预约演示
更新于:2026-02-28
SU3327
更新于:2026-02-28
概要
基本信息
在研机构- |
权益机构- |
最高研发阶段无进展药物发现 |
首次获批日期- |
最高研发阶段(中国)- |
特殊审评- |
结构/序列
分子式C5H3N5O2S3 |
InChIKeyNQQBNZBOOHHVQP-UHFFFAOYSA-N |
CAS号40045-50-9 |
关联
100 项与 SU3327 相关的临床结果
登录后查看更多信息
100 项与 SU3327 相关的转化医学
登录后查看更多信息
100 项与 SU3327 相关的专利(医药)
登录后查看更多信息
29
项与 SU3327 相关的文献(医药)2026-01-01·INTERNATIONAL JOURNAL OF ANTIMICROBIAL AGENTS
SU3327: A multi-target compound targeting bacterial menaquinone and DNA
Article
作者: Wang, Yang ; Sun, Chengtao ; Wang, Shuge ; Liu, Dejun ; Zhao, Ke ; Wu, Congming ; Chen, Hongliang ; Chen, Ziqi
OBJECTIVES:
The rapid emergence of antibiotic resistance (AMR) threatens global health by rendering existing antibiotics ineffective, demanding novel antimicrobial agents with unique mechanisms of action. SU3327 (also known as Halicin), identified through deep learning-based drug screening, has demonstrated broad-spectrum antibacterial activity and high safety in vivo. However, its precise mechanism of action remains unclear. This study aimed to elucidate the antibacterial mechanism of SU3327 and to explore its potential as a multi-target therapeutic.
METHODS:
We employed microbiological, biochemical/biophysical, mass spectrometry, electrochemical, and transcriptomic analyses to study SU3327's effects. We examined its impact on proton motive force (PMF), intracellular ATP synthesis, and electron transport chain (ETC) function, including Complex I inhibition and menaquinone interaction. We also evaluated nitroreductase bioactivation, identified the hydroxylamine metabolite, and quantified resulting oxidative DNA damage.
RESULTS:
SU3327 disrupts bacterial energy metabolism by targeting menaquinone (MK) in the ETC. It inhibits Complex I, impeding electron transfer, disrupting the PMF, and impairing ATP synthesis. Additionally, SU3327 enters Escherichia coli ATCC 25922, undergoing nitroreductase bioactivation, forming hydroxylamine compounds (R-NHOH) that react with DNA, generating hydroxyl radicals (•OH) and inducing oxidative DNA damage. These results indicate that SU3327 operates via a dual-target mechanism: disrupting bacterial respiration and causing DNA damage.
CONCLUSIONS:
SU3327 is a first-in-class multi-target antibacterial agent acting via two synergistic mechanisms. Its dual action enhances efficacy and may reduce resistance likelihood by requiring mutations in distinct pathways. This study advances understanding of SU3327 and supports its development as a novel antibiotic for veterinary medicine.
2025-06-17·ACS Omega
Halicin Reduces the Warburg Effect and Overcomes Drug Resistance by Activating the Pyruvate Kinase M2 Pathway for Triple-Negative Breast Cancer Treatment
Article
作者: Xia, Lei ; Huo, Shao-Hu ; Gu, Han ; Feng, Yu-Bin ; Nie, Xuan ; You, Ye-Zi ; Zhan, Xiang ; Zhou, Xiao-Hong ; Wang, Zhe ; You, Wei
Breast cancer represents the most common cancer among women worldwide. Triple-negative breast cancer (TNBC), a particularly aggressive, metastatic, and drug-resistant subtype of breast cancer, poses a significant clinical challenge because of its resistance to treatment. The efficacy of commonly employed chemotherapeutic agents, such as paclitaxel, is severely limited. In this study, we identified the chemical compound Halicin, which inhibits the proliferation of various cancers, including TNBC. In vitro and in vivo experiments demonstrated that Halicin could activate pyruvate kinase M2 from its inactive dimers to active tetramers, changing the metabolism of tumor cells and suppressing tumor growth. Moreover, Halicin disrupted mitochondria and downregulated the antiapoptotic gene Bcl-2 to overcome multidrug resistance. Further experiments revealed that Halicin inhibits the growth of orthotopic breast tumors through intraperitoneal injection. The discovery of Halicin as an anticancer drug holds great promise for the treatment of TNBC.
2025-03-04·JAC-Antimicrobial Resistance
SAAP-148 and halicin exhibit synergistic antimicrobial activity against antimicrobial-resistant bacteria in skin but not airway epithelial culture models
Article
作者: Nibbering, Peter H ; Dorin, Julia R ; Lennard, Patrick R ; Hiemstra, Pieter S
Abstract:
Background:
The escalating global threat of antimicrobial resistance (AMR) necessitates the development of novel antimicrobial agents, innovative strategies, and representative infection models to combat AMR bacterial infections. Host defence peptides (HDPs) and their derivatives have been proposed as complements to conventional antibiotics due to their antibacterial activity and modulation of the immune response.
Objectives:
This study investigated the novel use of the HDP-derived synthetic antibacterial and anti-biofilm peptide (SAAP)-148 as a pretreatment in epithelial tissue models to prevent colonization by AMR bacteria. The combined activities of SAAP-148 pretreatment with post-infection halicin to treat infections were also explored.
Methods:
Employing cultured human skin equivalents (HSEs) and primary bronchial epithelial cells (PBECs) as models of tissue infection, we examined the prophylactic and therapeutic effects of SAAP-148, both singularly and in combination with the repurposed antibiotic halicin, against AMR bacteria. We additionally interrogated the response of HSE and PBEC cultures to SAAP-148 treatment via confocal microscopy and quantitative PCR of native HDPs and inflammatory cytokine genes.
Results:
Our findings demonstrated that pretreatment with SAAP-148 significantly reduces colonization of HSEs and PBECs by AMR Staphylococcus aureus and Pseudomonas aeruginosa. Confocal microscopy revealed differential uptake and localization of SAAP-148 in these tissues, correlating with its distinct activity in these tissues. SAAP-148 exposure temporarily increased expression of the HDPs cathelicidin (CAMP) and β-defensin 1 (DEFB1), and the cytokine IL-8 (CXCL8), which did not correlate with the transient antibacterial activity observed. Sequential treatment with SAAP-148 prior to infection with AMR S. aureus and post-infection halicin treatment demonstrated synergistic activity in HSEs, whereas this combined activity was indifferent in PBEC cultures.
