预约演示
更新于:2025-11-23
Repirinast
瑞吡司特
更新于:2025-11-23
概要
基本信息
最高研发阶段临床2期 |
首次获批日期 日本 (1987-01-01), |
最高研发阶段(中国)临床2期 |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C20H21NO5 |
InChIKeyNFQIAEMCQGTTIR-UHFFFAOYSA-N |
CAS号73080-51-0 |
关联
3
项与 瑞吡司特 相关的临床试验CTR20131347
瑞吡司特片治疗支气管哮喘有效性和安全性的随机双盲双模拟、平行阳性对照、多中心临床试验
评价瑞吡司特片治疗支气管哮喘患者的有效性和安全性.
开始日期- |
申办/合作机构 |
CTR20131348
瑞吡司特颗粒人体生物等效性试验方案
研究河南辅仁堂制药有限公司研制的瑞吡司特颗粒(受试制剂,50 mg/袋)与河南辅仁堂制药有限公司研制的瑞吡司特片(参比制剂,150 mg/片)的人体生物等效性。
开始日期- |
申办/合作机构 |
CTR20131346
瑞吡司特片人体药代动力学试验方案
确定瑞吡司特片单次和多次口服给药在中国健康男性和女性受试者中的药代动力学研究。
开始日期- |
申办/合作机构 |
100 项与 瑞吡司特 相关的临床结果
登录后查看更多信息
100 项与 瑞吡司特 相关的转化医学
登录后查看更多信息
100 项与 瑞吡司特 相关的专利(医药)
登录后查看更多信息
35
项与 瑞吡司特 相关的文献(医药)2025-10-01·WATER RESEARCH
Resistome and microbiome shifts in catfish rearing water: the influence of temperature and antibiotic treatments
Article
作者: Roy, Luke A ; Wang, Hongye ; Li, Xiran ; Abdelrahman, Hisham A ; Wang, Luxin ; Soto, Esteban ; Kelly, Anita M
The increasing reliance on aquaculture for sustainable protein production highlights the need for responsible antibiotic use to manage bacterial infections, particularly in intensive farming systems. This study investigated the effects of three FDA-approved antibiotics (Aquaflor®, Romet®, Terramycin®) at common fish bacterial disease outbreak temperatures (20 °C, 25 °C, and 30 °C) on the microbiome and resistome of aquaculture water using a catfish model system. Metagenomic analyses evaluated the abundance, diversity, and mobility of antimicrobial resistance genes (ARGs) and antibiotic-resistant bacteria (ARB). The impact of temperature on Aquaflor- and Romet-induced changes in ARG abundance, richness, and resistome composition followed a U-shaped trend, with the least effect observed at 25 °C. Of the three antibiotics tested, Terramycin exerted the most significant influence on the water microbiome and resistome, enriching tetracycline resistance genes and co-selecting for floR, sul, and dfrA genes. Temperature also induced notable shifts in the ARB population, with Mantel tests revealing strong correlations between ARG profiles and changes in the overall bacterial community and ARB populations. While certain ARG classes consistently remained associated with specific host phyla, others shifted, highlighting the potential for horizontal gene transfer (HGT) as a critical mechanism for disseminating resistance genes like tet(C), particularly after antibiotic treatment. This is further supported by the observed reduction in plasmid numbers following treatment, which coincided with increased HGT events. Our findings highlight the pivotal role of temperature in influencing resistome dynamics, emphasizing the importance of accounting for environmental factors when applying antibiotics to effectively mitigate antimicrobial resistance in aquaculture systems.
2009-04-27·Journal of chemical information and modeling2区 · 化学
Analysis of the Calculated Physicochemical Properties of Respiratory Drugs: Can We Design for Inhaled Drugs Yet?
2区 · 化学
Article
作者: Luscombe, Christopher N. ; Ritchie, Timothy J. ; Macdonald, Simon J. F.
From an analysis of calculated physicochemical properties for 81 currently marketed respiratory drugs, compounds administered via the inhaled/intranasal routes have a higher polar surface area, a higher molecular weight, and a trend toward lower lipophilicity, when compared with their orally administered counterparts. Ranges of physicochemical space are described for the 29 drugs administered by the inhaled or intranasal routes.
