预约演示
更新于:2025-05-07
PODXL
更新于:2025-05-07
基本信息
别名 GCTM-2 antigen、gp135、Gp200 + [10] |
简介 Involved in the regulation of both adhesion and cell morphology and cancer progression. Functions as an anti-adhesive molecule that maintains an open filtration pathway between neighboring foot processes in the podocyte by charge repulsion. Acts as a pro-adhesive molecule, enhancing the adherence of cells to immobilized ligands, increasing the rate of migration and cell-cell contacts in an integrin-dependent manner. Induces the formation of apical actin-dependent microvilli. Involved in the formation of a preapical plasma membrane subdomain to set up initial epithelial polarization and the apical lumen formation during renal tubulogenesis. Plays a role in cancer development and aggressiveness by inducing cell migration and invasion through its interaction with the actin-binding protein EZR. Affects EZR-dependent signaling events, leading to increased activities of the MAPK and PI3K pathways in cancer cells. |
关联
4
项与 PODXL 相关的药物靶点 |
作用机制 PODXL抑制剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期1800-01-20 |
靶点 |
作用机制 PODXL抑制剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期1800-01-20 |
靶点 |
作用机制 PODXL抑制剂 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期1800-01-20 |
100 项与 PODXL 相关的临床结果
登录后查看更多信息
100 项与 PODXL 相关的转化医学
登录后查看更多信息
0 项与 PODXL 相关的专利(医药)
登录后查看更多信息
1,001
项与 PODXL 相关的文献(医药)2025-04-01·Biological Trace Element Research
Zinc Deficiency Causes Glomerulosclerosis and Renal Interstitial Fibrosis Through Oxidative Stress and Increased Lactate Metabolism in Rats
Article
作者: Zhang, Qianyu ; Ye, Shang ; Li, Ningxu ; Liu, Xiaoxin ; Liao, Yajie ; Zheng, Yunxi ; Yu, Xiaohong ; Huang, Zixuan
2025-04-01·Journal of traditional Chinese medicine = Chung i tsa chih ying wen pan
Yishen Tongluo formula ameliorates kidney injury modulating inflammation and apoptosis in streptozotocin-induced diabetic kidney disease mice.
Article
作者: Yufan, Meng ; Jiangyan, X U ; Genlin, L I ; Jiayao, Yuan ; Suhui, W U ; Hanbing, L I
2025-03-24·Protein & Peptide Letters
Unveiling the Potential Role of Hesperetin and Emodin as a Combination Therapy to Inhibit the Pancreatic Cancer Progression against the C-Met
Gene
Article