Conclusions:
These results support SAAP-148 as a candidate for pre-infection prophylaxis and synergistic antibiotic therapy with halicin in skin, broadening the potential of both agents to address AMR bacterial infection.
24
项与 SU3327 相关的新闻(医药)2026-02-17
(来源:麻省理工科技评论)
在塞萨尔・德拉富恩特(César de la Fuente)十几岁时,为了规划日后的职业发展方向,他曾列出一份全球热门行业清单。他按照各国政府投入解决资金的多少反向排序,微生物耐药性排在首位。
二十年过去,这一问题并未消失,反而变得更加严峻。由细菌、真菌和病毒进化出耐药机制所引发的感染,如今每年造成超过 400 万人死亡。发表在《柳叶刀》上的一项近期分析预测,到 2050 年,这一数字可能突破 800 万。如今已是生物工程师与计算生物学家的德拉富恩特,与合成生物学家詹姆斯・柯林斯(James Collins)在 2025 年 7 月《物理评论快报》的文章中警告,一个“后抗生素时代”正在逼近。届时,大肠杆菌、金黄色葡萄球菌等常见细菌的耐药菌株所引发的感染,即便目前仍有药物可以治疗,未来也可能致命。他们写道,抗生素研发渠道依旧极度匮乏,高研发成本、漫长周期与低投资回报率共同阻碍了相关进展。
但德拉富恩特正利用 AI 开创不一样的未来。他在宾夕法尼亚大学的团队正在训练 AI 工具,广泛且深入地搜索基因组,寻找具有抗菌特性的多肽。他的目标是将这些由最多 50 个氨基酸连接而成的多肽分子组合成不同结构,其中包括自然界中从未出现过的形式。他希望,这些成果能够保护人体,抵御那些对传统疗法产生耐受的微生物。
他的研究在出人意料的地方找到了具备潜力的候选分子。2025 年 8 月,他所带领的宾夕法尼亚大学机器生物学团队共 16 名科学家,在名为古菌的古老单细胞生物的基因序列中发现了隐藏的多肽。在此之前,他们还从蛇、胡蜂与蜘蛛的毒液中筛选出一批候选分子。在一项德拉富恩特称之为“分子去灭绝”的持续项目中,他与合作者正在扫描已发表的灭绝物种基因序列,寻找具备潜在功能的分子。这些物种包括尼安德特人、丹尼索瓦人等人科物种,猛犸象等知名巨型动物,以及古代斑马与企鹅。德拉富恩特认为,在地球生命演化史上,某些生物可能进化出了如今依然有用的抗菌防御机制。这些早已消失的基因序列,让猛犸素-2、大地懒素-2、远古海牛素-1 等化合物得以“复活”。过去几年间,这场分子层面的大规模探索,让德拉富恩特积累了超过百万条基因序列库。
现年 40 岁的德拉富恩特,已斩获美国微生物学会、美国化学学会等机构颁发的多项荣誉。2019 年,《麻省理工科技评论》曾将他评为“35 岁以下科技创新 35 人”,以表彰他将计算方法应用于抗生素发现领域。他被公认为将 AI 用于解决现实问题的领军人物。任职于麻省理工学院的柯林斯表示,他真正推动了这一领域的开拓。两人并未在实验室开展合作,但柯林斯长期处于 AI 药物研发包括抗生素探索的前沿。2020 年,柯林斯团队利用 AI 模型预测出一种广谱抗生素哈利辛,目前该药物已进入临床前研究阶段。
柯林斯表示,抗生素研发领域需要研究者拿出尽可能多的创意与创新。而德拉富恩特在多肽方面的工作推动了整个领域的发展。他评价道,塞萨尔才华出众,极具创新精神。
探索之路
德拉富恩特将微生物耐药性称为一个“几乎不可能解决”的问题,但他认为“几乎”二字中仍有巨大探索空间。他说自己喜欢挑战,而这正是终极挑战。
他表示,抗生素的使用、过度使用与滥用,共同推动了微生物耐药性的产生。而传统的药物发现、制备与检测方法成本极高,且常常走入死胡同,导致问题持续恶化。过去许多尝试开展抗生素研发的公司最终都以倒闭收场,因为最终无法获得可观的投资回报。
抗生素发现从来都是一条混乱且充满噪音的道路,依赖偶然发现,充满不确定性与错误方向。几十年来,研究者主要依靠蛮力式的机械方法开展工作。德拉富恩特说,科学家深入土壤与水体,再从这些复杂有机物中尝试提取抗菌分子。
但分子结构可能异常复杂。研究者估算,可被合成的有机组合数量约在 10 的 60 次方量级。作为对比,地球上的沙粒数量约为 10 的 18 次方。加拿大麦克马斯特大学的化学生物学家乔纳森・斯托克斯(Jonathan Stokes)表示,任何领域的药物研发都是一场统计游戏,必须有足够多的尝试,才有可能命中一个目标。斯托克斯一直在利用生成式 AI 设计可在实验室合成的新型抗生素候选物,并与柯林斯合作开展哈利辛相关研究。
不过这些尝试必须是有效的尝试,而 AI 恰好能提升研究者的精准度。德拉富恩特解释,生物学本身就是一种信息来源,就像一整套代码。DNA 代码由四种碱基组成,蛋白质与多肽则由 20 种氨基酸构成,每一个“字母”对应一种氨基酸。德拉富恩特表示,他的工作本质上是训练 AI 模型,识别编码抗菌肽的序列。他说,如果从这个角度思考,就可以设计算法挖掘这些代码,识别出具备功能的分子,它们可以是抗菌药物、抗疟疾药物,也可以是抗癌药物。
德拉富恩特表示,从实际应用来看,目前仍未达到目标。这些多肽尚未转化为能帮助患者的可用药物,剂量、递送方式、具体作用靶点等大量细节仍需梳理。但抗菌肽具备独特优势,因为人体本身就在使用这类物质。它们是免疫系统的重要组成部分,通常是抵御病原体感染的第一道防线。传统抗生素往往只依靠单一机制杀灭细菌,而抗菌肽通常采用多重作用模式。它们可以同时破坏细胞壁、内部遗传物质以及多种细胞过程。一种细菌病原体可能对传统药物的单一作用机制产生耐药性,但未必能抵御抗菌肽的多重攻击。
从发现到临床