2007-02-01·Photodermatology
作者: Holzle, Erhard ; Horio, Takeshi
A review discusses solar urticaria focusing mainly on the pathomechanism of the disease.Solar urticaria is an uncommon type of phys. urticaria; the action spectrum ranges from UVB to visible light.It is most probably a type I allergic reaction to an as yet to be defined photoallergen.Spectrum of electromagnetic radiation that could inhibit, or augment, the development of solar urticaria is present in many patients.Exogenous agents such as tar and pitch, benoxaprofen, repirinast, and chlorpromazine have been reported to cause solar urticaria in rare instances.Association with PMLE, chronic actinic dermatitis, atopy, and elevated IgE has been reported in some studies.The course is chronic; 15-50% of patients are clear of disease within five years of the onset.Management includes photoprotection, oral antihistamines, desensitization with UVA or psoralen and UVA, cyclosporine, plasmapheresis, and photopheresis.
9
项与 瑞吡司特 相关的新闻(医药)2025-09-04
VANCOUVER, British Columbia, Sept. 04, 2025 (GLOBE NEWSWIRE) -- Algernon Pharmaceuticals Inc. (the “Company” or “Algernon”) (CSE: AGN) (FRANKFURT: AGW0) (OTCQB: AGNPF), a Canadian healthcare company announces that it is changing its name to Algernon Health. The planned name change reflects the Company’s recent announcement that it will be focusing on the Alzheimer’s Disease (AD) diagnostic market with plans to establish specialized neuroimaging clinics across North America as its lead program. The clinics will feature the latest technology of U.S. FDA-cleared, optimized brain-specific Positron Emission Tomography (PET) scanning systems to detect amyloid plaques in patients, providing significantly lower radiation compared to standard PET/CT machines currently in operation globally. The PET scans are covered by Medicare, Medicaid, and private insurance in the U.S. Amyloid plaques are aggregates of mis-folded proteins that form in the spaces between nerve cells and are known to play a central role in AD and are associated with neural degeneration. The amyloid plaques typically first develop in the areas of the brain concerned with memory and other cognitive functions. A positive amyloid confirmation through a PET scan or a spinal tap must be first established before a patient can be treated with either Kisunla or Leqembi, the two U.S. FDA-approved, AD monoclonal antibody treatment therapies that remove amyloid plaque from the brain. These new treatments were developed by Eli Lilly, and Eisai and Biogen, respectively, and are also covered by U.S. Medicare, Medicaid, and private insurance providers. After the first AD drug treatment was approved, in a post-earnings call, GE HealthCare CEO Peter Arduini called it a “profound growth opportunity” for all providers offering PET scans and molecular imaging. The introduction of dedicated neuroimaging clinics provides the Company with a clear pathway to revenue utilizing a technology already U.S. FDA-cleared and covered by U.S. Medicare and Medicaid and private insurance. The Algernon clinics will also offer additional brain PET scans for other forms of dementia, epilepsy, neuro-oncology, and movement disorders, to further expand the provision of healthcare to patients in need and expand clinic revenues. This strategic entry by Algernon into dedicated neuroimaging clinics is significantly based on the fact that the vast majority of PET/CT scanners in the U.S. are prioritized as cancer diagnostic and theranostic tools, and are also dedicated to cardiac imaging. Consequently, there is an insufficient supply of scanners to serve the rapidly expanding AD diagnostics market. The shortage of PET scanners for brain specific scanning also has an impact beyond immediate patient care. With 162 AD drugs under development, there is also a significant opportunity to provide brain specific PET scan imaging services to drug development companies and contract research organizations engaged in clinical trials, as another source of revenue for Algernon. Algernon will update the market shortly on its upcoming expansion and growth plans including the location of it first U.S. flagship neuroimaging clinic. Algernon continues to be the parent company of a private subsidiary called Algernon NeuroScience that has been advancing a psychedelic program investigating a proprietary form of DMT for stroke and traumatic brain injury. The Company also owns a 20% equity position in Seyltx, a private U.S. based drug development company advancing a chronic cough drug called Ifenprodil. Algernon is currently reviewing its repurposed chronic kidney disease drug program which features the drug repirinast as its lead molecule. Christopher J. Moreau CEO Algernon Pharmaceuticals Inc. 604.398.4175 ext 701 info@algernonpharmaceuticals.com investors@algernonpharmaceuticals.com www.algernonpharmaceuticals.com
About Algernon Health
Algernon Health is a Canadian healthcare company focused on the provision of brain specific PET scanning services through a planned network of new clinics in North America for the detection of Alzheimer’s disease, as well as other forms of dementia, epilepsy, neuro-oncology, and movement disorders. Algernon is also the parent company of a private subsidiary called Algernon NeuroScience that has been advancing a psychedelic program investigating a proprietary form of DMT for stroke and traumatic brain injury. The Company also owns a 20% equity position in Seyltx, a private U.S. based drug development company advancing a chronic cough drug called Ifenprodil. Neither the Canadian Securities Exchange nor its Market Regulator (as that term is defined in the policies of the Canadian Securities Exchange) accepts responsibility for the adequacy or accuracy of this release.
