预约演示
更新于:2026-02-26
Vidarabine
阿糖腺苷
更新于:2026-02-26
概要
基本信息
权益机构- |
最高研发阶段批准上市 |
首次获批日期 美国 (1976-11-26), |
最高研发阶段(中国)批准上市 |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C10H15N5O5 |
InChIKeyZTHWFVSEMLMLKT-CAMOTBBTSA-N |
CAS号24356-66-9 |
关联
2
项与 阿糖腺苷 相关的临床试验JPRN-UMIN000006026
Effects of anti-viral drug (vidarabine) on atrial fibrillation and ventricular arrhythmia - Treatment of arrhythmia by vidarabine
开始日期2011-08-01 |
申办/合作机构- |
NCT00000985
Treatment of Acyclovir-Resistant Mucocutaneous Herpes Simplex Infection in Patients With the Acquired Immunodeficiency Syndrome: A Randomized Multicenter Study of Foscarnet Versus Vidarabine
To compare the safety and effectiveness of foscarnet and vidarabine treatments for AIDS patients who have herpes simplex virus infections that are resistant to standard treatment with acyclovir.
Foscarnet is a drug that inhibits viruses and has been shown to be effective against infection with Cytomegalovirus and also against infection with the Herpes simplex virus in several patients with AIDS. Vidarabine has been shown to have activity against the Herpes simplex virus in patients who do not have AIDS, but it has not been studied in patients who do have AIDS. This study compares foscarnet and vidarabine treatments for AIDS patients who have herpes simplex infection that has not responded to therapy with acyclovir in the hope that one of these two drugs will help to stop further progression of the herpes simplex infection and may have fewer side effects.
Foscarnet is a drug that inhibits viruses and has been shown to be effective against infection with Cytomegalovirus and also against infection with the Herpes simplex virus in several patients with AIDS. Vidarabine has been shown to have activity against the Herpes simplex virus in patients who do not have AIDS, but it has not been studied in patients who do have AIDS. This study compares foscarnet and vidarabine treatments for AIDS patients who have herpes simplex infection that has not responded to therapy with acyclovir in the hope that one of these two drugs will help to stop further progression of the herpes simplex infection and may have fewer side effects.
100 项与 阿糖腺苷 相关的临床结果
登录后查看更多信息
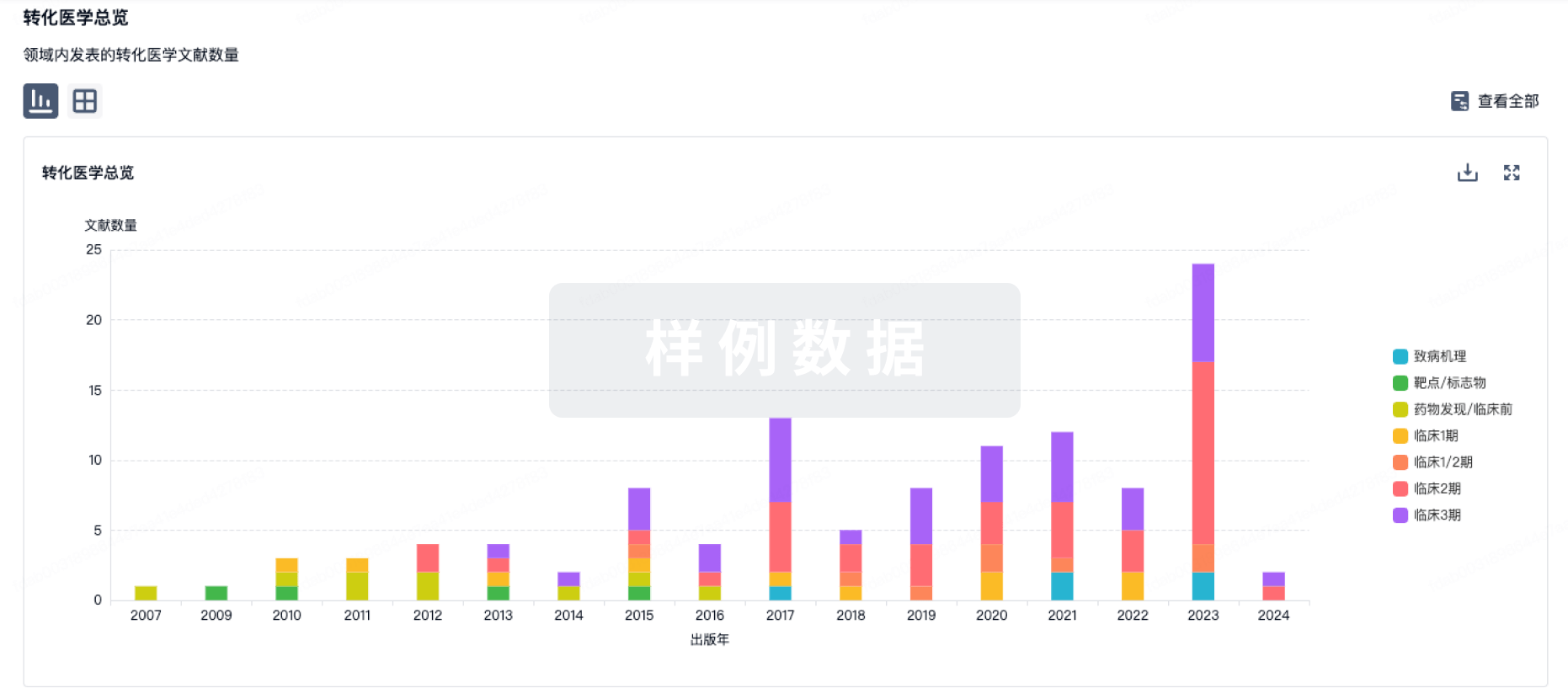
100 项与 阿糖腺苷 相关的转化医学
登录后查看更多信息
100 项与 阿糖腺苷 相关的专利(医药)
登录后查看更多信息
1,612
项与 阿糖腺苷 相关的文献(医药)2025-11-01·Food Science & Nutrition
From Wild Vegetable to Renal Protector: The Therapeutic Potential of
Aralia elata
(Miq.) Seem
Article
作者: He, Weiming ; Tan, Huifeng ; Liu, Yue ; Du, Zhenfang ; Xu, Ziyun ; Huang, Min ; Wei, Minggang ; Qiang, Sheng ; Wu, Jingyi ; Jiang, Chunbo
ABSTRACT:
Aralia elata
(Miq.) Seem. (
A. elata
) is a medicinal and edible wild vegetable, which has potential applications in the pharmaceutical and health food industries. Araloside A (Ara A) is a natural triterpenoid saponin extracted from
A. elata
. Chronic kidney disease (CKD) is an important public health problem worldwide. There is still a lack of effective drugs to delay the progression of renal injury. We experimentally found that Ara A can alleviate pathological renal injury, reduce proteinuria and improve renal dysfunction in model mice. This is associated with its ability to mitigate oxidative stress and regulate ferroptosis‐related proteins. In vitro experiments, we found that Ara A can inhibit ferroptosis in podocytes and protect functional proteins. Mechanically, Ara A alleviated iron accumulation and lipid peroxidation in vivo and in vitro, and played a renal protective role by influencing the solute carrier family 7a member 11 (SLC7A11)/glutathione (GSH)/glutathione peroxidase 4 (GPX4) signaling pathway. Through cell and animal experiments, we have verified that Ara A, a component derived from
A. elata
, can delay glomerulosclerosis, with the inhibition of oxidative stress and ferroptosis as its potential mechanisms of action. These results provide evidence for the utilization of
A. elata
as a nutraceutical for the treatment of CKD.
