预约演示
更新于:2026-02-08

Fakultní nemocnice Hradec Králové
更新于:2026-02-08
概览
关联
7
项与 Fakultní nemocnice Hradec Králové 相关的临床试验CTIS2023-505788-35-00
Complex immunomodulatory therapy for patients with immune thrombocytopenia (T-MEM)
开始日期2025-06-11 |
CTIS2023-509261-20-00
Modulation of the Extent of Ischemic Brain Damage by Protecting the Endothelial Glycocalyx with Low-Dose Albumin Administration (MOZEG)
开始日期2025-03-03 |
EUCTR2021-002014-14-CZ
Antibody and cellular response 28 days after the first dose of COVID-19 vaccine (Moderna) and then 33 weeks after the second dose and 8-12 weeks after the third dose in clinically stable patients with functional kidney transplantation.
开始日期2021-04-16 |
100 项与 Fakultní nemocnice Hradec Králové 相关的临床结果
登录后查看更多信息
0 项与 Fakultní nemocnice Hradec Králové 相关的专利(医药)
登录后查看更多信息
148
项与 Fakultní nemocnice Hradec Králové 相关的文献(医药)2026-01-27·ACS Omega
Targeting
Mycobacterium tuberculosis
: The Role of Alkyl Substitution in Pyrazinamide Derivatives
Article
作者: Juhás, Martin ; Paterová, Pavla ; Bárta, Pavel ; Bouz, Ghada ; Kučerová-Chlupáčová, Marta ; Zitko, Jan ; Weeks, Stephen D. ; Konečná, Klára ; Halířová, Martina ; Doležal, Martin ; Pang, Luping ; Jand́ourek, Ondřej
Tuberculosis (TB) remains a significant global health challenge due to the rapid emergence of drug resistance. Despite substantial progress in anti-TB drug development, effective treatment options are limited. In this study, we report the synthesis and biological evaluation of pyrazinamide (PZA) derivatives with 5-alkyl and 5-alkanamido modifications, designed to enhance antimycobacterial activity by increasing lipophilicity and improving penetration of the lipid-rich mycobacterial cell wall. A positive correlation between the length of the 5-alkyl chain and antimycobacterial activity was observed, with maximal potency achieved with the heptyl substituent (4: 5-heptylpyrazine-2-carboxamide, MIC_M. tuberculosis H37Rv = 3.13 μg/mL). In series C with phenyl substitution on the C-2 carboxamide, different simple substituents were tolerated on the benzene ring (both electron-donating and electron-withdrawing, both lipophilic and hydrophilic), and the length of the alkyl chain was the main determinant of the antimycobacterial activity. Compound 23 (5-hexyl-N-(3-trifluoromethylphenyl)-pyrazine-2-carboxamide) exerted MIC = 3.13 μg/mL and selectivity index (SI, compared to HepG2 cells) >25. Notably, the tested compounds exhibited significant activity against multidrug-resistant (MDR) Mycobacterium tuberculosis strains while maintaining favorable selectivity profiles and low cytotoxicity. In contrast, 5-alkanamido derivatives (series B and D) were devoid of antimycobacterial activity. Mechanistic investigations revealed that unlike PZA, the 5-alkyl pyrazinamide derivatives are not hydrolyzed by mycobacterial pyrazinamidase (PncA), indicating a distinct mode of action. While molecular modeling initially suggested enoyl-ACP reductase (InhA) as a potential target of series C, subsequent experimental validation disproved this hypothesis; thus, the precise mechanism of action remains to be elucidated.
2025-12-31·Journal of Maternal-Fetal & Neonatal Medicine
Clinical utility of a glycosylated fibronectin test (Lumella
TM
) for assessment of impending preeclampsia
Article
作者: Jorgensen, Jan Stener ; Wielgos, Miroslaw ; Gravett, Michael G. ; Di Renzo, Gian Carlo ; Stanirowski, Pawel ; Froeliger, Alizee ; Bartha, Jose Luis ; Gao, Lina ; Arduini, Maurizio ; Kacerovsky, Marian
OBJECTIVE:
Preeclampsia is a major pregnancy complication that results in significant maternal and infant mortality and morbidity, yet difficulties remain in the diagnosis of preeclampsia based on clinical parameters alone. The objective was to assess the performance of a hand-held point-of-care (POC) immunoassay in a clinical environment for glycosylated fibronectin (GlyFn) for the prediction of preeclampsia within 4 weeks of sampling.