德拉富恩特的团队是众多推动 AI 抗生素研发边界的团队之一。他主要聚焦多肽,柯林斯则专注小分子发现。麦克马斯特大学的斯托克斯同样研究小分子,其模型可以筛选出有潜力的新分子,并预测它们能否被合成。柯林斯表示,研究者真正将 AI 有效用于药物研发只有短短几年时间。
与斯托克斯和柯林斯有过合作的斯坦福大学计算机科学家邹嘉(James Zou)表示,即便在这么短的时间里,相关工具也已发生变化。研究者从使用预测模型转向开发生成式方法。邹嘉解释,在预测模式下,研究者会从已知有潜力的大型候选库中进行筛选。生成式方法则提供了另一种可能,即从头设计全新分子。例如去年,德拉富恩特团队使用一种生成式 AI 模型设计一系列合成多肽,再用另一种模型进行评估。团队选取其中两种化合物,在感染耐药鲍曼不动杆菌的小鼠身上进行测试。世界卫生组织已将这种病菌列为耐药性研究的 “极重要优先对象”。两种化合物均安全有效地治愈了感染。
但该领域仍处于发现阶段。在当前研究中,德拉富恩特正努力让候选药物更接近临床试验。为此,他的团队正在开发一个名为 ApexOracle 的多模态模型。该模型旨在分析新病原体,精准定位其基因弱点,匹配可能对其有效的抗菌肽,再预测由这些多肽构成的抗生素在实验室测试中的表现。他表示,这一模型整合了化学、基因组学与语言模型的认知。他补充说,目前研究仍处于初步阶段,但即便无法完美运行,也将帮助引导下一代 AI 模型朝着攻克耐药性这一最终目标前进。
他相信,借助 AI,人类研究者现在有机会直面这一威胁。这项技术已经节省了数十年的研究时间,如今,他希望它也能拯救生命。
他说,这就是我们今天所处的世界,令人难以置信。
原文链接:
https://www.technologyreview.com/2026/02/16/1132516/cesar-de-la-fuente-using-ai-antibiotics-hunt/
微生物疗法
2026-02-06
● 这场大会为何值得传统医药人关注?
2026年1月12日至15日,旧金山。
全球医药界最具影响力的年度盛会——第44届摩根大通医疗健康年会(J.P. Morgan Healthcare Conference)如期举行。
业内有一句话:"想知道未来一年医药行业往哪走,先看JPM怎么说。"每年一月,全球顶级药企CEO、生物科技创业者、华尔街分析师和机构投资者齐聚旧金山,在四天时间里完成数百场企业路演、数千名机构投资者。这里释放的信号——谁在融资、谁在并购、哪些赛道被反复提及——往往预示着整个行业接下来一年的资本流向与战略重心。
过去几年,这场大会的议题对传统医药从业者来说多少有些"隔岸观火"的意味:mRNA疫苗的狂飙突进、GLP-1的资本神话、AI制药的风口轮转……那是另一个世界的游戏,规则由合成分子和临床数据书写,动辄以十亿美元计。
但行业风向正在发生微妙的变化。
作为闭门性质的盛会,外界无从得知每一场讨论的具体内容。然而,结合本届大会的公开议程、头部企业近期的战略布局,以及过去一年行业研究的演进脉络,一些信号值得关注:当创新药研发成本高企、临床失败率居高不下、专利悬崖逼近,资本开始重新审视一个问题——"创新"是否必须意味着从零开始?那些经过人类长期使用验证的天然活性成分,是否可能成为新一轮价值发现的起点?
这正是本文想要探讨的视角:JPM 2026透露了哪些行业趋势?这些趋势与传统医药与大健康产业之间,是否存在被低估的连接点?
三大趋势信号:
为什么说传统医药的"时间窗口"正在打开?
趋势一
AI革命的意外转向
从"合成新分子"到"挖掘老宝藏"
1月12日下午的专题研讨"AI革命与药物发现"(AI Revolution in Drug Discovery)是本届大会最受关注的议题之一。
值得注意的是,整个行业对AI制药的期待正在发生微妙转向。 早期的热情集中在"AI如何设计全新的化学结构",而现在一个更务实的问题开始浮现:如何用AI去重新解析那些已经被人类使用了数百甚至数千年的天然化合物?
正如《Nature Reviews Drug Discovery》一篇由60余位国际科学家联合撰写的文献所指出的:天然产物"作为天然代谢物的起源,使其很可能成为转运系统的底物,从而使药物能够到达其靶点",且具有"相对较高的三维复杂性"——这些特性使其在调控具有挑战性的药物靶点方面具备独特优势。[1]
这一转向的背后是冷峻的现实:
Source: Deloitte Centre for Health Solutions. "Be brave, be bold: Measuring the return from pharmaceutical innovation." 15th Edition. March 2025.[2]
当从零开始的创新变得如此昂贵和高风险时,资本开始问一个简单的问题:有没有"捷径"?
答案指向了天然产物(Natural Products)——那些已经经过人类长期使用验证的化合物。
AI如何改变天然产物研究的游戏规则?
AI在天然产物药物发现中的应用正在多个关键领域取得突破[1]:
基因组与代谢组挖掘: 深度学习方法如DeepBGC、GECCO和SanntiS能够识别传统基于规则的方法无法捕获的新型生物合成基因簇(BGCs)。例如,decRiPPter算法发现了属于全新镧肽类别的pristinin;DeepRiPP则实现了核糖体合成天然产物的自动化发现。
结构表征的AI加速: 卷积神经网络工具SMART 2.0指导了新型大环内酯symplocolide A等新化合物家族的发现和结构解析。CANOPUS等深度神经网络能够从质谱数据系统分类未知代谢物。
靶点与活性预测: 基于深度学习的chemprop消息传递神经网络成功预测了合成化合物halicin和abaucin的杀菌活性,以及其他八种结构上与已知抗生素类别明显不同的抗生素分子。
这对传统医药意味着什么?