2025-02-01·FOOD RESEARCH INTERNATIONAL
Comprehensive metabolite profiling reveals the dynamic changes of volatile and non-volatile metabolites in albino tea cultivar ‘Ming guan’ (MG) during white tea withering process
Article
作者: Chen, Changsong ; Zhang, Hui ; Wang, Xiuping ; Zhong, Qiusheng ; Zhang, Yinggen ; Chen, Quanbin ; Huang, Ting
'Ming guan'(MG), an elite albino cultivar deriving from the progeny of the traditional albino cultivar 'Bai jiguan', is a promising candidate for white tea production due to its favorable amino acid to phenol ratio. In this study, a comprehensive metabolomics analysis using ultra-high performance liquid chromatography tandem mass spectrometry (UHPLC-MS/MS) and headspace solid-phase microextraction-gas chromatography mass spectrometry (HS-SPME-GC-MS) were conducted to reveal the dynamic changes of non-volatile and volatile organic compounds (VOCs) throughout the withering processing of MG white tea. Meanwhile, multivariate statistical analyses were applied to screen for the characteristic components in the flavor and aroma of MG white tea. A total of 625 non-volatile metabolites and 118 VOCs were determined, of which 90 non-volatile metabolites (VIP ≥ 1, FC ≥ 2 or ≤ 0.5) were identified as key flavor components significantly changed throughout the withering process. The relative odor activity value (ROAV) analysis highlighted 22 VOCs (ROAV ≥ 1) with substantial effect on aroma formation, of which geraniol, (E)-2-hexenal, 4-methoxy-benzaldehyde and guaiacol emerging as the most key aroma constituents of MG white tea, endowing MG white tea with fruity and floral odor notes. This study offered a comprehensive investigation into metabolite changes in MG white tea, contributing valuable insights for the innovation of new white tea products utilizing albino tea plant mutants.
2025-01-01·JOURNAL OF ETHNOPHARMACOLOGY
Yanghe decoction inhibits inflammation-induced lung metastasis of colorectal cancer
Article
作者: Ding, Kai ; Liu, Songyu ; Zhang, Lu ; Li, Bo ; Zhou, Jinyi ; Su, Xiaosan ; Li, Jv ; Zeng, Bin ; Sun, Ruifen ; Wang, Junliang
ETHNOPHARMACOLOGICAL RELEVANCE:
Positive deficiency and cancer toxicity are the main pathogenesis of colorectal cancer (CRC) lung metastasis. Yanghe decoction (YHD), a traditional Chinese medicine, has the effects of warming yang, tonifying blood, dispersing cold and clearing stagnation, adopting a treatment method that combines supporting the right and dispelling the wrong, which has remarkable efficacy in anti-tumor.Although, its precise mechanism of inhibiting the metastasis of colorectal cancer to the lung is still poorly understood.
AIM OF THE STUDY:
This study aimed to elucidate the antitumor properties of YHD within the context of colorectal cancer lung metastasis.
MATERIALS AND METHODS:
Ultrahigh-performance liquid chromatography coupled with mass spectrometry (UHPLC-MS) was utilized to analyze the chemical composition of YHD. The anticancer activity of YHD was evaluated in a CRC lung metastasis mouse model by quantifying pulmonary metastatic nodules. The effects of YHD on CRC cell proliferation, apoptosis, cell cycle progression, and invasion were assessed using CCK-8 assays, flow cytometry, and Transwell assays. YHD-mediated immune modulation in tumor-bearing mice was evaluated by analyzing antitumor immunity, immunosuppressive cells, and cytokines in peripheral blood and tumor tissue. Gut microbiota analysis was conducted to determine the impact of YHD on the gut microbiota in mice.
RESULTS:
Our analysis identified 1801 chemical markers in YHD. CFA exacerbated lung metastasis in CRC, whereas oral administration of YHD significantly mitigated this effect, as evidenced by the reduced number of metastatic lung nodules in CRC tumor-bearing mice. In vitro experiments demonstrated that YHD inhibits CRC cell proliferation, induces apoptosis, and suppresses invasion. In the lung tissues of mice with CRC metastasis treated with CFA, there was a significant reduction in NK cells and IL-21, along with an increase in M2 macrophages and IL-6. Following YHD treatment, there was a notable increase in NK cells and IL-21, accompanied by a decrease in M2 macrophages and IL-6 in lung tissues. YHD administration was also associated with an increase in beneficial bacterial species such as Bacillus and a decrease in deleterious bacterial species such as Oscillibacter.
CONCLUSION:
Our findings demonstrate that YHD inhibits lung metastasis in CRC by suppressing CRC cell proliferation and invasion, in addition to modulating the tumor microenvironment to favor antitumor immunity. These results provide a scientific basis for the clinical application of YHD in the treatment of CRC patients.