METHODS:
Multinational European prospective observational pilot study of predominantly high-risk patients in the second half of pregnancy to assess a point-of-care immunoassay for GlyFn in predicting preeclampsia within 4 weeks of sampling. GlyFn was measured using a second generation hand held POC immunoassay. Results were considered normal for GlyFn concentrations of < 350 µg/mL, positive for GlyFn concentrations of 351-600 µg/mL, and high-positive for GlyFn concentrations > 600 µg/mL.
RESULTS:
Preeclampsia developed in 16 (19%) of 84 subjects and was associated with a shorter gestational age at delivery 35.3 weeks vs. 37.3 weeks for non-preeclamptics, n = 82; p = 0.001), a higher risk of fetal growth restriction (FGR; 31.2% vs. 10.3% for non-preeclamptics, p = 0.046), and an increased risk of preterm birth < 37 weeks gestation (83.3% vs. 33.3% for non-preeclamptics, (n = 78; p = 0.003). GlyFn positive or high positive was seen in 13/16 (81%) and in 35/68 (51.5%), yielding a sensitivity of 81%, a specificity of 49%, a positive predictive value of 27%, and a negative predictive value of 92%. GlyFn positive or high positive was also associated with preterm birth < 37 weeks in singleton pregnancy non-preeclamptic patients. Preterm birth occurred in 4.8% of those with normal GlyFn, in 26.7% with positive GlyFn, and in 50% of those with high GlyFn in singleton gestations without preeclampsia (p = 0.008).
CONCLUSION:
The ability to use this test in a POC format provides a method for practitioners to quickly determine risk for preeclampsia in their pregnant patients and offers an affordable alternative, as a single analyte to other diagnostic or screening tests that require laboratory-based testing or ultrasound equipment. Independent of preeclampsia, an elevated GlyFn was also correlated with preterm delivery and requires further study.
2025-12-31·PERSOONIA
Phylogeny, taxonomy and geographic distribution of novel and known fungi with holoblastic-denticulate conidiogenesis in
Rhamphoriales
and
Pleurotheciales
(
Sordariomycetes
)
Article
作者: Nekvindová, J. ; Hradilová, M. ; Kolařik, M. ; Réblová, M. ; Hernández-Restrepo, M.
As part of a broader survey of lignicolous saprobic fungi, we investigated fungal taxa from the class
Sordariomycetes
displaying holoblastic-denticulate conidiogenesis, a distinct developmental process and phylogenetically informative trait. Although these fungi appear morphologically
similar in culture, they represent distinct evolutionary lineages. This taxonomic study integrates comparative morphological analyses, phylogenetic to introduce novel taxa in the
Pleurotheciales
and
Rhamphoriales
. A new genus and species
Echinodenticula allantospora
and
three new species,
Phaeoisaria parallela
,
Rhamphoriopsis cuprea
and
Rh. denticulata
, are described. A rarely encountered species
Rhamphoria separata
is reported, along with its previously undocumented asexual morph. Furthermore, we successfully demonstrate the utility
of two protein-coding genes,
rpb2
and
tef1
, as complementary barcodes for distinguishing closely related
Phaeoisaria Rhamphoriales
and a prevalent trait in the
Pleurotheciales
. An unknown ascomycete that produced only sterile mycelium in culture is described here
as
Melanocrypta curvata
and placed at an incertae sedis position within the
Sordariomycetes
. Additionally, we present short-read whole-genome sequencing data for the ex-type strains of the newly described species, providing a valuable genomic resource for future taxonomic, phylogenetic,
and functional studies. Environmental DNA data from the GlobalFungi database bring new perspective into the biogeographical patterns of
Phaeoisaria
,
Rhamphoria
, and
Rhamphoriopsis
. The distribution of
E. allantospora
and
M. curvata
remains poorly understood,
as no records for these species were found in GlobalFungi. This study provides new insights into the molecular systematics, taxonomy, and biogeography of the
Rhamphoriales
and
Pleurotheciales
, and highlights the role of environmental DNA metabarcoding in uncovering fungal diversity
and distribution patterns.