传统医药体系——无论是中医药、藏医药、阿育吠陀还是其他民族医药——本质上是人类历史上规模最大、持续时间最长的"真实世界临床试验"。
以藏医药为例,其经典典籍《四部医典》成书于公元8世纪,记载了数千种药物的性味、功效、配伍禁忌。这不是"古老的传说",而是经过1200多年持续使用和迭代优化的"临床数据库"。
用资本市场能听懂的语言来说:
传统医药积累的不是"经验",
而是"数据资产"——
一种经过数千年真实世界验证的、
尚未被结构化挖掘的"大数据"。
当AI具备了从海量文本中提取结构化信息、从复杂成分中预测作用靶点的能力时,这些"数据资产"的价值开始显现。
然而,AI在天然产物药物发现领域的进展,主要受限于缺乏大规模、高质量的数据集,而非创新算法的匮乏。这为传统医药企业提出了一个关键问题:如何将这些"沉睡"的知识资产转化为AI可以处理的结构化数据?
该文献特别指出了两个数据缺口:一是生物合成修饰酶的催化活性数据(用于预测天然产物结构),二是生物活性数据(用于理解构效关系)。这恰恰是传统医药体系可能贡献独特价值的领域——千年临床实践中积累的"适应证-方剂-疗效"关联,本质上就是一种未被数字化的生物活性数据库。
这可能是未来五年传统医药企业最重要的战略投资方向之一:不仅投资AI算法,更要投资数据基础设施建设。
阿如拉藏医药集团旗下金诃藏药核心产品系列,65个国药准字号批文,覆盖13大治疗领域
趋势二
2000亿美元肥胖市场的"第二入口"
当GLP-1遇上"药食同源"
1月13日下午的专题研讨"2000亿美元肥胖治疗市场"(The $200 Billion Obesity Treatment Market)吸引了大量投资者参与。
GLP-1类药物(如司美格鲁肽、替尔泊肽)无疑是当前这一赛道的绝对主角。根据IQVIA最新报告,2020年全球抗肥胖药物(AOMs)市场规模约为30亿美元,到2024年已飙升至300亿美元以上——四年内增长超过十倍。预计到2034年,全球抗肥胖药物市场规模将达到约1300亿美元,年复合增长率为13-15%[3]。
但本届大会的讨论已经开始触及更深层的问题:
供应瓶颈:即使产能全开,也难以满足爆发式增长的需求;
支付压力:每月数百至上千美元的费用,长期可负担性存疑;
使用门槛:注射给药、需要处方、潜在副作用,限制了普及范围;
停药反弹:部分研究显示停药后体重回升明显。
这意味着什么?
GLP-1类药物正在定义肥胖治疗的"医疗级"市场,但这个市场的边界是有限的。值得注意的是,IQVIA报告指出,目前GLP-1和/或GIP受体激动剂占据了96%的市场份额,但到2034年,非GLP-1/GIP类药物的市场份额预计将增长7倍,达到28%,年复合增长率高达36-39%[3]。这意味着市场正在寻找多元化的解决方案。
真正的"蓝海"可能在于:
Source:医疗级数据基于IQVIA Forecast Link 2025年7月报告[2];亚临床级与消费级为基于全球肥胖流行趋势的估算
市场背景的严峻性:
根据IQVIA报告援引的数据,到2035年,全球预计将有15.3亿成年人患有肥胖症(BMI≥30),如果加上超重人群(BMI 25-30),总数将达到33亿——占全球成年人口的54%以上。这些状况预计将使全球经济每年损失超过4万亿美元,相当于全球GDP的近3%[3]。
传统医药在"亚临床级"和"消费级"市场具有天然优势:
"药食同源"的文化基础:在亚洲市场,"以食养身""调理体质"的理念深入人心,消费者对天然来源的产品接受度高;
安全性背书:与新合成药物相比,传统药材经过长期使用验证,安全性边界相对清晰;
差异化定位:区别于GLP-1类药物的"快速减重",传统医药可以强调"整体调理""代谢平衡""可持续的生活方式";
价格优势:相比每月数百美元的处方药,传统医药产品的可负担性更强,有利于长期使用;
口服制剂趋势的契合:IQVIA报告指出,约40%的在研肥胖药物为口服制剂[3],这反映了市场对更便捷给药方式的需求——而传统医药天然以口服为主。
藏医药的独特机遇:藏医药在代谢健康领域有着丰富的理论和实践积累。以"培根"(藏医体液理论中与代谢功能密切相关的概念)失调为例,藏医药发展出了一系列针对性的调理方法,包括饮食指导、生活方式调整和药物干预。
阿如拉藏医药集团旗下藏有引力健康集团大健康产品矩阵,涵盖7大品类近100款产品
与新兴疗法的差异化定位:IQVIA报告详细列出了当前肥胖治疗的创新方向,包括双靶点/三靶点肠促胰素激动剂(如替尔泊肽、retatrutide)、β2-肾上腺素能激动剂(如bimagrumab,可促进脂肪氧化并保留肌肉)、中枢神经系统靶向疗法等[3]。这些疗法追求的是"更强效的减重",而传统医药可以占据一个差异化的生态位:"更温和的调理"与"更可持续的生活方式"。
藏有引力·藏式健康管理中心,提供个性化健康调理方案
趋势三
中国创新药的"出海时刻"
对传统医药的启示
今年JPM最明显的变化之一,是中国创新药企业的存在感。百济神州、传奇生物、信达生物的路演场场爆满,讨论的不再是"中国市场有多大",而是"你们的全球三期数据什么时候出来"[4]。
2025年,中国创新药对外授权交易实现历史性突破:全年交易总金额达1356.55亿美元,交易数量157笔,较2024年的519亿美元、94笔实现翻倍式增长,首次超越美国成为全球创新药授权交易规模最大的国家[5]。在肿瘤、自身免疫领域,已经有中国企业开发的药物在美国和欧洲获批上市。
几个值得注意的变化:
首先是话语方式的转变。五年前中国企业在JPM讲的是"我们有成本优势""中国市场很大",现在讲的是作用机制、临床终点、与竞品的头对头数据。投资者的态度也变了,从"你们是谁"变成"你们的分子在全球竞争中排第几"。
其次是监管路径逐渐清晰。FDA对中国临床数据的态度从怀疑到接受,"中国数据支持全球获批"的案例越来越多。一条可复制的路径正在形成:国内做早期开发,全球做关键试验,多地区同步上市。
第三是资本市场的态度。2025年生物医药融资环境依然分化明显,但有差异化管线和全球能力的中国企业持续获得资本青睐。信达生物与武田制药达成总金额最高114亿美元的合作,恒瑞医药与GSK的合作突破120亿美元,三生制药与辉瑞的交易以12.5亿美元首付款刷新国内纪录——这些数字本身就是估值认可的最直接证明[4]。部分公司的交易估值已经和欧美同类资产相当,"中国公司打折"的逻辑正在被打破。
这对传统医药意味着什么?