26
项与 阿糖腺苷 相关的新闻(医药)2026-02-04
·重良社
破产重整动态
Bankruptcy Reorganization
2月3日晚间*ST景峰(300091)发布公告,其重整计划获得法院裁定批准,该上市公司进入重整计划执行阶段。
当天,*ST景峰收到湖南省常德市中级人民法院(以下简称“常德中院”)送达的《民事裁定书》,常德中院裁定批准*ST景峰《重整计划》。
2025年10月21日,公司收到常德中院送达的《民事裁定书》和《决定书》,裁定受理彭东钜、上海鑫绰投资管理有限公司对公司的重整申请,并指定北京市中伦律师事务所担任公司管理人。
2025年10月23日和 2025 年 11 月 21 日,公司分别披露了《关于公司重整债权申报通知及召开债权人会议的公告》和《关于公司召开第一次债权人会议的提示性公告》。
2025年12月3日,景峰医药召开了第一次债权人会议。 2026年1 月29日,《重整计划(草案)》《重整计划(草案)之出资人权益调整方案》已分别经公司第二次债权人会议和出资人组会议审议通过。
权益调整
鉴于景峰医药已不能清偿到期债务,且明显缺乏清偿能力,生产经营和财务状况均已陷入困境。同时,普通债权人不能在重整计划中全额获得清偿。因此,本重整计划安排对出资人权益进行调整。景峰医药在确定出资人权益调整方案时主要考虑了如下因素:
1.本重整计划偿债方案中所需的偿债资源。
2.景峰医药经营方案中涉及未来经营恢复、业务升级、内部管理系统改造升级等所需的资金。
3.充分考虑中小投资者利益,避免过度转增导致对中小投资者权益的稀释。
综合上述因素,本次出资人权益调整以景峰医药现有总股本为基础,按照每 10 股转增10 股的比例实施资本公积转增股本。
本次权益调整以景峰医药现有总股本879,774,351股为基数,按照每 10 股转增 10 股的比例实施资本公积金转增股本,共计可转增 879,774,351 股股票(最终实际转增的股票数量以中证登深圳分公司实际登记确认的数量为准)。转增后,景峰医药总股本将增至 1,759,548,702 股。
上述转增股票不向原股东分配,全部由重整投资人支付现金对价受让,具体受让股票数以后续重整计划执行阶段的司法协助执行通知书载明的内容及中证登深圳分公司实际登记确认的数量为准。
债权清偿
1.职工债权
职工债权由景峰医药在法院裁定批准重整计划且债权经法院裁定确认后 20 个工作日内以现金方式一次性清偿完毕。
2.税款债权
税款债权由景峰医药在法院裁定批准重整计划且债权经法院裁定确认后 20 个工作日内以现金方式一次性清偿完毕。
3.普通债权
普通债权以债权人为单位,每家普通债权人的普通债权在 200 万元以下(含 200 万元)的部分,由景峰医药在后续重整计划执行期限内以现金方式一次性清偿完毕。每家普通债权人的普通债权超过200 万元的部分,可选择如下方案进行受偿:
(1)留债方案
针对每家普通债权人的普通债权超过200 万元的部分,以现金方式予以留债清偿,留债最长期限为2 年,留债期限内不计息。留债期限内,不影响债权人根据有关合同约定以及法律规定,就其债权未获清偿部分(含合同约定或法定的 26 利息)向其他债务人(包括但不限于主债务人)主张有关权益,亦不影响其他担保人或主债务人根据有关合同约定以及法律规定向债权人清偿债务。
(2)现金清偿方案
每家普通债权人按 90%的清偿率在法院裁定批准重整计划之日起 90 日内获得现金清偿;剩余未获清偿债权部分在重整计划执行完毕后豁免清偿,景峰医药不再承担清偿责任。
债权金额超过 200 万元的普通债权人(含暂缓确认债权人),应自法院裁定批准重整计划之日起10 个工作日内按照重整计划要求的书面格式提供债权清偿方式选择告知书。债权人逾期未告知的,视为全额选择按以普通债权清偿方式中的“现金清偿方案”获得清偿。
经营计划
基于“优化存量,培育增量”的未来发展思路,景峰医药未来业务板块将包括中成药为主、化药为辅的存量业务,以及生物医药为主的增量业务,积极拓展合成生物产业。具体发展举措如下:
(一)优化存量,培育增量,创新引领、双轮驱动
1.聚焦存量优势产品,提升营销能力,创新创研新产品
重整完成后,景峰医药将启动生产设备和生产过程的革新、研发方向与产业趋势的融合,提升管理水平,重塑品牌优势,打造具有国际先进水平的重点产业化项目。具体举措如下:
(1)优化产品管线,聚焦存量核心
产品公司产品种类丰富,未来将逐步淘汰长期缺乏竞争力的品种,按照潜力和目前的产品结构,公司将聚焦心血管领域产品,并持续不断优化产品结构。具体如下:
一是公司将成立心血管销售部,聚焦心血管产品在等级医院的终端拓展、零售终端拓展及学术推广工作。心脑宁胶囊、参芎葡萄糖注射液、乐脉丸的等级医院渠道划归心血管销售部管理和销售;心脑宁和参芎除了稳定现有代理商的终端和纯销以外,要重点提升全国top100 医院的学术推广覆盖和纯销提升速度,另一方面要积极拓展空白市场和代理商广度,设立终端开发目标进行考核,在精细化招商的指引下逐步提升终端数量和纯销数量,为未来销售夯实基础。公司将推动参芎葡萄糖注射液纳入国家医保目录作为景峰医药重点工作任务。
二是对于重点产品心脑宁胶囊(全国独家),目前为医保乙类药品,未来拟研发增加其适应症(针对精准干预急性冠状动脉综合征后状态合并颈动脉斑块残余风险研究),加速完成药品再评价、逐步提升药品价值并申请进入基药目录。
三是重启参芎葡萄糖注射液三期临床相关研究,扩大产品适应症重新申报国家医保药品目录,同时加速金鸡丸、乐脉丸等中药产品的多元发展;对于通过一致性评价的品种,深度挖掘销售价值,增加产品的中标概率。
四是完善升级生产线,拓展玻璃酸钠制剂产品(包括玻璃酸钠美容填充项目和水光针项目)产能。淘汰生产端陈旧设备,改善生产自动化水平,通过委托加工等多种方式,高效解决工厂产能不足现状,保证旺季销售供应。
五是围绕榄香烯开展系统性研究,开展质量标准提升研究,并推进提取后副产物的资源化利用;在此基础上,开发榄香烯系列衍生物及相关制剂,拓展其在抗肿瘤及其他疾病治疗领域的应用范围;同时,通过优化提取工艺,降低生产成本,提高资源利用效率,最终构建从原料到制剂的全产业链价值提升路径。
(2)拓展销售渠道,提升营销能力
一是调整营销模式。将由各子公司独立经营销售转变为母公司“统一规划,统一销售,统一管理,统一团队”的四统一销售模式,促进存量产品发展。按照产品治疗领域和销售渠道的属性进行划分形成条线管理,通过组织架构的调整,使销售队伍在执行力、专业度上快速提升。打造心脑血管、抗肿瘤等领域专业化的营销团队。
二是继续拓展基层及民营医疗机构等三终端,提升产品销售和市场覆盖比例。将参芎葡萄糖注射液、乐脉丸、金鸡丸、通迪胶囊、妇平胶囊、儿童回春颗粒、注射用克林霉素磷酸酯、阿糖腺苷、奥美拉唑等列为发力品种,参芎重点拓展代理商数量,填补空白市场,加强代理商管理,要求更多市场覆盖和纯销提升;梳理、整合其他产品的三终端渠道和代理商现状,积极开发新的市场机会,做好代理商选择及优化,尽快完成三终端销售的布局和价值体现;普药部分做好现有市场覆盖的维护及提升,利用积极有效的市场调研和价格杠杆,努力培育 5-10 个战略合作伙伴,加快提升普药的市场份额和销售金额。