CAUTIONARY DISCLAIMER STATEMENT: No Securities Exchange has reviewed nor accepts responsibility for the adequacy or accuracy of the content of this news release. This news release contains forward-looking statements relating to product development, licensing, commercialization and regulatory compliance issues and other statements that are not historical facts. Forward-looking statements are often identified by terms such as “will”, “may”, “should”, “anticipate”, “expects” and similar expressions. All statements other than statements of historical fact, included in this release are forward-looking statements that involve risks and uncertainties. There can be no assurance that such statements will prove to be accurate and actual results and future events could differ materially from those anticipated in such statements. Important factors that could cause actual results to differ materially from the Company’s expectations include the failure to satisfy the conditions of the relevant securities exchange(s) and other risks detailed from time to time in the filings made by the Company with securities regulations. The reader is cautioned that assumptions used in the preparation of any forward-looking information may prove to be incorrect. Events or circumstances may cause actual results to differ materially from those predicted, as a result of numerous known and unknown risks, uncertainties, and other factors, many of which are beyond the control of the Company. The reader is cautioned not to place undue reliance on any forward-looking information. Such information, although considered reasonable by management at the time of preparation, may prove to be incorrect and actual results may differ materially from those anticipated. Forward-looking statements contained in this news release are expressly qualified by this cautionary statement. The forward-looking statements contained in this news release are made as of the date of this news release and the Company will update or revise publicly any of the included forward-looking statements as expressly required by applicable law.
放射疗法
2025-04-11
April 08, 2025 -- Algernon Pharmaceuticals Inc. (the “Company” or “Algernon”) (CSE: AGN) (FRANKFURT: AGW0) (OTCQB: AGNPF), a Canadian clinical stage pharmaceutical development company, is pleased to announce that it has received a notice of allowance from the European Patent Office (EPO) for patent application 19827430.0 for its lead chronic kidney disease (CKD) program drug NP-251 (Repirinast).
The invention claims the use of Repirinast, either alone or in combination with telmisartan, for the treatment or prophylaxis of renal fibrosis or kidney disease. The base claims of the patent will be valid through 2038, excluding any patent term adjustments or extensions which may provide additional protection.
The Company has previously announced the allowance of this patent application in the U.S., Japan and China. The patent is currently pending in Canada.
Repirinast is the Company’s lead candidate for the treatment of CKD based on data showing it reduced fibrosis by 51% with statistical significance and showed an additive benefit to telmisartan in a unilateral ureteral obstruction (UUO) mouse model.
Algernon’s intellectual property strategy for its repurposed drug program includes protecting its compounds by filing patent applications including method of use, dosing, and formulations, and for new composition of matter patents based on novel salt forms.
“This is our first patent issued by the EPO and marks another key milestone for the Company,” said Christopher J. Moreau, CEO of Algernon Pharmaceuticals.
The Company also announces a grant of 50,000 restricted share units (each, an "RSU") and 50,000 stock options to consultants of the Company. Each RSU entitles the recipient to receive one common share of the Company or a cash payment equal to the equivalent of one common share of the Company on vesting. The stock options are exercisable at $0.09 for five years from the date of grant. Both the RSUs and stock options vest on the date of grant.
The RSUs and stock options are subject to approval by regulatory authorities.