中国创新药走出去,本质上走的是"在别人的赛道上证明自己"这条路——相同的靶点、相同的适应症、相同的评价标准。这条路已经证明走得通,但也注定拥挤。
传统医药的机会可能不在于复制这条路,而在于开辟另一种可能。
传统医药的位置在哪里?
根据Grand View Research报告,美国补充与替代医学市场规模2025年已达528亿美元,预计到2033年将增长至3755亿美元,年复合增长率高达27.8%[6]。消费者对"自然""整体""个性化"的需求是真实存在的,但市场上缺少有科学背书、品质稳定的高端产品。这个空白和传统医药的优势高度重合。
慢性病管理是另一个方向。和急性病不同,慢病管理要的不是"单点突破",而是"长期可持续"。传统医药讲的"调理""平衡""治未病",和慢病管理的思路天然接近。代谢健康、消化功能、睡眠、情绪——这些领域有大量需求没被满足。
还有老龄化。全球都在讨论怎么让人"活得长"变成"活得好"。传统医药的整体观和养生理念,和这个方向是吻合的。
阿如拉藏医药集团大健康产品线GMP标准生产车间
几点现实的考量:
创新药出海用了二十年走到今天,传统医药的国际化不会更快。文化隔阂、认知障碍、监管的不确定性、研究需要的长期投入——这些都是绕不过去的坎。
但创新药的经验至少说明一件事:当科学表达、质量标准和商业逻辑能够对接全球体系的时候,"中国来源"不是减分项,反而可以变成独特的标签。
传统医药从来不缺价值,缺的是让这种价值被全球市场理解和接受的方式。创新药正在回答这个问题的前半部分,传统医药需要找到自己的答案。
● 结语:窗口期的意义
回到文章开头的问题:为什么说传统医药的"时间窗口"正在打开?
三个趋势给出了不同角度的答案:
AI的转向,让传统医药数千年的实践积累有机会被重新"看见"。当资本市场开始为"天然产物数据库"定价,藏医药典籍中那些经过1200多年验证的组方和配伍逻辑,就有了进入现代药物研发体系的可能。《四部医典》不再只是文化遗产,而是一座尚未被结构化挖掘的临床数据库。
GLP-1的边界,在证明一个巨大市场的同时,也暴露出单一路径的局限。2000亿美元的肥胖治疗市场不会只属于注射针剂,"亚临床"和"消费级"的广阔地带,正是传统医药可以施展的空间。藏医药在代谢健康领域有独特的理论积累——"培根"失调的调理体系、针对血脂和血糖的多靶点食疗方案、强调体质辨识的个体化干预——这些都与当前市场寻找的"GLP-1之外的选择"高度契合。
创新药出海的成功,则提供了一个参照系:当科学表达和质量标准能够对接全球体系,"中国来源"可以从折价因素变成溢价标签。藏医药面临的不是"有没有价值"的问题,而是"如何让价值被理解"的问题。创新药用了二十年走通这条路,藏医药需要找到属于自己的节奏和方式。
这三条线索指向同一个判断:传统医药正在获得一个重新定义自身的机会,而藏医药的特点——系统完整的理论体系、丰富的天然药物资源、在高原代谢适应方面的独特经验——让它在这个窗口期有独特的位置。
来源:阿如拉
信使RNA疫苗并购核酸药物
2026-02-02
·精准汇
一家公司上市募资20亿港元,背后是什么趋势?
导语
2025年12月31日,一家中国AI制药公司登陆港交所,募资超20亿港元,腾讯、礼来等15家行业巨头成为基石投资者。这家公司的核心药物将于2026年上半年启动关键临床试验。这不仅仅是一次融资,更是AI制药从概念走向价值兑现的标志性事件。英矽智能的成功上市,标志着AI制药正式进入"黄金时代"。第一部分:AI如何颠覆传统"双十魔咒"传统药物研发的"三高"困境
新药研发领域长期存在一个令人望而却步的"双十魔咒":平均耗时超10年,耗资逾10亿美元,而成功率却不足10%。这三重门槛如同三座大山,阻碍着创新药的诞生。
更令人沮丧的是,90%的候选药物会在临床试验阶段折戟沉沙。科学家们像在大海中捞针,依靠经验、直觉和大量资源堆砌出极少数成功药物。这种"经验试错"模式在过去几十年里虽屡建奇功,却也越来越难以满足日益增长的医疗需求。AI带来的三大革命性突破时间维度:从10年缩短至4-6年
人工智能通过深度学习海量生物医学文献、基因组数据和蛋白质数据库,能够在虚拟的分子宇宙中快速筛选潜在药物靶点。罗氏制药整合2000万种化合物数据构建AI筛选平台,将KRAS G12C抑制剂的先导化合物发现周期从4.2年缩短至1.8年,研发时间缩短了57%。成本维度:从10亿美元降至4亿美元
德勤调查显示,78%的生物制药高管认为AI是驱动行业变革的核心力量。AI不仅能降低实验成本,还能避免大量无效试错。MIT与DeepMind合作发现的新型抗生素Halicin,被《Cell》杂志评价为"近30年抗生素领域重大突破",而其发现成本远低于传统方法。成功率维度:从10%提升至30%以上
AI系统能够预测分子在体内的吸收、分布、代谢、排泄和毒性(ADMET)特性,在早期就筛选掉高风险分子。山东2025年规划显示,AI赋能新药研发项目占比将超30%,覆盖小分子药物、重组蛋白药物等重点领域。到2027年,AI参与的新药一期临床成功率有望突破50%,二期达30%。AI在药物研发全流程中的具体应用
靶点发现阶段
中国科学院团队开发的"脸谱识别"算法,成功发现了抗肿瘤老药甲氨蝶呤的新免疫靶点。AlphaFold已把人类98.5%的蛋白结构算出来,为"无结构"靶标提供了可药化的起点。分子设计与优化阶段
英矽智能的AI引擎"InClinico"准确预测多项临床试验II/III期转化结果,大幅提高研发成功率。生成式AI能够从零开始设计全新的分子结构,ED2Mol、ResGen等算法把结合口袋当作"模板",逐原子"3D打印"出兼顾结合力与成药性的候选分子。临床试验阶段
AI系统如"Deep Patient"能够分析电子健康记录中的非结构化文本,自动识别患者的深层表征,精准匹配符合试验条件的个体。这种基于真实世界数据的智能分型,不仅提高了招募效率,还能发现隐藏的疾病亚型,支持个性化医疗的发展。第二部分:中国在AI制药领域的全球地位与代表企业中国AI制药的全球排名