三是规划集采药品销售方案。成立集采销售部,规划仿制药布局,积极探索和参加国家集采及省际联盟集采,力争更多产品进入国家集采通道和中选更多省份的集采招标。其中,重点推进海南锦瑞公司仿制药一致性评价的过审,以及玻璃酸钠注射液重点开发上量。另外,市场准入团队积极学习和解读集采政策,结合公司产品结构,响应国家集采号召,积极参与、力争中选,统一配送、跟进销售。后续公司将更加关注多家中选省份的销售策略制定和备选省份的销售拓展计划。
四是建立和壮大零售队伍,打造公司销售业务新增长点。零售连锁市场逐年增长,公司将建立专门的零售队伍,通过外部引进和内部培养的方式快速建立零售专业销售团队,产品聚焦心脑宁胶囊、镇痛活络酊、冰栀伤痛气雾剂、乐脉丸、金鸡丸等产品;全国 top10 连锁力争直接合作或者贴牌销售,其他连锁选择专业的代理商合作,快速上架,专业推广,迅速提升销售,为后续销售增长带来强劲动力。
(3)创新创研新产品,实现产业升级
借鉴产业投资人的研发创新优势,公司未来医药研发将以“建立完整、高效、国内领先的研究开发系统,制定并持续更新科学、稳健的产品线规划”为目标,建立健全与国际接轨的研发质量管理体系,加强建设研发项目管理体系。
一是加快推进化药新药项目落地,NMPA1类新药交联玻璃酸钠注射液根据 III 期临床CED 沟通会的要求进行补充研究,推动完成 III 期临床试验样品的生产与工艺验证,并启动 III 期临床研究。
二是推动普通注射剂项目完成一致性评价及工艺验证, 推进玻璃酸钠注射液、注射用氯诺昔康开展一致性评价工作。
三是持续推进抗肿瘤产品研发,以多种方式解决抗肿瘤药物伊立替康注射液、注射用吉西他滨、注射用异环磷酰胺等产品的产能不足及产销问题。四是公司在产品立项上将继续保持谨慎态度,依据自身研发优势及核心竞争力,从多维度进行产品评估,结合未来产品市场竞争态势的预测,以及未来销售量等市场数据,确定研发方向,推进临床研究以及里程碑的逐步实现,以实现产品上市后的收益。
2.培育生物医药第二增长曲线
(1)常德市政策支持生物医药产业,已形成产业集聚生物制药产业在常德市拥有深厚的产业基础。常德在生物制造领域底蕴深厚,拥有亚洲最大的甾体原料药和中间体生产出口基地、全国最大的酶制剂生产出口基地、全省第一方阵的生物医药企业、全省规模领先的食品加工产业。目前,常德市已建成国家级合成生物制造产业创新平台1个、省级创新平台 5 个、省级科技成果转化中试基地1个。常德经开区和津市市的合成生物中试转化基地即将投入使用,安乡县合成生物产业应用研发中心已经建成。常德市和湖南文理学院共建合成生物学公共技术平台,一体布局生物合成、分子检测、生物提取、生物发酵和生物医学等高能级实验室,争创合成生物省级、国家级重点实验室。生物制药产业在常德市拥有完备的园区配套支撑。按照“一中心三基地”产业布局,津市市、常德经开区产业承载 44 优势,安乡县原料资源优势,全力打造全省引领、全国有名的常德“生物制造谷”,推动合成生物制造产业高端化、绿色化、融合化、协同化发展。
(2)石药控股及景峰医药在生物制药领域拥有深厚积累 石药控股在生物制药领域拥有深厚积累,研发实力雄厚。石药控股及其下属企业依托纳米制剂药物、细胞治疗、长效注射剂等八大技术平台,聚焦肿瘤、精神神经、心血管等领域,并在中药和合成生物学等领域有所布局。景峰医药研发中心已分别打造生物药开发技术平台、大分子交联技术平台、纳米分散系DDS平台、MUPS(MultipleUnitPellet System)OSD 平台、预灌封制剂产业化平台等研发平台,致力于建立高技术壁垒的生物药品研发与生产平台,为拓展生物制药提供研发基础保障。
(3)依托区位优势及企业积累,培育第二增长曲线重整完成后,景峰医药将依托常德市对生物医药行业的政策支持、资金保障、产业集聚,并凭借石药控股和公司在生物医药行业的积累与研发实力,着力培育生物医药作为第二增长曲线,拓展合成生物产品及关联赛道,逐步成长为双轮驱动的优质公司。具体举措包括但不限于:
一是聚焦合成生物学与创新药械,推动具体项目落地。公司将重点引入与现有业务具有战略协同效应的优质合成生物学项目,在常德逐步构建从研发、中试到产业化的合成生物制造产业链,为业绩贡献新增量。
二是联合设立专项产业基金,构建项目孵化与引入平台。该基金将借助石药控股的产业资源和基金管理经验,主要投向生物医药及相关联的高成长性领域,在追求财务回报的同时,积极引导被投企业将研发成果和生产基地落户常德,形成“基金投资—项目孵化—产业落地”的良性循环。
三是整合内外部生物医药资源,打造“研发-生产-销售”一体化生态。推动石药控股与景峰医药的生物医药协同研发项目在常德优先转化及生产落地,并借助石药控股整合两湖两广销售资源的优势,在常德设立销售总部统一管理,从而全面强化常德总部的职能,打造“研发-生产-销售”的生物医药全链路,实现第二曲线扎实增长。
(二)优化组织架构,提升管理水平,降本增效
借鉴产业投资人丰富的企业管理经验,景峰医药将进一步优化组织运营管理架构,强化企业管理,加强景峰医药及子公司在战略、财务、人才、业务等方面的规范运作管理,不断提升内控管理体系运行质量,通过建立和运行常态化的管理提升机制,降低管理成本及费用,推进和保障公司可持续发展。具体如下:
1. 优化组织架构,强化总部管控
一是公司将继续严格执行《公司法》、《证券法》及相关规定,严格履行股东会及董事会的监督指导作用,充分发挥独立董事的风险把控作用。二是重构组织架构,通过销售集中、采购集中、财务集中三集中模式,实现扁平化管理。强化母公司管控力度,梳理人员配置,定岗定编,先推动实现财务专业和营销专业纳入母公司的集中体系化管理,再逐步拓展至生产、研发、采购等专业管理;发挥母公司计划、调度、考核等管理职能,合理科学调配母公司内部资源,实现公司整体利益最大化。
2. 完善管理制度,提升管理水平
一是优化业务流程。加强制度和流程的执行管理,提高组织运营效率;完善预算管理体系与制度,实现管理的横向到边、纵向到底的全覆盖;完善健全投资管理制度,以理性的、科学的投资决策和管理制度引领企业稳健增长;建立高效管理机制,搭建周报、半月会和经营分析会等方式,确保重点工作、核心指标的达成。
二是强化财务风险管控。包括重点加强资金审批控制、完善内部会计稽核制度、严格执行预算和收支管理等;公司将合理运用法规及制度筹措资金,并提高现金周转率和回款速度、降低回款周期;将安排工作组加大对应收账款的清欠回收力度,进一步加强应收账款管理,加速资金回笼以缓解资金压力。
三是制定人才发展战略。公司将全面梳理人员配置,完善薪酬考核体系,在培养人才的同时留住人才。公司将围绕经营目标,落地预算管理,加强指标与绩效考核的管理,分配机制重点向业务线、市场线倾斜,激发员工的工作热情,强化以结果为导向的考核机制,实行内部竞争上岗制和轮岗制,激发内部活力。
3. 加强成本管控,盘活低效资产
一是落地成本领先战略。公司将持续抓好生产组织和原料采购,降低运营成本并认真做好开源节流、增收节支工作。
二是盘活低效资产,提升资产运营效率,降低资产运营压力。公司将继续对低效业务、低效资产与闲置资产进行重整与盘活,充分提高资产利用率、提高存货周转率,不断强化主营业务,提升经营现金流。
-END-
|素材来源:公司公告
推荐阅读
原正部级李微微,无期徒刑!