作者: Kaviyaprabha, Rangaraj ; Muthusami, Sridhar ; Miji, Thandaserry Vasudevan ; Arulselvan, Palanisamy ; Bharathi, Muruganantham ; Sreelakshmi, Puthupparambil Shaji
17
项与 PODXL 相关的新闻(医药)2025-03-28
OTC2025论坛深度聚焦类器官与疾病建模、新药发现/研发、3D细胞培养、类器官培养及质控。展位咨询请联系:王晨 180 1628 8769。一、类器官研究:国自然申请的“黄金赛道”近年来,类器官技术以“黑马”姿态席卷科研界,成为国自然申请的焦点领域。据统计,2024年PubMed上“Organoid”相关论文突破3000篇,较2023年增长40%,技术迭代速度惊人。国自然基金委连续六年将类器官列为优先资助方向,2024年资助金额超4000万元,项目中标率达15.7%,创历史新高。类器官研究已从“前沿探索”跃升为“战略必争”,成为攻克临床难题、推动转化医学的核心工具。政策红利与技术迭代的双重驱动类器官的爆发式增长不仅是技术突破的结果,更是国家战略布局的体现。2024年《“十四五”生物经济发展规划》明确提出,将类器官列为“颠覆性生物技术”重点支持方向,旨在通过该技术实现疾病模型、药物筛选和再生医学的跨越式发展。此外,国自然基金委在《2025年项目指南》中首次提出“类器官与器官芯片整合计划”,鼓励跨学科团队联合攻关。这种政策倾斜不仅为科研人员提供了资金支持,更释放出明确的信号:类器官研究需从单一技术开发转向系统化平台构建,强调临床转化与产业化落地。值得注意的是,2024年国际竞争格局也在悄然变化。美国NIH启动“Human Biomimetic Organ Initiative (HBOI)”,计划投入2亿美元推动类器官标准化与数据共享;欧盟则通过“Organoid Innovation Hub”促进产学研合作。中国若要在这一领域保持领先,必须加强原创技术研发(如血管化类器官、动态微环境模拟)和伦理规范制定,避免陷入“跟跑”困境。二、类器官技术:从“模型革命”到“临床破局”技术优势:精准医学的“体外器官”类器官凭借三维结构、遗传稳定性和高度仿生性,突破传统模型的局限。以肿瘤研究为例,2024年《Cell》子刊研究显示,类器官对化疗药物敏感性的预测准确率达92%,远超动物模型(65%)。其核心优势在于:1. 个体化诊疗平台:利用患者来源类器官筛选个性化治疗方案,助力精准医疗;2. 疾病机制“解码器”:如2025年Nature首篇类器官论文中,团队通过脑类器官揭示阿尔茨海默病中tau蛋白的跨神经元传播机制,颠覆传统认知;3. AI赋能的高通量筛选:结合机器学习算法,类器官可同时评估数千种化合物,效率提升10倍以上,成本降低60%。类器官的“技术天花板”与突破路径尽管类器官技术优势显著,但其局限性不容忽视。例如,现有类器官多缺乏功能性血管系统,难以模拟器官间相互作用;长期培养中遗传漂变问题尚未完全解决。2024年,哈佛大学团队在《Science》发表突破性成果,通过3D生物打印技术将内皮细胞嵌入肝类器官,成功构建血管网络,并实现类器官移植后的小鼠体内存活。这一进展提示,未来类器官研究需聚焦“复杂性”与“功能性”双重提升,结合材料科学、工程学等多学科技术,突破现有技术瓶颈。此外,伦理争议仍是类器官发展的潜在风险。2024年,日本学者利用人脑类器官模拟意识活动的实验引发广泛讨论。对此,国自然基金委在资助项目中明确要求:涉及神经类器官的研究需通过伦理审查,并遵循“14天规则”(即类器官发育不得超过14天)。科研人员需在技术创新与伦理约束间找到平衡点。政策导向:国自然聚焦四大热点2024年国自然类器官资助项目呈现三大趋势:1. 临床痛点导向:如耐药机制、罕见病建模、器官再生;2. 技术融合创新:类器官芯片、单细胞空间组学、光遗传操控;3. 转化应用落地:类器官指导临床试验(如FDA已批准3项类器官辅助的肿瘤新药试验);4. 伦理规范化:基金委要求类器官研究需符合《生物医学新技术临床研究伦理审查指南》。从“单点突破”到“生态构建”国自然的资助导向已从支持单一技术开发,转向构建“类器官研究生态”。例如,2024年重点专项“类器官标准化与资源共享平台”要求建立全国统一的类器官样本库、数据共享平台和质量评价体系。这一转变意味着,科研团队需具备“平台化思维”,既能解决具体科学问题,又能贡献于行业基础设施建设。例如,上海瑞金医院联合中科院搭建的“类器官临床转化联盟”,已实现3000例肿瘤类器官数据共享,为全国研究者提供标准化模型和比对基准。三、类器官研究“五步法”:国自然中标模板锚定临床“卡脖子”问题以近期热点的“肿瘤免疫治疗耐药”为例,PD-1抑制剂在实体瘤中有效率不足30%,且耐药机制不明。申请者可聚焦“肿瘤类器官-免疫细胞共培养模型”,解析微环境中T细胞耗竭的关键信号通路,直击临床痛点。当前许多类器官研究仍停留在表型观察阶段,缺乏机制深度。以肿瘤耐药研究为例,多数论文仅报道类器官对药物的反应差异,却未揭示耐药基因的调控网络。2024年,斯坦福大学团队在《Cell》提出“耐药进化树”概念,通过连续传代类器官模拟肿瘤耐药演化过程,并结合单细胞测序追踪耐药克隆的基因突变轨迹。此类研究设计不仅阐明机制,更为逆转耐药提供了动态靶点,值得借鉴。构建高质量类器官资源库2024年中科院团队发表于《Science Translational Medicine》的研究提供范本:1. 样本多样性:涵盖不同分期、分子分型及治疗反应的肿瘤组织;2. 