100 项与 Fakultní nemocnice Hradec Králové 相关的药物交易
登录后查看更多信息
100 项与 Fakultní nemocnice Hradec Králové 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年03月18日管线快照
无数据报导
登录后保持更新
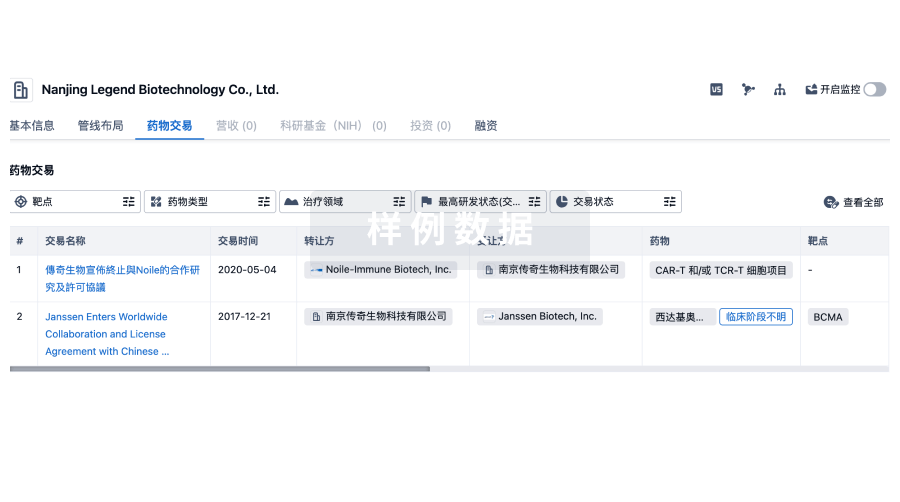