Repirinast was originally developed by Mitsubishi Tanabe Pharma (“Mitsubishi”) and was sold and marketed in Japan under the brand name RometTM for the treatment of Asthma. RometTM was marketed for over 25 years in Japan. Mitsubishi discontinued manufacturing and sales of the drug in 2013. Accordingly, Algernon has retained Zhejiang Ausun Pharmaceutical in Zhejiang, China to manufacture a cGMP Repirinast supply.
Mast cells are recruited to sites of cellular damage, and degranulation of mast cells leads to release of a myriad of proinflammatory chemical mediators which lead to tissue damage in a self-propagating cascade. NP-251 binds to receptors on mast cells and prevents their degranulation, which the Company believes could help prevent fibrosis in multiple organ classes including the kidneys.
Algernon Pharmaceuticals is a Canadian clinical stage drug development company investigating multiple drugs for unmet global medical needs. Algernon Pharmaceuticals is also the parent company of a private subsidiary called Algernon NeuroScience, that is advancing a psychedelic program investigating a proprietary form of DMT for stroke and traumatic brain injury.
The content above comes from the network. if any infringement, please contact us to modify.
上市批准专利到期
2024-01-31
VANCOUVER, British Columbia, Jan. 31, 2024 (GLOBE NEWSWIRE) -- Algernon Pharmaceuticals Inc. (the “Company” or “Algernon”) (CSE: AGN) (FRANKFURT: AGW0) (OTCQB: AGNPF) a Canadian clinical stage pharmaceutical development company, is pleased to announce that it has received a notice of intention to grant from the Chinese Patent Office for patent application No. 201980043698.6 for its antifibrotic drug candidate NP-251 (Repirinast). The invention claims the administration of Repirinast, either alone or in combination with telmisartan, for use in the prophylaxis or treatment of renal fibrosis or kidney disease. The base claims of the patent will be valid through 2038, excluding any patent term adjustments or extensions, which may provide additional protection. The Company was recently issued patents for Repirinast in chronic kidney disease (CKD) from Japan and has also filed corresponding patent applications in Canada, Europe and the United States. Repirinast is the Company’s lead candidate for the treatment of CKD based on data showing it reduced fibrosis by 51% with statistical significance and showed an additive benefit to telmisartan in a unilateral ureteral obstruction (UUO) mouse model. Algernon’s intellectual property strategy for its repurposed drug program includes protecting its compounds by filing patent applications including method of use, dosing and formulations, and for new composition of matter patents based on novel salt forms. “We are once again pleased to receive another notice of intent to grant a patent from the Chinese Patent Office,” said Christopher J. Moreau CEO of Algernon Pharmaceuticals. “This will be the second patent granted to the Company in China, and it complements multiple other patents issued in other major world markets as part of our comprehensive global intellectual property strategy.” The Company also announces a grant of 1,625,000 restricted share units pursuant to its RSU Plan (each, an "RSU") to executives and directors, of the Company. Each RSU entitles the recipient to receive one common share of the Company or a cash payment equal to the equivalent of one common share of the Company on vesting. The RSUs vest on the date of grant. The Company has also granted 100,000 stock options exercisable at $0.075 for two years from date of grant, to a consultant of the Company. The stock options vest on the date of grant. The RSUs and stock options are subject to approval by regulatory authorities. About Algernon Pharmaceuticals Algernon Pharmaceuticals is a Canadian clinical stage drug development company investigating multiple drugs for unmet global medical needs. Algernon Pharmaceuticals has active research programs for chronic cough, chronic kidney disease and NASH, and is the parent company of a private subsidiary called Algernon NeuroScience, that is advancing a psychedelic program investigating a proprietary form of psychedelic DMT for stroke and traumatic brain injury. CONTACT INFORMATION Christopher J. MoreauCEOAlgernon Pharmaceuticals Inc.604-398-4175 ext 701 info@algernonpharmaceuticals.cominvestors@algernonpharmaceuticals.comwww.algernonpharmaceuticals.com Neither the Canadian Securities Exchange nor its Market Regulator (as that term is defined in the policies of the Canadian Securities Exchange) accepts responsibility for the adequacy or accuracy of this release.