标准化流程:采用自动化类器官培养系统(如OrganoPlate®),确保批次稳定性;3. 多模态验证:通过单细胞测序、空间转录组和活细胞成像,证实类器官与原发肿瘤的一致性(相似度>90%)。类器官库可分为“静态库”(仅保存类器官)和“动态库”(整合患者临床数据与多组学信息)。2024年,北京协和医院首创“伴随诊断型类器官库”,在培养类器官的同时,持续收集患者治疗反应数据,并通过AI模型预测药物敏感性。这种“活库”不仅能服务基础研究,还可直接指导临床用药,显著提升资源库的附加值。多组学驱动靶点挖掘典型案例:复旦大学团队利用类器官转录组-表观组联合分析,发现结直肠癌中LIN28B通过调控糖代谢重编程诱导耐药,相关成果入选2024年“中国生命科学十大进展”。多组学分析易陷入“数据沼泽”,即产出海量数据却缺乏生物学意义。2024年《Nature Methods》提出“靶点挖掘三原则”:1. 临床相关性:靶点需在独立临床队列中验证;2. 进化保守性:靶点应在不同物种或疾病模型中保守;3. 可干预性:靶点需具备成药潜力或基因编辑可行性。例如,浙江大学团队在肝癌类器官中发现Wnt/β-catenin通路异常激活,进一步通过跨物种分析(类器官-小鼠-斑马鱼)证实其保守性,并设计小分子抑制剂成功逆转耐药。临床队列验证与AI预测需整合大规模临床样本(如≥200例)和AI模型。例如,北大团队开发DeepOrganoid系统,通过类器官影像特征预测患者5年生存率(AUC=0.87),显著提升研究转化价值。尽管AI在类器官数据分析中表现优异,但其“黑箱”特性常被诟病。2024年,MIT团队提出“双通道验证法”:在AI预测后,通过类器官基因编辑实验反向验证关键特征。例如,若AI判定类器官形态不规则与预后不良相关,则通过CRISPR修复相关基因,观察形态是否恢复正常。这种“假设-验证”闭环极大提升了AI模型的可信度。机制解析与干预策略运用CRISPR筛选、类器官移植模型(如PDX-Organoid)等验证靶点功能。2025年《Nature Medicine》研究通过基因编辑修复囊性纤维化类器官的CFTR突变,并利用mRNA疗法恢复离子通道功能,为临床提供全新方案。传统机制研究多聚焦于“纠正异常”,而最新趋势是“功能重建”。例如,剑桥大学团队在胰腺类器官中引入合成生物学线路,使其在感知高血糖时自动分泌胰岛素。这种“智能类器官”不仅揭示了疾病机制,更直接转化为治疗手段,代表未来研究的核心方向。四、2024年高分案例:国自然“标杆式”设计案例1:类器官芯片破解肿瘤异质性清华团队在《Nature Biotechnology》发表研究,将肺癌类器官与微流控芯片结合,模拟血管化肿瘤微环境,发现缺氧诱导的ANGPTL4蛋白促进转移。该研究获2024年国自然重点专项资助,关键点在于:1. 技术创新:首创“器官芯片+单细胞代谢组学”平台;2. 临床关联:ANGPTL4抑制剂已进入Ⅰ期临床试验。类器官芯片的价值不仅在于模拟复杂微环境,更在于实现“动态干预”。例如,2024年荷兰Hubrecht研究所开发“光控类器官芯片”,通过光遗传学手段实时调控特定信号通路,直接观察类器官响应。这种技术将机制研究从“静态关联”升级为“因果论证”,极大提升论文说服力。案例2:脑类器官揭示自闭症新机制上海交大团队利用患者iPSC构建脑类器官,结合空间蛋白组技术,发现星形胶质细胞分泌的SPARCL1异常导致突触修剪失衡,成果登上《Cell》并获杰青项目支持。该研究的突破性在于,团队并未止步于机制解析,而是进一步筛选出可上调SPARCL1的小分子化合物,并在自闭症患儿类器官中验证疗效。这种“一站式”研究模式(机制-靶点-药物)完美契合国自然“转化优先”导向,值得效仿。五、申请书撰写“三大黄金法则”创新性:直击基金委“兴趣点”1. 标题公式:前沿技术+临床场景+科学问题,如《基于时空组学的肝癌类器官免疫逃逸机制研究》;2. 摘要策略:用数据凸显创新,如“首次建立类器官动态缺氧模型,解析HIF-1α/PD-L1轴在耐药中的功能”。国自然评审专家期待“渐进式创新”,而非“空中楼阁”。建议采用“三级跳”策略:技术层面:改进现有类器官培养方法(如添加机械应力模拟心脏搏动);方法层面:整合多学科技术(如类器官+空间代谢组);理论层面:提出新机制或新范式(如“类器官免疫编辑假说”)。例如,中山大学2024年获批项目《力学微环境调控的肝类器官纤维化逆转机制》,即通过三级创新设计脱颖而出。可行性:细节决定成败1. 技术路线图:细化类器官培养参数(如基质胶浓度、氧梯度设置);2. 团队背书:突出多学科合作(如临床医生+生物信息学家);3. 风险预案:针对类器官血管化难题,预实验已优化VEGF梯度灌注方案。顶层设计需要明确研究框架与核心问题;中层支撑需要展示预实验结果、合作单位资源;底层保障需要详述设备、样本、经费的具体分配。例如,华中科技大学团队在申请书中附上预实验的类器官成像数据、合作医院的样本收集协议,以及超算中心的数据处理协议,形成完整证据链。科学意义:呼应国家战略1. 临床价值:如“建立类器官指导的胃癌精准治疗体系,缩短新药研发周期2年以上”;2. 学科贡献:推动类器官标准化(参照2024年ISO发布的《类器官质量评价指南》)。将“学科发展”与“社会效益”螺旋式结合。例如:“本研究通过建立帕金森病类器官模型,不仅揭示α-synuclein蛋白传播机制(学科贡献),还将联合药企开发神经保护剂,预计惠及全球1000万患者(社会效益)。”