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
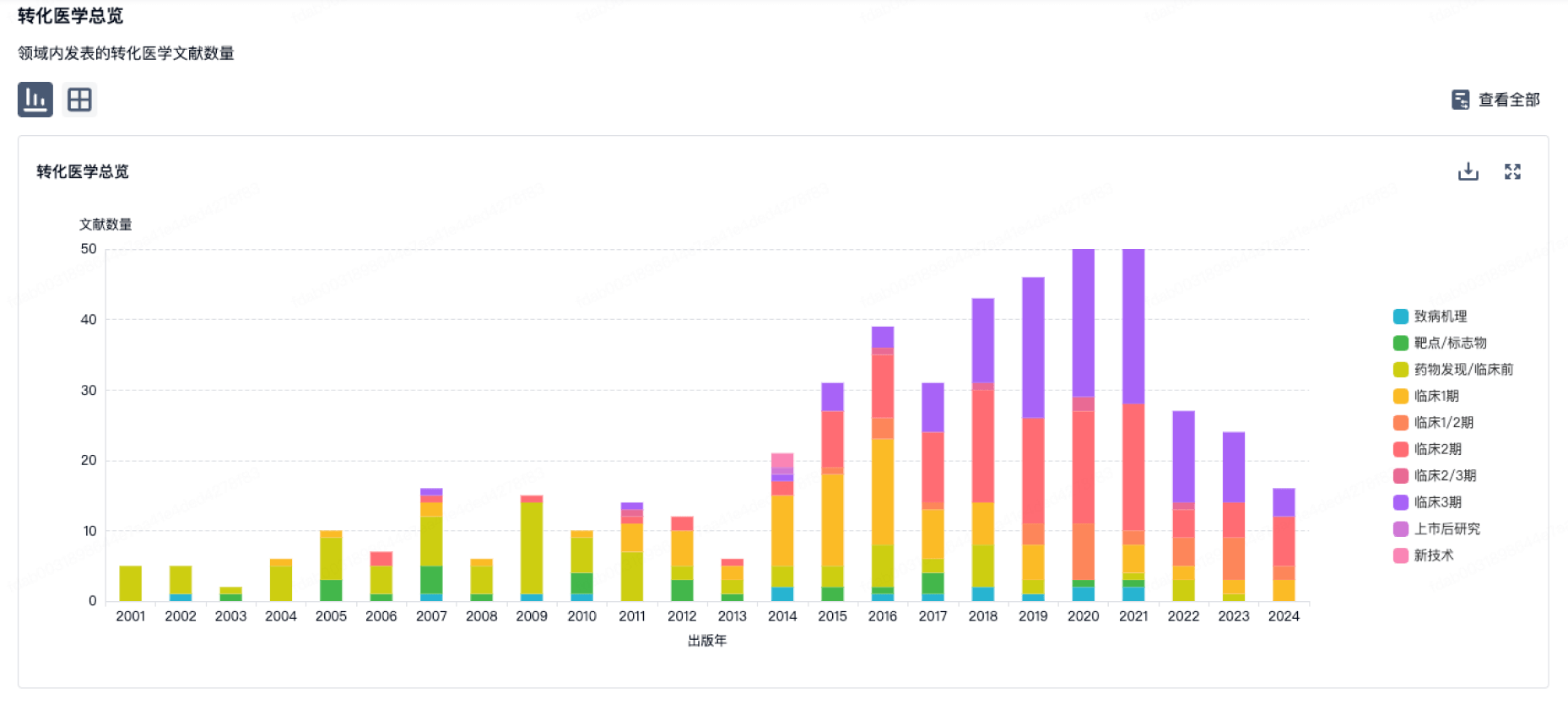
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
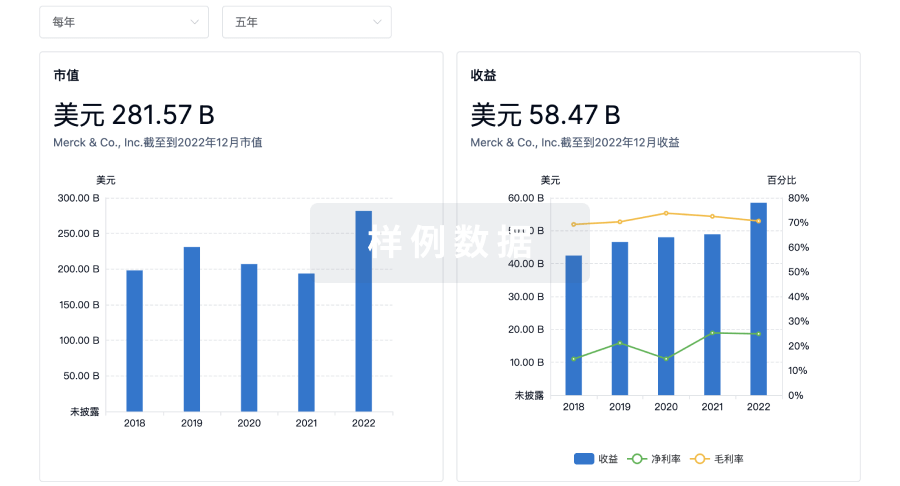
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