截至2025年,全球共有102条AI辅助药物管线进入临床阶段,中国占34条,占比达到33.3%。英矽智能的ISM001-005成为国内首个进入临床的AI研发药物,已在临床一期显示良好安全性。中国AI制药企业已从"跟跑者"向"关键贡献者"转变。代表性企业深度解析英矽智能(Insilico Medicine)
上市时间:2025年12月31日
募资规模:超20亿港元
基石投资者:腾讯、礼来等15家行业巨头
核心药物:Rentosertib,将于2026年上半年在中国启动IIb/III期试验
全球布局:在中国、美国、英国等多个国家设有研发中心
英矽智能建成了全球首个AI辅助决策的全自动化机器人实验室,实现从虚拟设计到实验验证的无缝衔接。其商业模式包括AI SaaS平台服务、AI CRO外包合作与AI Biotech自主管线开发三种主导模式。晶泰科技(XtalPi)
核心优势:量子物理+AI+云计算
代表案例:仅用六周就协助辉瑞确认新冠口服药Paxlovid的优势晶型
技术平台:智能药物晶型预测与筛选系统
合作伙伴:辉瑞、默克等国际制药巨头
晶泰科技的技术能够将药物晶型筛选从传统的试错转变为精准预测,大幅缩短研发周期。其AI筛选平台将候选化合物筛选效率提升100倍,成本却只需原来的1%。其他值得关注的中国AI制药企业
望石智慧(StoneAI):专注小分子药物设计,已完成多轮融资,估值超10亿美元深度智耀(DeepWisdom):提供AI驱动的临床研发管理解决方案德睿智药(MindRank AI):专注于肿瘤和神经系统疾病领域的AI药物发现中国AI制药的政策支持环境审评审批加速
2025年9月12日,国家药监局发布《关于优化创新药临床试验审评审批有关事项的公告》,明确设立"30日审评审批通道",将符合条件的创新药临床试验申请审评时间压缩50%。这一政策将加速AI制药企业的临床进程。医保支持创新
2025年6月,国家医保局、国家卫生健康委联合印发《支持创新药高质量发展的若干措施》。该文件从医保谈判续约规则优化、支付考核豁免、诊疗配套支持等角度为创新药发展提供政策保障。研发投入持续增长
恒瑞医药自2011年以来,已有18款新分子实体药物(一类新药)在国内获批上市,累计研发投入超400亿元。其中2021年、2022年、2023年研发投入均突破60亿元,占营收比保持在20%以上。第三部分:未来挑战:数据质量、算法可解释性、监管伦理挑战一:数据质量与偏差问题
AI模型依赖大量高质量、标准化的数据进行训练。若训练集存在种族、性别或实验条件偏差,预测结果可能"以偏概全",甚至加剧健康不平等。
当前面临的主要问题包括:
天然产物-靶标复合物数据稀缺
实验噪声干扰模型泛化能力
复杂生物系统协同效应建模不足
多源异构数据的标准化程度不一挑战二:算法"黑箱"问题
许多深度学习模型虽能给出预测,却难以解释其判断依据。在医疗领域,"可解释性"至关重要,医生需要理解AI为什么推荐某个分子、预测某种风险。
当前的可解释AI(XAI)技术包括:
注意力机制可视化:展示AI关注的分子特征
基于规则的解释:将复杂模型转化为可理解的规则
反事实推理:回答"如果分子结构改变,结果会如何变化"挑战三:数据隐私与安全
医疗数据包含患者的个人健康信息,涉及高度敏感的隐私问题。如何在保护隐私的前提下实现数据共享,是亟待解决的伦理难题。
当前的解决方案包括:
联邦学习:数据不离开本地,只共享模型参数
差分隐私:在数据中加入噪声,保护个体隐私
同态加密:在加密状态下进行计算,保护数据安全挑战四:监管框架滞后
AI制药作为一个新兴领域,监管框架仍在完善中。当前面临的监管挑战包括:
如何评估AI生成药物的安全性和有效性
AI在临床试验中的角色界定
AI决策出错时的责任划分
国际监管标准的协调统一挑战五:算力与成本
训练大模型需要巨大算力,效率与成本不容忽视。若算法效率低,可能出现"省时间却不省钱"的尴尬。此外,人体是动态复杂的系统,AI目前仍难以预测长期毒性、罕见副作用或多靶标、多通路的复杂疾病网络效应。第四部分:AI制药的未来图景技术融合趋势
AI制药正在向三个关键方向演进:
大分子药物开发
AI向抗体设计领域拓展。全球已有20余家公司布局AI抗体药物,Exscientia已将抗体设计纳入技术平台。中国信华生物设计的First-in-class多功能抗体药物即将进入临床前开发阶段。自动化闭环实验室
"干湿结合"模式成为趋势。英矽智能建成全球首个AI辅助决策的全自动化机器人实验室,实现从虚拟设计到实验验证的无缝衔接。实验室能够24小时不间断运行,大幅提升研发效率。虚拟患者与数字孪生
斯坦福大学团队开发的多尺度神经网络模型已能高精度模拟肿瘤细胞对药物的反应。1635例"虚拟乳腺癌患者"成功用于药物测试,极大降低了临床试验的成本和风险。商业模式创新平台化服务
越来越多的AI制药企业从单纯的药物开发转向平台化服务。通过开放API接口,为科研机构和制药公司提供AI药物发现能力,形成"AI即服务"的商业模式。生态协同构建
AI制药企业、传统制药公司、科研院所、医疗机构形成创新生态。通过数据共享、技术合作、联合开发等方式,共同推动AI制药的产业化进程。全球化布局
中国AI制药企业加速国际化布局。一方面,通过FDA、EMA等国际监管机构的审评;另一方面,与国际制药巨头达成战略合作,共同开发全球市场。写在最后
AI制药不是要取代科学家,而是成为他们手中最强大的工具。在这场静默的革命中,人类智慧与机器智能正携手前行,共同开启医学的下一个篇章。
从阿尔茨海默病的早期预警,到抗病毒药物的快速发现,再到临床试验的智能设计,AI正在重塑医药创新的图景。它让药物研发从"试错的艺术"变成"创造的科学"。
未来,随着数据质量的提升、算法的优化、监管框架的完善,AI制药有望攻克更多人类顽疾,为全球患者带来更多来自中国的治疗方案。
但这并不意味着医生会被AI取代。相反,AI会成为医生的最强助手,帮助他们做出更精准的诊断、制定更个性化的治疗方案、发现更多创新药物。
你认为AI会让医生失业还是成为最强助手?欢迎在评论区留言,分享你的观点~
数据来源:
英矽智能招股说明书
德勤《AI制药行业调查报告》
国家药监局官方网站
山东《AI赋能新药研发规划》
各企业公开披露信息
欢迎转发给关注AI制药的朋友~
本文仅为个人观点,如有错误、侵权,请联系修改、删除!