六、答辩制胜:从“答”到“辨”的升华1. PPT设计:用动态视频展示类器官药物响应过程,取代静态图表;2. 应答策略:针对“模型局限性”质疑,可回应:“本研究同步建立患者源性异种移植(PDOX)模型,双模型验证确保结论可靠性”。首先用1分钟视频展示研究亮点(如类器官动态响应);随后聚焦核心数据(如靶点验证的生存曲线);最后以临床转化蓝图收尾(如“预计3年内启动临床试验”)。这种结构化表达能牢牢抓住评委注意力。七、结语类器官研究正处于“技术爆发”与“政策红利”的双重窗口期。研究者需紧扣国自然“需求导向、交叉融合、转化优先”的资助逻辑,以临床问题为锚点,以技术创新为引擎,方能在激烈竞争中脱颖而出。2025年国自然申请将结束,抓紧最后的时间完善申请书的内容,唯有深耕技术、精研策略,方能以类器官为支点,撬动生物医学研究的未来!附:2024年类器官领域国自然中标项目名单:END免责声明:本文仅作知识交流与分享及科普目的,不涉及商业宣传,不作为相关医疗指导或用药建议。文章如有侵权请联系删除。OTC2025论坛深度聚焦类器官与疾病建模、新药发现/研发、3D细胞培养、类器官培养及质控。展位咨询请联系:王晨 180 1628 8769。
2025-02-06
2025年2月5日,中山大学附属第三医院的研究团队在期刊《Cell Death Discovery》上发表了题为“TMEM45A enhances palbociclib resistance and cellular glycolysis by activating AKT/mTOR signaling pathway in HR+ breast cancer”的研究论文。研究结果表明,TMEM45A/AKT/mTOR轴通过促进BRCA细胞中的糖酵解和上皮-间质转化(EMT),成为乳腺癌帕博西尼耐药性的关键介导因子,凸显了其作为治疗靶点的潜力。
2025年2月5日,中山大学附属第三医院的研究团队在期刊《Cell Death Discovery》上发表了题为“TMEM45A enhances palbociclib resistance and cellular glycolysis by activating AKT/mTOR signaling pathway in HR+ breast cancer”的研究论文。研究结果表明,
TMEM45A/AKT/mTOR轴通过促进BRCA细胞中的糖酵解和上皮-间质转化(EMT),成为乳腺癌帕博西尼耐药性的关键介导因子,凸显了其作为治疗靶点的潜力。
https://www.nature.com/articles/s41420-025-02336-9
https://www.nature.com/articles/s41420-025-02336-9
关于乳腺癌的耐药性
关于乳腺癌的耐药性
01
01
乳腺癌(BRCA)是女性最常见的恶性肿瘤,其中激素受体阳性(HR+)BRCA是最常见的亚型,约占所有乳腺癌的70%。CDK4/6抑制剂(CDK4/6i),包括palbociclib、ribociclib和abemaciclib,可以改善晚期HR+BRCA患者的无进展生存期(PFS)、总生存期(OS),延长化疗时间,提高生活质量。然而,尽管晚期疾病的治疗有了明显改善,但仍有相当一部分患者最终会对CDK4/6i产生耐药性。
乳腺癌(BRCA)是女性最常见的恶性肿瘤,其中激素受体阳性(HR+)BRCA是最常见的亚型,约占所有乳腺癌的70%。CDK4/6抑制剂(CDK4/6i),包括palbociclib、ribociclib和abemaciclib,可以改善晚期HR+BRCA患者的无进展生存期(PFS)、总生存期(OS),延长化疗时间,提高生活质量。然而,尽管晚期疾病的治疗有了明显改善,但仍有相当一部分患者最终会对CDK4/6i产生耐药性。
CDK4/6i治疗的抗药性与基因调控异常密切相关,这些分子可作为治疗靶点。递送小干扰RNA(siRNA)或非编码RNA,可能是靶向治疗耐药性的一种有前途的方法。研究表明,从癌症患者原代细胞中提取的外泌体可作为siRNA靶向递送的有效方法,提高化疗和免疫治疗药物的敏感性。因此,利用从患者肿瘤原代细胞中提取的外泌体作为siRNA载体,将治疗用siRNA运送到肿瘤部位是一种有效的方法。
CDK4/6i治疗的抗药性与基因调控异常密切相关,这些分子可作为治疗靶点。递送小干扰RNA(siRNA)或非编码RNA,可能是靶向治疗耐药性的一种有前途的方法。研究表明,从癌症患者原代细胞中提取的外泌体可作为siRNA靶向递送的有效方法,提高化疗和免疫治疗药物的敏感性。因此,利用从患者肿瘤原代细胞中提取的外泌体作为siRNA载体,将治疗用siRNA运送到肿瘤部位是一种有效的方法。
生物工程外泌体在PDX模型中的抗肿瘤功能
生物工程外泌体在PDX模型中的抗肿瘤功能
02
02
