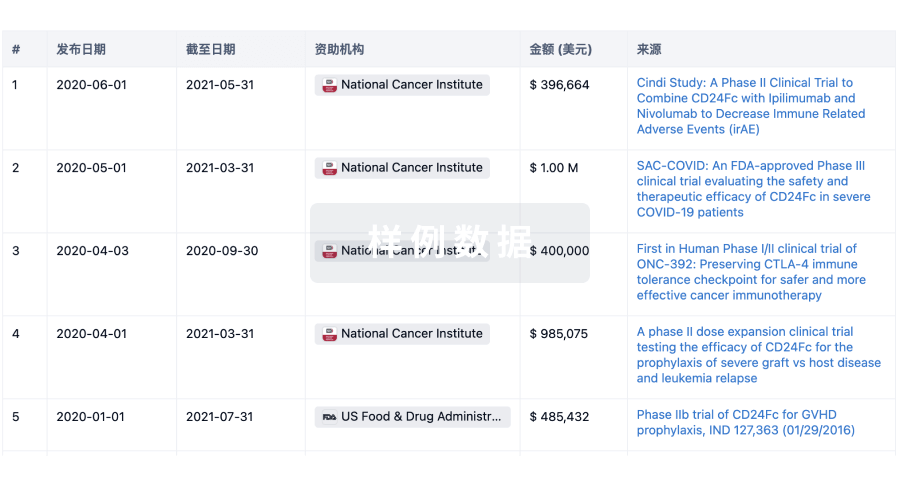
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
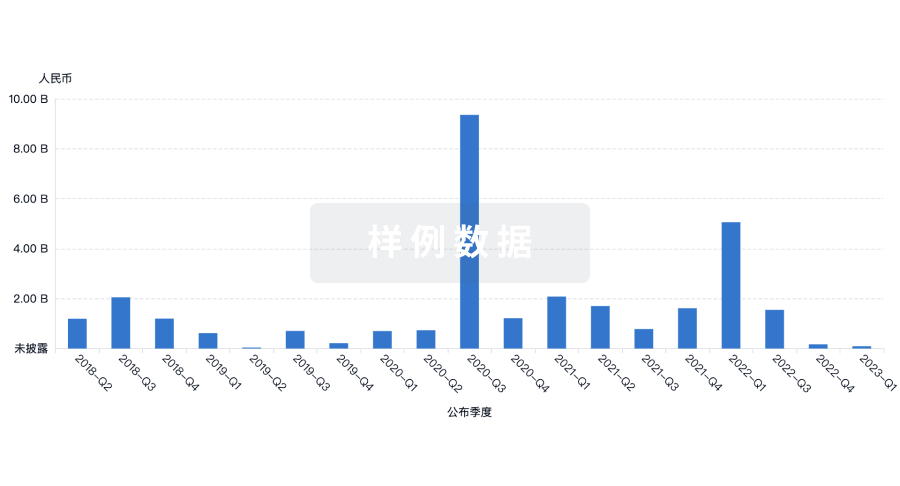
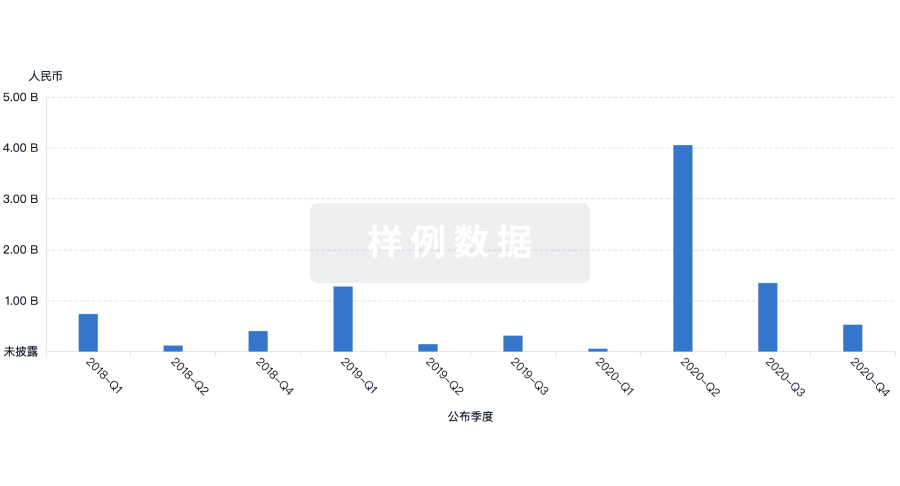
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用