每一篇文字,只为与您共鸣。
点击关注,让我们走得更远。
长按扫码关注
点击蓝字,关注我们
临床2期
100 项与 SU3327 相关的药物交易
登录后查看更多信息
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 细菌感染 | 药物发现 | 美国 | 2019-08-27 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
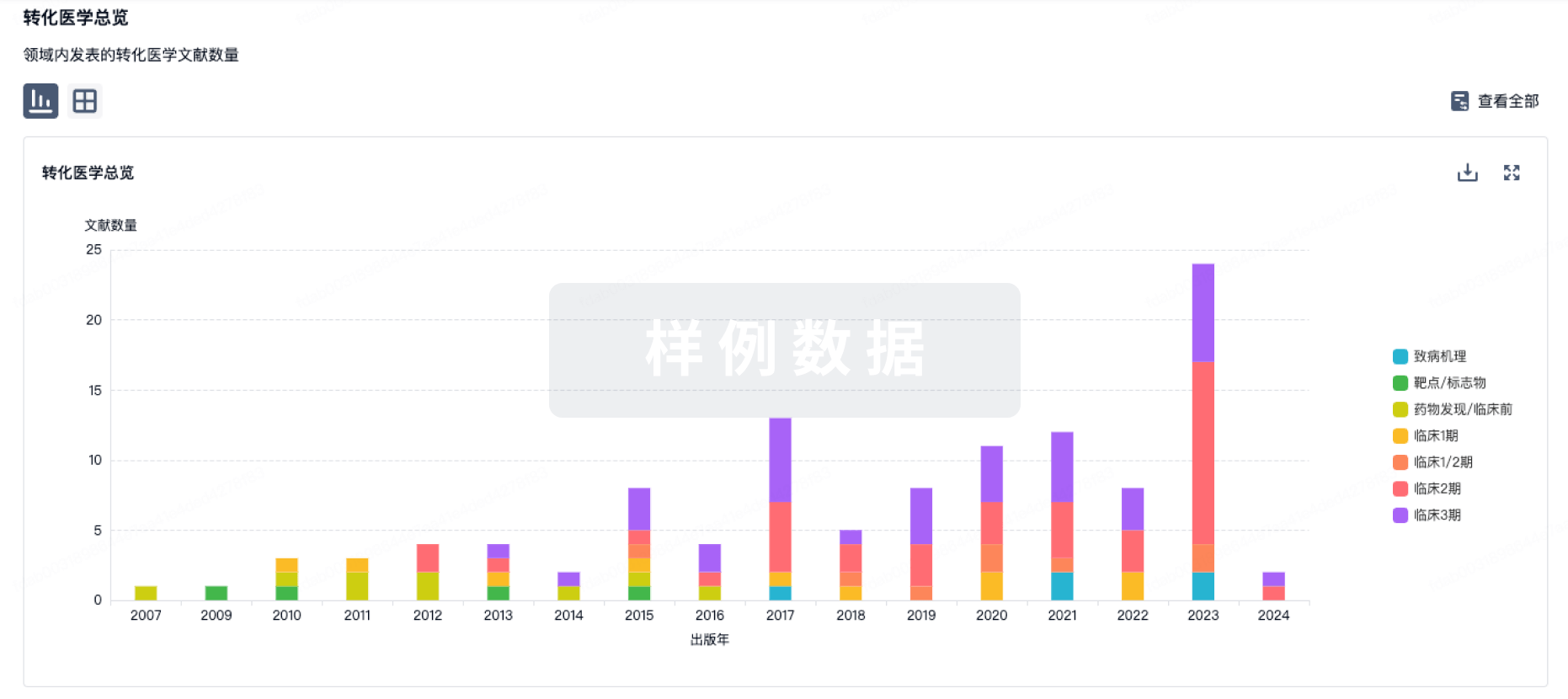
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
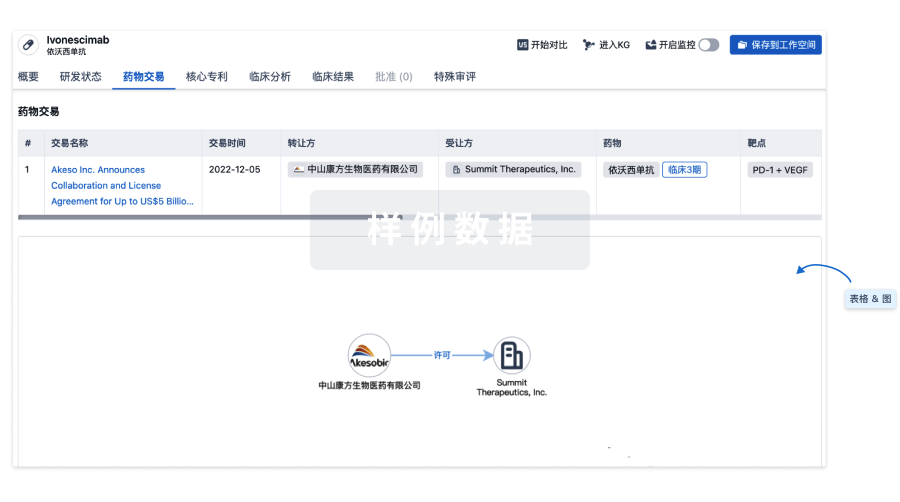