在PDX-pa小鼠中,单独使用palbociclib(150mg/kg)和单独使用Exo-siTMEM45A可抑制肿瘤生长,而联合使用Exo-siTMEM45A和palbociclib(150mg/kg)可显著抑制肿瘤大小。此外,减少帕博西来(palbociclib)的用量(50毫克/千克)与Exo-siTMEM45A合用也能有效抑制肿瘤生长。在对帕博西尼耐药的PDX-PalR小鼠模型中,帕博西尼单药治疗(150毫克/千克)并不能显著缩小肿瘤大小。然而,Exo-siTMEM45A和palbociclib(150毫克/千克)联合治疗,即使浓度较低(50毫克/千克),也能明显抑制肿瘤生长。这一结果表明,Exo-siTMEM45A和palbociclib的联合用药明显有利于肿瘤CDK4/6抑制剂的治疗,而且通过工程外泌体递送siTMEM45A可能会大大减少palbociclib的用量。IHC染色显示,Exo-siTMEM45A能降低PDX肿瘤组织中TMEM45A的水平,表明工程外泌体在体内具有良好的敲除效率。联合使用palbociclib和Exo-siTMEM45A还能抑制肿瘤组织中增殖标志物Ki67的表达。
在PDX-pa小鼠中,单独使用palbociclib(150mg/kg)和单独使用Exo-siTMEM45A可抑制肿瘤生长,而联合使用Exo-siTMEM45A和palbociclib(150mg/kg)可显著抑制肿瘤大小。此外,减少帕博西来(palbociclib)的用量(50毫克/千克)与Exo-siTMEM45A合用也能有效抑制肿瘤生长。在对帕博西尼耐药的PDX-PalR小鼠模型中,帕博西尼单药治疗(150毫克/千克)并不能显著缩小肿瘤大小。然而,Exo-siTMEM45A和palbociclib(150毫克/千克)联合治疗,即使浓度较低(50毫克/千克),也能明显抑制肿瘤生长。这一结果表明,
Exo-siTMEM45A和palbociclib的联合用药明显有利于肿瘤CDK4/6抑制剂的治疗,而且通过工程外泌体递送siTMEM45A可能会大大减少palbociclib的用量。
IHC染色显示,Exo-siTMEM45A能降低PDX肿瘤组织中TMEM45A的水平,表明工程外泌体在体内具有良好的敲除效率。
联合使用palbociclib和Exo-siTMEM45A还能抑制肿瘤组织中增殖标志物Ki67的表达。
工程外泌体在PDX模型中的肿瘤抑制作用。
工程外泌体在PDX模型中的肿瘤抑制作用。
总结
总结
03
03
1. TMEM45A作为潜在治疗靶点:研究建立了帕博西尼耐药细胞系,通过体外和体内实验验证了TMEM45A是克服帕博西尼耐药的潜在治疗靶点。
1. TMEM45A作为潜在治疗靶点:
研究建立了帕博西尼耐药细胞系,通过体外和体内实验验证了TMEM45A是克服帕博西尼耐药的潜在治疗靶点。
2. TMEM45A的生物学作用机制:TMEM45A通过激活AKT/mTOR信号通路促进糖酵解,介导帕博西尼耐药性。AKT/mTOR通路在CDK4/6抑制剂耐药中起关键作用。
2. TMEM45A的生物学作用机制:
TMEM45A通过激活AKT/mTOR信号通路促进糖酵解,介导帕博西尼耐药性。AKT/mTOR通路在CDK4/6抑制剂耐药中起关键作用。
3. 外泌体siRNA递送系统:基于患者来源的原代细胞构建的外泌体用于递送靶向TMEM45A的siRNA,提高了帕博西尼耐药PDX模型对帕博西尼的敏感性,表明TMEM45A是治疗BRCA帕博西尼耐药的有前景的靶点。
3. 外泌体siRNA递送系统:
基于患者来源的原代细胞构建的外泌体用于递送靶向TMEM45A的siRNA,提高了帕博西尼耐药PDX模型对帕博西尼的敏感性,表明TMEM45A是治疗BRCA帕博西尼耐药的有前景的靶点。
4. TMEM45A与肿瘤进展的关系:TMEM45A可抑制肿瘤进展中的细胞周期停滞和细胞凋亡,与肿瘤的迁移和侵袭能力相关。耐药细胞更依赖糖酵解,TMEM45A可能通过激活AKT/mTOR通路促进耐药和糖酵解。
4. TMEM45A与肿瘤进展的关系:
TMEM45A可抑制肿瘤进展中的细胞周期停滞和细胞凋亡,与肿瘤的迁移和侵袭能力相关。耐药细胞更依赖糖酵解,TMEM45A可能通过激活AKT/mTOR通路促进耐药和糖酵解。
5. 外泌体作为siRNA载体的优势:外泌体具有天然的生物相容性和低免疫原性,能保护siRNA不被核酸酶降解,是基因治疗的理想载体。研究利用患者肿瘤原代细胞提取的外泌体递送siRNA,特异性地减少了肿瘤细胞内TMEM45A的表达。
5. 外泌体作为siRNA载体的优势:
外泌体具有天然的生物相容性和低免疫原性,能保护siRNA不被核酸酶降解,是基因治疗的理想载体。研究利用患者肿瘤原代细胞提取的外泌体递送siRNA,特异性地减少了肿瘤细胞内TMEM45A的表达。
参考资料:
参考资料:
1.Hong R, Xu B. Breast cancer: an up-to-date review and future perspectives. Cancer Commun. 2022;42:913–36.
1.Hong R, Xu B. Breast cancer: an up-to-date review and future perspectives. Cancer Commun. 2022;42:913–36.
2.Beaver JA, Amiri-Kordestani L, Charlab R, Chen W, Palmby T, Tilley A, et al. FDA Approval: Palbociclib for the treatment of postmenopausal patients with estrogen receptor-positive, HER2-negative metastatic breast cancer. Clin Cancer Res. 2015;21:4760–6.