CAUTIONARY DISCLAIMER STATEMENT: No Securities Exchange has reviewed nor accepts responsibility for the adequacy or accuracy of the content of this news release. This news release contains forward-looking statements relating to product development, licensing, commercialization and regulatory compliance issues and other statements that are not historical facts. Forward-looking statements are often identified by terms such as “will”, “may”, “should”, “anticipate”, “expects” and similar expressions. All statements other than statements of historical fact, included in this release are forward-looking statements that involve risks and uncertainties. There can be no assurance that such statements will prove to be accurate and actual results and future events could differ materially from those anticipated in such statements. Important factors that could cause actual results to differ materially from the Company’s expectations include the failure to satisfy the conditions of the relevant securities exchange(s) and other risks detailed from time to time in the filings made by the Company with securities regulations. The reader is cautioned that assumptions used in the preparation of any forward-looking information may prove to be incorrect. Events or circumstances may cause actual results to differ materially from those predicted, as a result of numerous known and unknown risks, uncertainties, and other factors, many of which are beyond the control of the Company. The reader is cautioned not to place undue reliance on any forward-looking information. Such information, although considered reasonable by management at the time of preparation, may prove to be incorrect and actual results may differ materially from those anticipated. Forward-looking statements contained in this news release are expressly qualified by this cautionary statement. The forward-looking statements contained in this news release are made as of the date of this news release and the Company will update or revise publicly any of the included forward-looking statements as expressly required by applicable law.