大润发母公司高鑫零售:公司暂时无法与执行董事兼首席执行官李卫平取得联系
突发!百亿量化私募创始合伙人沈显兵去世,年仅40岁
坏报 | A股一夜超30家可能终止上市!
【擎天须重器 济世待良臣】
END
声明
本微信公众平台致力于好文推送,欢迎投稿,所发表内容注明来源,版权归原出处所有(无法查证版权或未注明出处均来源于网络搜集)。转载内容只以信息传播为目的,仅供学习交流参考,不代表本平台认同其观点和立场,内容的真实性、准确性和合法性由原作者负责。如转载涉及版权等问题,请发消息至公众号后台与我们联系,我们将在第一时间处理,非常感谢。
关于我们
关注特殊资产领域投资
高管变更
2026-02-03
·中财网
原标题:*ST景峰:湖南景峰医药股份有限公司重整计划
湖南景峰医药股份有限公司
重整计划
湖南景峰医药股份有限公司
二〇二六年一月
目录
释义..............................................................................................5前言............................................................................................10摘要............................................................................................12正文............................................................................................14一、景峰医药基本情况...........................................................14(一)基本信息.......................................................................14(二)股本结构.......................................................................14(三)预重整及重整的申请及受理情况...............................15(四)资产情况.......................................................................16(五)负债情况.......................................................................17(六)偿债能力分析...............................................................18二、出资人权益调整方案.......................................................19(一)出资人权益调整的必要性、合理性...........................19(二)出资人权益调整的范围...............................................20(三)出资人权益调整的内容...............................................21(四)除权与除息...................................................................21(五)出资人权益调整方案的执行效果...............................22三、债权分类及调整方案.......................................................23(一)债权分类方案...............................................................23(二)债权调整方案...............................................................23四、债权清偿方案...................................................................25(一)偿债资源.......................................................................25(二)债权清偿方案...............................................................25(三)劣后债权清偿方案.......................................................26(四)合并报表范围内关联债权的清偿方案.......................26(五)预计债权清偿方案.......................................................26五、重整投资人情况...............................................................28(一)重整投资人的招募情况...............................................28(二)重整投资人具体情况...................................................29(三)重整投资人受让转增股份情况...................................37六、经营方案............................................................................39(一)优化存量,培育增量,创新引领、双轮驱动...........39
(二)优化组织架构,提升管理水平,降本增效...............45
七、重整计划(草案)的表决和批准...................................47(一)重整计划(草案)的表决...........................................47(二)重整计划(草案)的批准...........................................48(三)重整计划批准生效.......................................................48(四)未获批准的后果...........................................................48八、重整计划的执行...............................................................49(一)执行期限.......................................................................49(二)执行期限的延长与提前...............................................49(三)执行完毕的标准...........................................................49(四)司法协助执行事项.......................................................49(五)执行完毕的效力...........................