2.Beaver JA, Amiri-Kordestani L, Charlab R, Chen W, Palmby T, Tilley A, et al. FDA Approval: Palbociclib for the treatment of postmenopausal patients with estrogen receptor-positive, HER2-negative metastatic breast cancer. Clin Cancer Res. 2015;21:4760–6.
siRNA临床结果
2024-11-28
OTC2024类器官前沿应用与3D细胞培养论坛圆满落幕,点击图片可查看会后报告,咨询OTC2025类器官论坛请联系:王晨 180 1628 8769。
文章来源:华夏源类器官
“
人类肠道内分泌细胞(EEC)是一种罕见的胃肠道上皮细胞,由于低表达,多亚型,种间差异大,缺乏有效的研究模型。近日,Hans Clevers团队在顶级期刊《Science》(IF:44.7)发表研究论文[1],全面描述并验证了肠道内分泌细胞传感器(受体)功能,揭示其在调节激素分泌中的关键作用。研究中开发的胃类器官模型,补充了现有肠道类器官培养协议,有望为糖尿病、肥胖症等代谢疾病治疗提供新策略。
肠道是重要的消化器官,也是人体最大的免疫器官,具有营养吸收、能量代谢、保护屏障等重要生理作用。这道身体“长城”既可以保护身体免受有害细菌和高度动态PH值的侵害,又可以通过消化、吸收食物中的营养成分,使其进入血液,为身体提供能量。更重要的是,肠道还是分泌多种激素以调节身体机能的内分泌细胞的“家园”。
人类肠道内分泌细胞(EEC)是肠道上皮细胞的一个特殊亚群,相对稀有,所占数量不足整个胃肠道(GI)上皮细胞的1%。EEC作为专门的激素分泌细胞,可细分为五个主要亚型,且每种亚型产生不同的肽类激素和/或神经递质(如GLP-1和GIP等),激素的释放通过代谢物传感蛋白调控。
EEC通过分泌激素来响应肠道内容物,从而调节包括食欲、胰岛素释放和肠道运动等重要生理活动。尽管如此,由于EEC的稀有性,低表达,种间差异大(小鼠 EEC 响应的信号可能与人类 EECs响应的信号不同),亚型多样,以及对EEC的药理学研究多集中在治疗糖尿病和肥胖症等方面,目前鲜有实验模型能全面反映EEC在代谢调节中的重要作用,为深入研究带来挑战。
令人惊喜的是,近日由荷兰Hubrecht研究所和罗氏人类生物学研究所(Institute of Human Biology,IHB)领导的研究团队开发了全新的胃肠类器官模型,全面描述并验证了人类肠内分泌细胞(EEC)的传感器(受体)功能,揭示了其在调节激素分泌中的作用。该研究区分了人类胃EEC,并整合了胃和肠道类器官的研究协议,超越了以往主要基于小鼠模型或仅专注于肠道类器官的研究。
该实验阐明了个别人类EEC传感器在激素分泌中的重要作用,这些传感器有望成为影响食欲、肠道运动、胰岛素敏感性和黏膜免疫的潜在药物靶点,有望为治疗代谢疾病(如糖尿病和肥胖症等)提供全新的干预途径。
01
胃类器官EEC功能验证
开辟代谢疾病治疗新路径
肠道内分泌细胞(Enteroendocrine cells, EEC)是分布在胃、小肠和结肠上皮中的激素分泌细胞。EEC根据产生的肽类激素(或神经递质)不同被分为5个亚型。例如,L细胞产生胰高血糖素样肽1(Glucagon-like peptide 1, GLP-1),而K细胞产生胃抑制蛋白(Gastric inhibitory protein, GIP)。这些激素通过G蛋白偶联受体(G protein-coupled receptors, GPCR)以及营养状态来控制细胞内钙水平,进而调节激素的分泌。
由“类器官鼻祖”Hans Clevers教授带领的Hubrecht研究团队在此前已经开发出在类器官中获取大量EEC的方法。考虑到EEC在肠道的不同区域具有不同的传感蛋白和激素谱,研究这些稀有细胞需要制作所有这些不同区域的富集EEC的类器官。