专利到期
100 项与 瑞吡司特 相关的药物交易
登录后查看更多信息
研发状态
批准上市
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| 哮喘 | 日本 | 1987-01-01 |
未上市
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 慢性肾病 | 临床前 | 尼日利亚 | 2024-04-09 | |
| 代谢功能障碍相关脂肪性肝炎 | 临床前 | 加拿大 | 2023-11-30 | |
| 特发性肺纤维化 | 临床前 | 加拿大 | - | |
| 终末期肾脏病 | 临床前 | 加拿大 | - |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
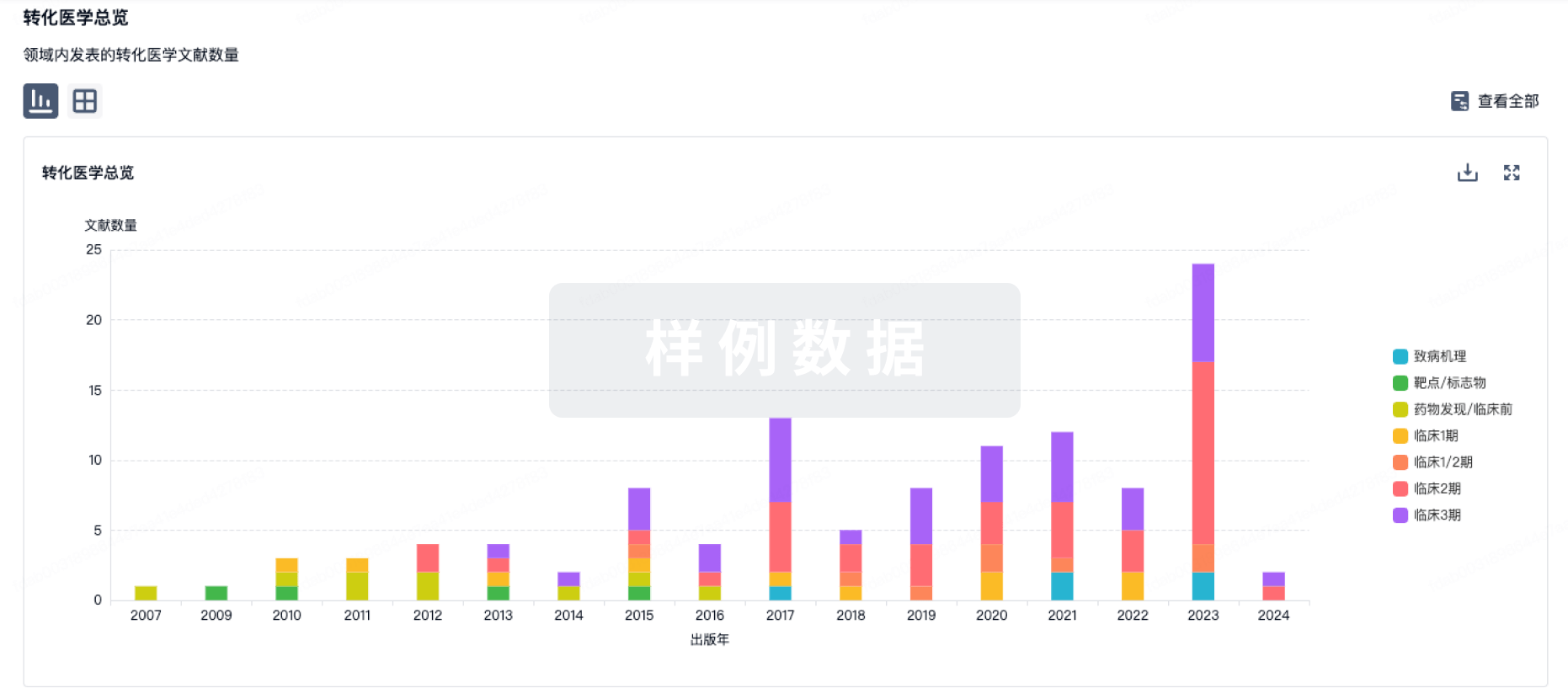
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
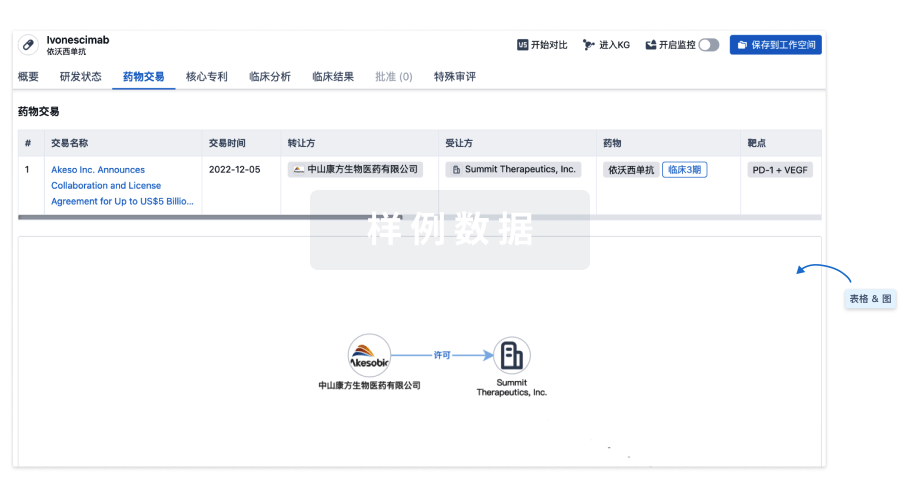
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
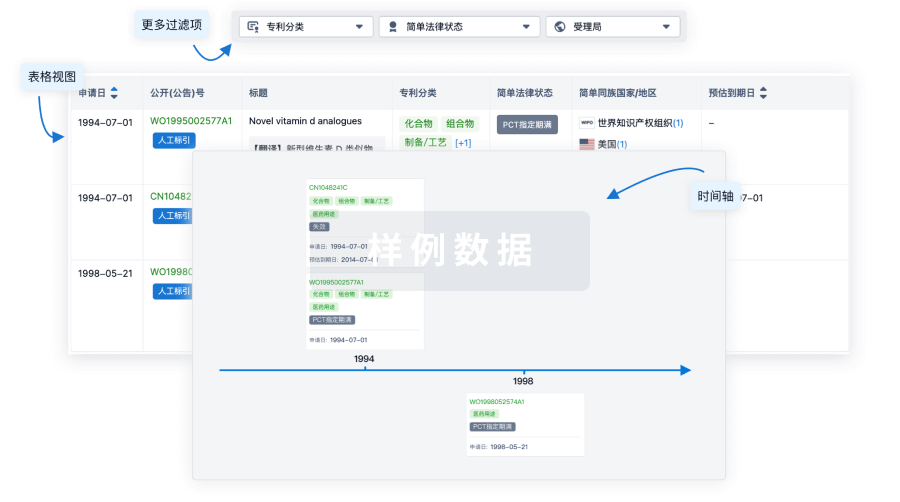
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
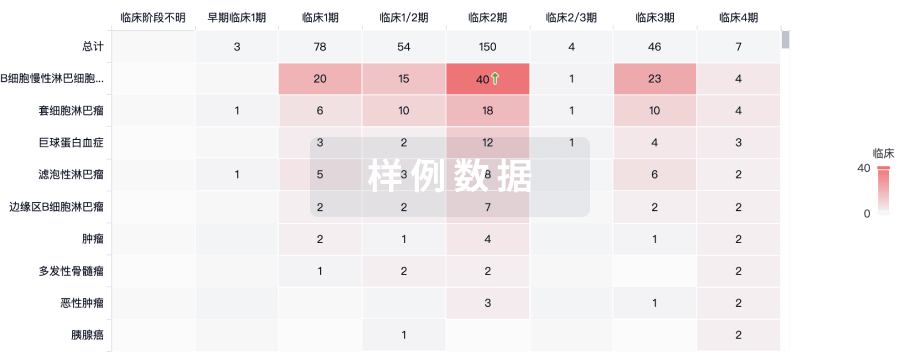
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
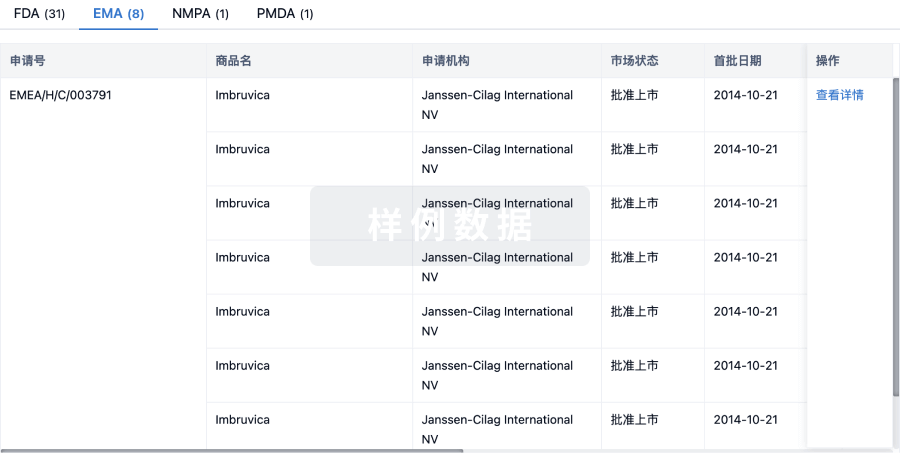
批准
利用最新的监管批准信息加速您的研究。
登录
或
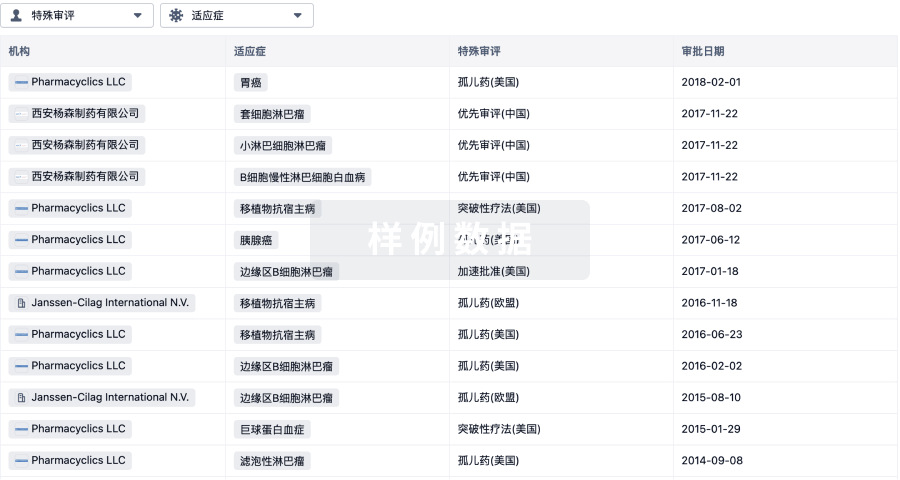
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用