................................50(六)重整计划的变更...........................................................50(七)有关重整计划执行的重大不确定事项.......................51九、重整计划执行的监督.......................................................51(一)监督期限.......................................................................51(二)监督期限的延长与提前...............................................51(三)监督期限的终止...........................................................51十、其他说明情况...................................................................52(一)重整计划的生效...........................................................52(二)偿债资源的提存及预留、处理...................................52(三)偿债资源的分配...........................................................53(四)财产保全措施的解除及信用等级的修复...................54(五)债权人对其他还款义务人的权利...............................54(六)对实控人原有股权调整的方案...................................55(七)破产费用的支付及共益债务的清偿...........................55(八)转让债权的清偿...........................................................56(九)重整计划的解释...........................................................56释义
景峰医药/公司/债务
人湖南景峰医药股份有限公司常德中院湖南省常德市中级人民法院深交所深圳证券交易所中证登深圳分公司中国证券登记结算有限责任公司深
圳分公司临时管理人、管理人由常德中院指定的景峰医药临时管
理人、管理人债权人符合《企业破产法》第四十四条之
规定的,景峰医药的某个、部分或
全体债权人出资人出资人组会议召开通知中所载明的
股权登记日在中证登深圳分公司登
记在册的景峰医药股东重整投资人经公开招募后确定的,与景峰医药
管理人、景峰医药签署《重整投资
协议》和/或《重整投资协议之补充
协议》的主体石药控股石药控股集团有限公司德源招商常德市德源招商投资有限公司重整计划(草案)依据《企业破产法》第七十九条之 规定,债务人制作并提交的重整计
划(草案)重整计划重整计划(草案)获债权人会议及
出资人组表决通过,或虽未获表决
通过但经常德中院裁定批准后的重
整计划破产费用《企业破产法》第四十一条规定的
人民法院受理破产申请后发生的相
关费用共益债务《企业破产法》第四十二条规定的
人民法院受理破产申请后发生的相
关债务有财产担保债权依据《企业破产法》第八十二条第
一款第(一)项之规定,对景峰医
药特定财产享有担保权的债权职工债权依据《企业破产法》第八十二条第
一款第(二)项等相关规定,包括
景峰医药所欠职工的工资和医疗、
伤残补助、抚恤费用,所欠的应当
划入职工个人账户的基本养老保
险、基本医疗保险费用,法律、行
政法规规定应当支付给职工的补偿
金普通债权依据《企业破产法》第八十二条第
一款第(四)项之规定,债权人对
景峰医药享有的债权审查确定的债权债权申报期限内经债权人申报并经
管理人依法审查确定的债权未申报债权与债务人构成债权债务关系,未在
重整计划(草案)提交债权人会议
表决前向管理人依法申报但可能受
法律保护的债权暂缓确定债权已向管理人申报但截至重整计划
(草案)提交之日,因诉讼未决、
需要补充证据材料、债权人提出异
议等原因尚未经管理人审查确定的
债权转增股票根据本重整计划规定的出资人权益
调整方案,以景峰医药股本
879,774,351股为基数,实施资本公
积转增股本形成的股票提存对应偿债资源按照本重整计划规定
暂支付至管理人或管理人指定方名
下,视为提存完毕评估机构为景峰医药重整提供资产评估和偿
债能力分析服务的北京中企华资产
评估有限责任公司审计机构为景峰医药重整提供审计服务的大
信会计师事务所(特殊普通合伙)《偿债能力分析报
告》评估机构以2025年9月30日为基
准日出具的《湖南景峰医药股份有
限公司重整项目偿债能力分析报
告》(中企华评咨字(2025)第5769号《资产清算价值报
告》评估机构以2025年9月30日为基
准日出具的《湖南景峰医药股份有 限公司重整涉及的资产清算价值项
目评估咨询报告》(中企华评咨字
(2025)第5768号)《资产市场价值报
告》评估机构以2025年9月30日为基
准日出具的《湖南景峰医药股份有
限公司重整涉及的资产市场价值项
目资产评估报告》(中企华评报字
(2025)第5766号)《审计报告》审计机构以2025年9月30日为基
准日出具的《湖南景峰医药股份有
限公司审计报告》(大信专审字
[2025]第39-00008号)重整计划的通过依据《企业破产法》第八十六条第
一款之规定,债权人会议各表决组
及出资人组会议均通过重整计划
(草案)时,重整计划即为通过重整计划的批准依据《企业破产法》第八十六条第
二款或第八十七条第三款之规定,
重整计划获得常德中院裁定批准重整计划的执行期
限根据《企业破产法》第八十一条第
(五)项之规定,在景峰医药重整
计划中载明的执行期限及法院裁定
延长的重整计划执行期限重整计划执行监督
期限依据《企业破产法》第九十条之规
定,本重整计划规定的管理人监督
重整计划执行的期限《企业破产法》自2007年6月1日起施行的《中华
人民共和国企业破产法》元本重整计划中除特别注明外,均为
人民币元前言
景峰医药为深交所上市公司,主营业务为以自有资产进
行医药、医疗项目投资;生物制药技术项目的研发与投资;
商品进出口贸易;企业管理咨询、医疗医药研发技术咨询(依
法须经批准的项目,经相关部门批准后方可开展经营活动)。
由于经营不善,景峰医药目前生产经营不佳,无法清偿到期
负债,且已不具备债务清偿能力。
常德中院根据债权人申请,于2024年7月2日对景峰
医药启动预重整并指定北京市中伦律师事务所作为公司预
重整期间的临时管理人,组织开展预重整各项工作。预重整
期间,经过公开招募和遴选程序,临时管理人、景峰医药分
别与石药控股、德源招商签署了《重整投资协议》。2025年
10月21日,常德中院作出(2024)湘07破申7号《民事
裁定书》,裁定受理彭东钜、上海鑫绰投资管理有限公司
对景峰医药的重整申请,并同日作出(2025)湘07破15
号《决定书》,指定北京市中伦律师事务所担任湖南景峰
医药股份有限公司重整管理人,具体开展重整各项工作。
景峰医药的重整工作得到了政府及常德中院的高度重
视和大力支持。为保证重整成功,避免景峰医药破产清算,
管理人在常德中院的监督和指导下,严格遵照《企业破产法》
的规定全面履行相关职责。预重整期间,临时管理人启动债
权、资产的调查工作,进行了第一轮债权审核,完成了重整
投资人招募工作,为进入法院正式重整做好准备工作,节省
正式重整时间、提高成功率。重整期间,管理人监督公司自
行管理财产和营业事务,确保公司生产经营及职工稳定,保
障重整工作的顺利推进;同时,景峰医药及管理人全力以赴