在最新研究中,研究团队设法在消化系统的其他部分(包括胃)构建类器官模型。Hans Clevers团队利用类器官技术和CRISPR基因筛选,揭示了调控EEC分化的关键因子。首先,他们通过诱导神经元生成因子-3——NEUROG-3(一种对EEC分化至关重要的转录因子)的过量表达,实现胃EEC的生产,并成功分化出人类胃类器官中的关键EEC亚型。
这些肠道内分泌细胞在受到刺激时能够分泌大约20种活性激素,包括5-羟色胺(5-HT)、生长抑素(somatostatin, SST)、神经肽Y(neuropeptide Y, NPY)等,它们通过内分泌和旁分泌途径作用于远处或邻近的靶器官。
△ 人类胃器官,可以看到一种罕见的紫色生长激素释放肽产生细胞
与真实的胃部一样,这些胃类器官也会对已知的激素释放诱导剂做出反应,并分泌大量的胃饥饿素(Ghrelin),这种激素向大脑发出饥饿信号方面起着关键作用。
这证实了胃类器官可用于研究EEC的激素分泌,以此作为现有肠道类器官培养技术的补充。
△ 胃EEC生成的示意图
由于人类肠道内分泌细胞较为罕见,研究团队一直在努力绘制许多EEC的轮廓图。
在最新实验中,研究人员在EEC表面上发现了一种名为CD200的标志物,利用这一表面标志物从类器官中分离出大量人类EEC,并研究它们的传感器(受体)蛋白。
CD200表面标志物用于不同区域(胃、小肠和结肠组织)人类胃肠道(GI)EEC的深度转录组分析,克服了之前研究此类罕见细胞的困难。研究人员结合大规模RNA测序和蛋白质组分析,绘制了人类EEC的单细胞转录组图谱,并验证了类器官和组织中EEC的受体表达概况。
△ 胃、小肠和结肠中人类EEC亚型的免疫荧光染色
此外,研究人员利用CRISPR/Cas9基因编辑技术构建了一个包含22个受体缺陷类器官生物样本库。引入一种主要针对G蛋白偶联受体(GPCR)功能的新型筛选方式,研究这些受体缺失的类器官在不同配体刺激下的激素分泌情况(如GLP-1、5-羟色胺和生长激素释放肽GHRL)。例如,识别了多种控制GLP-1分泌的GPCR和通道蛋白,包括磺脲类药物受体ABCC8和色氨酸受体CasR。
△ 生成用于功能验证的人类功能丧失GPCR生物样本库的实验概述图
GPCR的功能验证提供了对这些受体如何影响肠道激素分泌的机制性理解,为开发能够激活这些传感器的新药物提供了机会,特别是通过靶向局部激素分泌(如GLP-1)作为当前系统性激素模拟物疗法的替代方案,可能引发小分子口服药物的开发,或将开辟代谢疾病(如糖尿病和肥胖症等)的治疗新思路。
综上,这些研究不仅加深了我们对EEC在代谢和免疫调节中作用的理解,还为开发新的治疗代谢性疾病的策略提供了重要基础。类器官体外平台已然成为连接基础科研与临床应用之间的桥梁。
02
医学科研革新工具
类器官技术大放异彩
肠道内分泌细胞(EEC)在调节消化、吸收、食欲和代谢等方面的关键作用,通过人类胃类器官体外模型的构建,补充了现有肠道类器官培养技术,为研究EEC的分化和功能提供了新工具。通过全面描述和验证人类EEC传感器(受体)功能,揭示其在调节激素分泌中的作用,为调控食欲、肠道运动、胰岛素敏感性和粘膜免疫提供了潜在药物靶点。
作为体内疾病的绝佳体外研究场所,类器官模型的构建已成为更多疾病治疗研究策略的新途径。
胃食管癌独特类器官模型
胃食管交界,是指食管远端和胃近端贲门交界处一段非常短的解剖学区域。胃食管交界处(GEJ)癌就是发生在这一区域的恶性肿瘤。临床上,GEJ癌既不属于胃癌也不属于食管癌范畴,治疗方案也因此差异很大。
2022年11月,来自南加利福尼亚大学等机构的科学家们通过研究利用人类组织开发了一种在实验室生长的三维类器官模型和CRISPR编辑转化的GEJ类器官模型。[2]该研究提供了对早期 GEJ 肿瘤中与TP53/CDKN2A失活相关的原发性机制的见解,确定了GEJ模型肿瘤发生过程中发生的关键代谢和表观基因组变化,并揭示了潜在的癌症治疗策略。