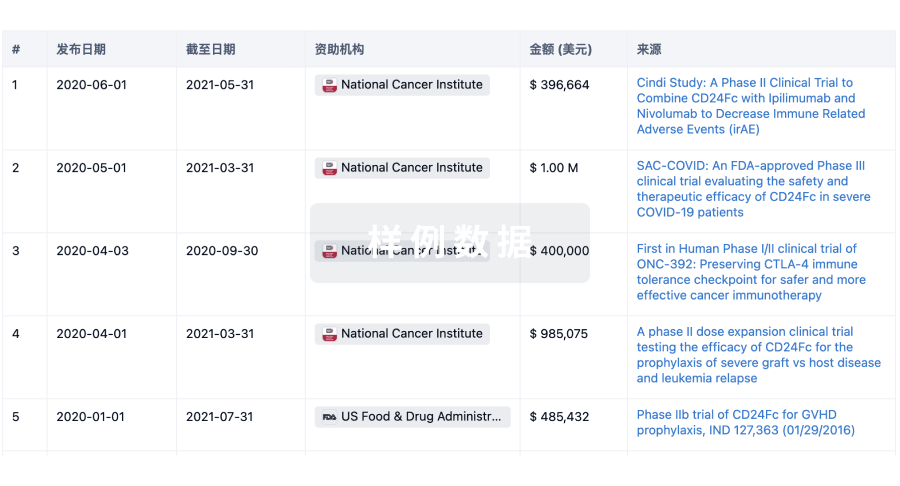
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
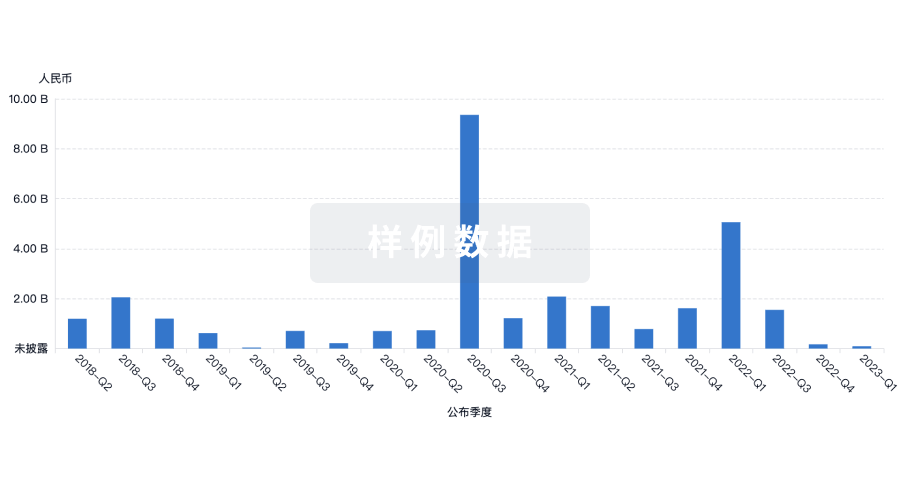
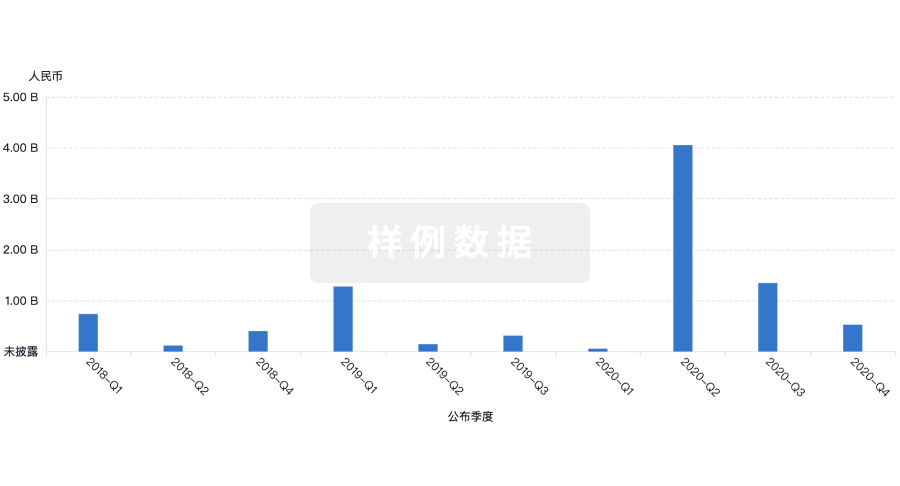
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用