做好与重整相关的各项具体工作,包括债权申报受理与审查、
财产调查、职工债权调查、重整投资人协商谈判、重整计划
(草案)的论证和制作等。在景峰医药预重整及重整各项工
作推进过程中,常德中院均依法监督,确保重整程序依法合
规开展,切实保障各方主体的合法权益。
截至目前,管理人已经完成重整所需各项基础工作,对
景峰医药的整体现状已有全面了解。在充分听取债权人、重
整投资人、股东等各方意见和建议的基础上,在充分尊重评
估机构出具的《资产市场价值报告》《资产清算价值报告》
及《偿债能力分析报告》结论的前提下,在充分进行法律上
的风险评估和论证、可行性预判和分析的条件下,景峰医药
严格依据《企业破产法》第七十九条、第八十条、第八十一
条之规定,在法律、法规及司法解释允许的范围内,结合景
峰医药实际情况,制作重整计划(草案),供债权人会议审
议、表决,并由出资人组会议对重整计划(草案)中涉及的
出资人权益调整事项进行表决。
摘要
一、重整完成后,景峰医药的企业法人性质及市场主体
资格不变,并将最大可能消除债务危机及其他风险,提升上
市公司质量。
二、本重整计划将以景峰医药现有股本879,774,351股
为基数,按照每10股转增10股的比例实施资本公积金转增
股本,共计可转增879,774,351股股票(最终实际转增的股
票数量以中证登深圳分公司实际登记确认的数量为准)。转
增后,景峰医药总股本将增至1,759,548,702股(最终转增的
准确股票数量以中证登深圳分公司实际登记确认的数量为
准)。前述转增的879,774,351股股票不再向原股东分配,全
部用于引入重整投资人,并由重整投资人提供资金认购,相
应资金用于根据重整计划的规定支付破产费用、清偿各类债
务、补充公司流动资金等。
三、职工债权、税款债权不作调整,在法院裁定批准重
整计划且债权经法院裁定确认后20个工作日内以现金方式
一次性清偿完毕。
四、普通债权的清偿方案如下:
普通债权以债权人为单位,每家普通债权人的普通债权
在200万元以下(含200万元)的部分,由景峰医药在后续
重整计划执行期限内以现金方式一次性清偿完毕。
每家普通债权人的普通债权超过200万元的部分,可选
择如下方案进行受偿:
(1)留债方案
针对每家普通债权人的普通债权超过200万元的部分,
以现金方式予以留债清偿,留债最长期限为2年,留债期限
内不计息。留债期限内,不影响债权人根据有关合同约定以
及法律规定,就其债权未获清偿部分(含合同约定或法定的
利息)向其他债务人(包括但不限于主债务人)主张有关权
益,亦不影响其他担保人或主债务人根据有关合同约定以及
法律规定向债权人清偿债务。
(2)现金清偿方案
每家普通债权人按90%的清偿率在法院裁定批准重整计
划后90日内获得现金清偿;剩余未获清偿债权部分在重整计
划执行完毕后豁免清偿,景峰医药不再承担清偿责任。
债权金额超过200万元的普通债权人(含暂缓确认债权
人),应自法院裁定批准重整计划之日起10个工作日内按照
重整计划要求的书面格式提供债权清偿方式选择告知书。债
权人逾期未告知的,视为全额选择按以普通债权清偿方式中
的“现金清偿方案”获得清偿。
本重整计划执行完毕后,全体债权人的债权将得到较高
清偿,公司的财务状况将得到根本改善,在减轻债务负担的
同时,公司可以有效提升经营效率和盈利能力,全体投资者
的利益将有望得到有效保护。
上述为本重整计划核心内容的摘录或总结,具体内容文
义以正文表述为准。
正文
一、景峰医药基本情况
(一)基本信息
景峰医药注册地址为湖南省常德经济技术开发区樟木
桥街道双岗社区桃林路661号(双创大厦1703室),营业期
限为1998年12月18日至无固定期限,统一社会信用代码
为914306007121062680,法定代表人为刘树林,注册资本
为87,977.4351万元人民币,总股本879,774,351股。景峰医
药是一家在深圳证券交易所A股公开发行股票的上市公司,
证券代码为000908。
景峰医药经营范围为:以自有资产进行医药、医疗项目
投资;生物制药技术项目的研发与投资;商品进出口贸易;
企业管理咨询、医疗医药研发技术咨询(依法须经批准的项
目,经相关部门批准后方可开展经营活动)。
(二)股本结构
1.前十大股东
截至2025年9月30日,景峰医药总股本为879,774,351
股,期末股东总数为27,854户,前10大股东持股情况如下:
股东名称/姓名持股比例中国长城资产管理股份有限公司12.92%叶湘武12.76%平江县国有资产事务中心1.26%武阳0.84%赵建锋0.65%股东名称/姓名持股比例陈永华0.60%陆琦0.57%侯健0.56%孙江波0.52%杜永生0.45%2.实际控制人及其持股情况
截至目前,叶湘武及其一致行动人合计持有公司股份数
量为114,684,310股,占公司股份总数的13.04%,景峰医药
的实际控制人为叶湘武。
(三)预重整及重整的申请及受理情况
2024年6月26日,申请人彭东钜、上海鑫绰投资管理
有限公司向常德中院申请对景峰医药进行重整,并同时申请
对景峰医药启动预重整。
2024年7月2日,常德中院作出(2024)湘07破申7
号《决定书》,常德中院决定对景峰医药启动预重整。
2024年7月30日,常德中院作出(2024)湘07破申7
号之一《决定书》,指定北京市中伦律师事务所担任景峰医
药预重整期间临时管理人。
2025年10月21日,常德中院作出(2024)湘07破申
7号《民事裁定书》,裁定受理债权人对景峰医药的重整申请,
并于同日作出(2025)湘07破15号《决定书》,指定北京
市中伦律师事务所为景峰医药重整管理人。
(四)资产情况
1.账面资产情况
截至2025年9月30日,景峰医药(母公司单体,下同)
的资产总额为11.41亿元。景峰医药主要资产由长期股权投
资、其他应收款等构成。景峰医药截至2025年9月30日的
账面资产明细情况具体如下:
类别占比流动资产合计9.82%其中:其他应收款9.34%其他流动资产0.42%货币资金0.05%非流动资产合计90.18%其中:长期股权投
资87.78%无形资产2.40%固定资产0.01%资产合计100.00%2.资产评估情况
根据评估机构出具的《资产市场价值报告》,截至2025
年9月30日,基于重整状态下的持续经营假设为前提,景峰
医药总资产的评估价值为59,180.50万元,具体如下:
类别流动资产合计非流动资产合计其中:长期股权投资固定资产无形资产类别资产合计(五)负债情况
结合审计机构出具的《审计报告》等,并经管理人对景
峰医药债权申报的审查,对于已申报债权,管理人已进行审
查认定,对于未申报债权,管理人已作预留相应偿债资源。
景峰医药负债情况具体如下:
1.债权申报情况
截至目前,共有60家债权人向管理人申报了债权(不含
职工债权),申报债权总额为233,295.05万元,其中,2家债
权人申报债权性质为税款及社保债权,对应申报金额为
137.94万元,4家债权人申报债权性质为有财产担保的债权
(含一家债权人申报债权性质为建设工程优先权债权),对
应申报金额为92,467.05万元,54家债权人申报债权性质为
普通债权,对应申报金额为140,690.06万元。
2.债权审查情况
(1)审查确定的债权
债权人已申报债权中,截至重整计划(草案)披露日,
经管理人审查确定的债权总额为97,750.52万元,涉及债权人
43家。其中包括1家税款债权,债权金额为131.77万元;
42家普通债权,债权金额为97,618.75万元。
(2)暂缓确认债权
截至重整计划(草案)披露日,不存在暂缓确认债权。
(3)不予确认的债权
债权人已申报债权中,截至重整计划(草案)披露日,
不予确认的债权对应41家债权人申报的债权(包含部分不
予确认的情形),涉及债权申报金额为135,521.13万元。
需要说明的是,对于在景峰医药预重整阶段申报的附利
息债权,管理人已将相应利息追加计算至重整受理日前一日。