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
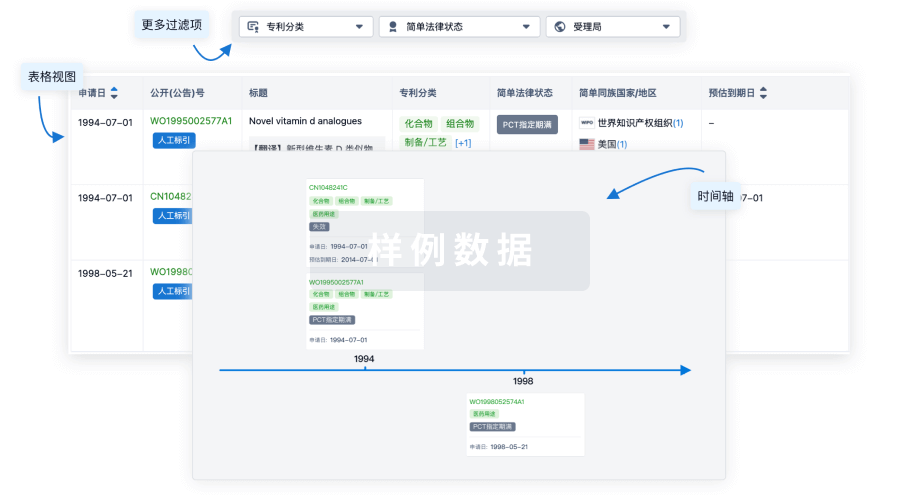
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
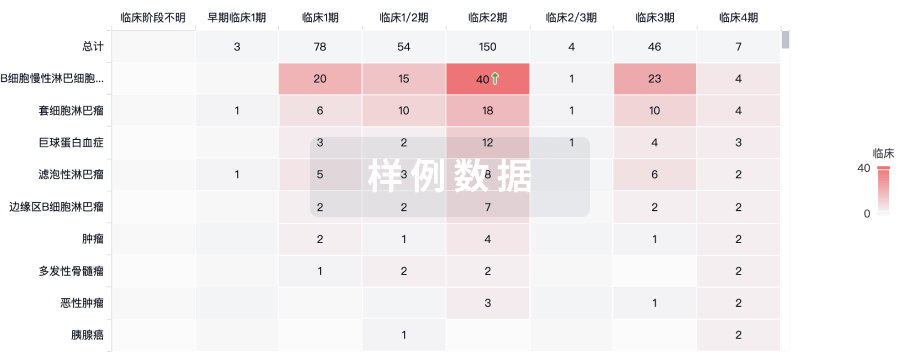
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
批准
利用最新的监管批准信息加速您的研究。
登录
或
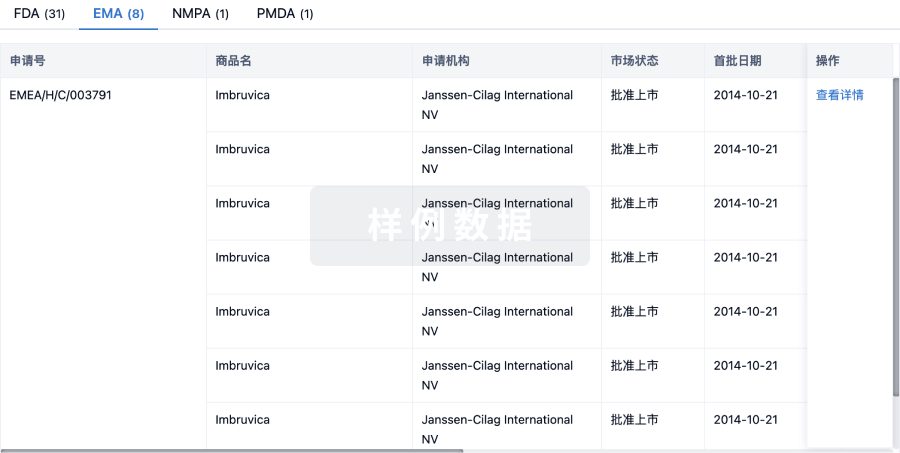
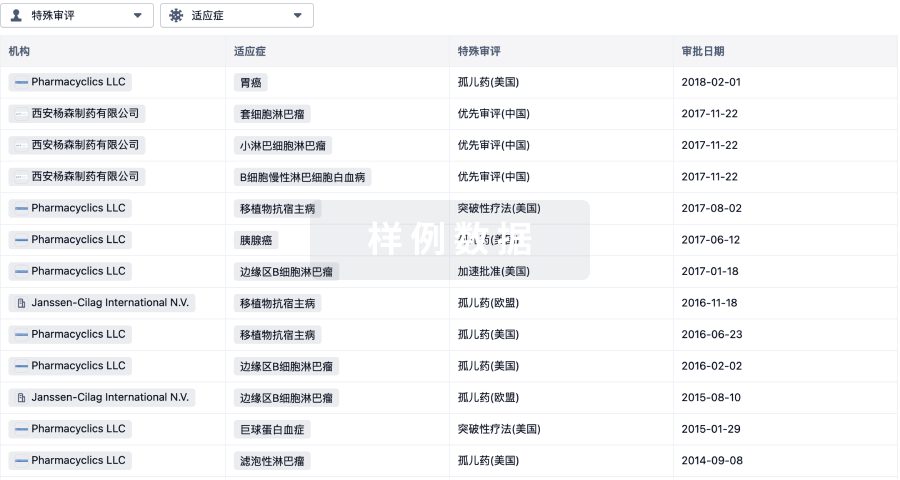
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用