△ 人类胃食管交界处来源的类器官图像,通过使用CRISPR/Cas9 基因编辑技术双敲除关键抑癌基因 (TP53/CDKN2A) 进行修饰,导致细胞变得更加癌变
腹膜假性粘液瘤类器官样本库
腹膜假黏液瘤(PMP)是一种罕见的恶性疾病,通常始于阑尾,由于黏液性肿瘤细胞在腹腔内广泛扩散所致。该疾病由于极其罕见,容易误诊,且治疗手段相对复杂,缺乏有效的体外研究模型,因而迫切需要新的研究模型和治疗策略。
2024年8月,西班牙Vall d 'Hebron肿瘤研究所等医疗机构的研究人员发表最新研究成果[3],他们通过120个PMP样本生成了50个患者衍生的类器官(PDO)和异种移植(PDX)模型,深入研究了该疾病并揭示了KRAS和BRFA药物靶点,为PMP的高通量药物筛选和个性化治疗奠定实验基础。通过实验数据和临床前模型的收集,为临床评估患者的系统性靶向治疗铺平道路。
△ 生成PMP-PDO、PDX和PDXO的模型集合,这些模型具有可识别的KRAS和BRAF药物靶点,以指导分子靶向治疗的选择
类器官与人体器官拥有高度相似的组织学特征,并能在体外重现其生理功能。不仅可以模拟疾病发育,也为更多疾病研究构建了全新体外模型,在个性化治疗,药物筛选,器官移植等方面表现出良好的应用潜力,成为再生医学领冉冉升起的新星。
Write in the last
写在最后
类器官技术将在医学基础研究和临床前转化方面扮演愈来愈重要的角色。随着类器官技术的持续精进和应用范围的拓展,预计未来将有更多类器官、患者来源类器官生物样本库的出现,有望推动医疗技术的革新,助力疾病治疗、药物筛选、新药研发、精准治疗等领域的进步,为疾病治疗提供靶向性、个性化的新选择。
参考资料(可上下滑动查看):
[1] Joep Beumer et al, Description and functional validation of human enteroendocrine cell sensors, Science (2024). DOI: 10.1126/science.adl1460. www.science.org/doi/10.1126/science.adl1460
[2] HUA ZHAO,YULAN CHENG,ANDREW KALRA, et al. Generation and multiomic profiling of a TP53/CDKN2A double-knockout gastroesophageal junction organoid model, Science Translational Medicine (2022). DOI:10.1126/scitranslmed.abq6146
[3] Jordi Martínez-Quintanilla et al, Precision oncology and systemic targeted therapy in Pseudomyxoma Peritonei, Clinical Cancer Research (2024). DOI: 10.1158/1078-0432.CCR-23-4072
END
免责声明:本文仅作知识交流与分享及科普目的,不涉及商业宣传,不作为相关医疗指导或用药建议。文章如有侵权请联系删除。
OTC2024类器官前沿应用与3D细胞培养论坛圆满落幕,点击图片可查看会后报告,咨询OTC2025类器官论坛请联系:王晨 180 1628 8769
抗体药物偶联物临床研究
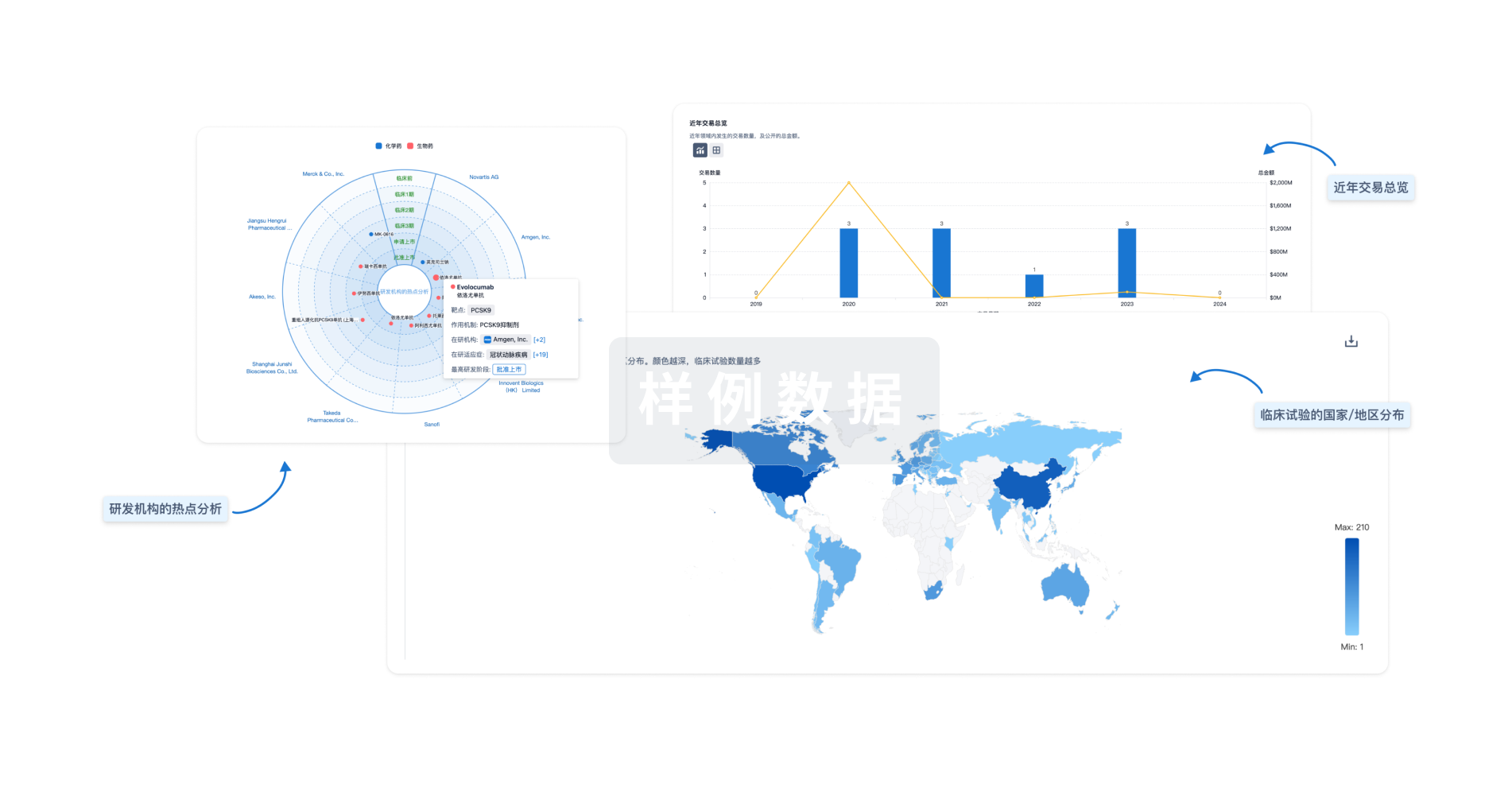
分析
对领域进行一次全面的分析。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用