债权最终审查结论以经常德中院裁定确认的债权表中记载
的为准。
3.职工债权调查情况
经管理人调查,截至2025年10月20日,景峰医药的
职工债权总额为272.81万元。
4.税款债权情况
经管理人调查,截至2025年10月20日,景峰医药的
税款债权为131.77万元,涉及1家债权人。
5.未申报债权情况
截至2025年10月20日,根据审计机构出具的《审计
报告》、公司的说明及管理人调查梳理,未在重整计划(草
案)提交债权人会议表决前申报但可能受法律保护的债权总
额约8,088万元。
(六)偿债能力分析
为给债权人表决重整计划(草案)提供必要参考,管理
人委托评估机构对景峰医药在假定破产清算条件下的偿债
能力进行了分析。根据评估机构出具的《偿债能力分析报告》,
截至评估基准日,在模拟破产清算状态下,假定景峰医药全
部资产能够按清算价值实际变现,按照《企业破产法》规定
的清偿顺序,破产财产的变现所得在支付必要的破产费用、
共益债务、职工债权、有财产担保的债权后,普通债权清偿
率约为21.61%。
考虑到景峰医药如实施破产清算,能够达到《偿债能力
分析报告》中普通债权清偿率的前提,一是资产处置时能够
按照评估价值变现,二是处置税费等破产费用能够控制在管
理人和评估机构预测的范围内。同时,司法实践中破产清算
程序耗时可能较长,将进一步产生超过预期的各项费用。基
于以上因素,景峰医药实际在破产清算状态下可用于偿债的
资金将可能比《偿债能力分析报告》预计的更低,导致债权
人的利益进一步受损。
而在本次重整中,对于普通债权的200万元以下(含200
万元)部分,将全额现金清偿,因此该部分普通债权,清偿
率为100%。对于普通债权超过200万元的部分,在“现金清
偿方案”中债权人超过200万元的部分为现金清偿,清偿率
为90%;在“留债方案”中债权人超过200万元的债权,留
债最长期限为2年,留债清偿后预计清偿率为100%。
二、出资人权益调整方案
(一)出资人权益调整的必要性、合理性
根据最高人民法院、中国证监会《关于切实审理好上市
公司破产重整案件工作座谈会纪要》“五、关于上市公司破
产重整计划草案的制定及表决”之“16.出资人权益调整”中
规定,“上市公司资产不足以清偿全部债务且普通债权人不
能在重整计划中全额获得清偿的,原则上应对出资人权益进
行调整。”
鉴于景峰医药已不能清偿到期债务,且明显缺乏清偿能
力,生产经营和财务状况均已陷入困境。同时,普通债权人
不能在重整计划中全额获得清偿。因此,本重整计划安排对
出资人权益进行调整。景峰医药在确定出资人权益调整方案
时主要考虑了如下因素:
1.本重整计划偿债方案中所需的偿债资源。
2.景峰医药经营方案中涉及未来经营恢复、业务升级、
内部管理系统改造升级等所需的资金。
3.充分考虑中小投资者利益,避免过度转增导致对中小
投资者权益的稀释。
综合上述因素,本次出资人权益调整以景峰医药现有总
股本为基础,按照每10股转增10股的比例实施资本公积转
增股本。该安排符合《上市公司监管指引第11号——上市
公司破产重整相关事项》的规定。
(二)出资人权益调整的范围
根据《企业破产法》第八十五条第二款之规定,重整计
划(草案)涉及出资人权益调整事项的,应当设出资人组,
对该事项进行表决。
景峰医药出资人组由截至出资人组会议召开公告所载
明确定的股权登记日在中证登深圳分公司登记在册的景峰
医药股东组成。上述股东在出资人组会议之股权登记日后至
本出资人权益调整方案实施完毕前,由于交易或非交易原因
导致持股情况发生变动的,本重整计划规定的出资人权益调
整方案的效力及于其股票的受让方及/或承继方。
(三)出资人权益调整的内容
本次权益调整以景峰医药现有总股本879,774,351股为
基数,按照每10股转增10股的比例实施资本公积金转增股
本,共计可转增879,774,351股股票(最终实际转增的股票
数量以中证登深圳分公司实际登记确认的数量为准)。转增
后,景峰医药总股本将增至1,759,548,702股。
上述转增股票不向原股东分配,全部由重整投资人支付
现金对价受让,具体受让股票数以后续重整计划执行阶段的
司法协助执行通知书载明的内容及中证登深圳分公司实际
登记确认的数量为准。
(四)除权与除息
重整计划(草案)经法院裁定批准后执行,根据本重整
计划规定实施的资本公积转增股本将全部用于引进重整投
资人;本次转增后,公司在总股本扩大的同时债务规模明显
减少、所有者权益明显增加;公司原每股股票所代表的企业
实际价值(以每股净资产计算)较重整前显著提升。这与转
增前后公司所有者权益不变、需要按照《深圳证券交易所交
易规则(2023修订)》第4.4.2条中载明的除权(息)参考价
计算公式对股票价格进行调整的一般情形存在本质差别。因
此,重整计划实施后,为反映出资人权益调整事项对景峰医
药股票价值的影响,需要对本次资本公积金转增股票除权计
算公式进行调整,并可能对本次资本公积转增股权登记日次
一交易日的股票开盘参考价进行调整。
根据《深圳证券交易所交易规则(2023修订)》第4.4.2
条的规定:“除权(息)参考价计算公式为:除权(息)参
考价=[(前收盘价-现金红利)+配股价格×股份变动比例]
÷(1+股份变动比例),证券发行人认为有必要调整上述计
算公式时,可以向本所提出调整申请并说明理由。经本所同
意的,证券发行人应当向市场公布该次除权(息)适用的除
权(息)参考价计算公式。”
公司已聘请财务顾问对本次重整中拟实施资本公积转
增股本除权参考价格的计算公式进行论证,最终以财务顾问
出具的专项意见为准。如果股权登记日公司股票收盘价高于
财务顾问专项意见中确认的本次重整资本公积转增股本的
平均价的,公司股票将按除权后价格于股权登记日次一交易
日调整开盘参考价。如果股权登记日公司股票收盘价格低于
或等于财务顾问专项意见中确认的本次重整资本公积转增
股本的平均价的,公司股权登记日次一交易日的股票开盘参
考价无需调整。
后续若上述拟调整的除权参考价格的计算公式或相关
计算参数因裁定批准的重整计划或根据监管规则要求需要
另行调整的,公司将按照前述要求进行调整。
(五)出资人权益调整方案的执行效果
根据上述出资人权益调整方案,景峰医药出资人所持有
的公司股票绝对数量不会因本次重整而减少。重整完成后,
景峰医药的基本面将发生根本性改善,并将逐步提升持续盈
利能力,公司价值也将进一步提升,有利于保护公司、债权
人及广大出资人的合法权益。
三、债权分类及调整方案
根据《企业破产法》规定,景峰医药债权将按照职工债
权、税款债权、有财产担保债权、普通债权以及劣后债权进
行分类,具体分类及调整原则如下:
(一)债权分类方案
1.职工债权
经管理人调查,截至2025年10月20日,景峰医药的
职工债权总额为272.81万元。
2.税款债权
经管理人审查,截至2025年10月20日,景峰医药的
税款债权为131.77万元,涉及1家债权人。
3.有财产担保债权
经管理人审查,截至2025年10月20日,景峰医药无
有财产担保债权人。
4.普通债权
经管理人审查确定,截至2025年10月20日,普通债
权总额为97,618.75万元,共计42家债权人。
5.劣后债权
截至2025年10月20日,经管理人审查,景峰医药无
劣后债权。
(二)债权调整方案
1.职工债权
职工债权不作调整,在法院裁定批准重整计划且债权经
法院裁定确认后20个工作日内以现金方式一次性清偿完毕。
2.税款债权
根据《企业破产法》第八十二条第一款第(三)项规定
的景峰医药欠付的税款本金,在本次重整中不作调整,在法
院裁定批准重整计划且债权经法院裁定确认后20个工作日
内以现金方式一次性清偿完毕。
3.有财产担保的债权
有财产担保债权人通过设定财产担保或依据相关法律
规定而对景峰医药特定财产享有优先受偿的权利,以担保财
产持续经营状态下的评估价值确定优先受偿范围。若担保财
产的评估价值不足以清偿所对应的有财产担保债权,则有财
产担保债权超出担保财产评估价值的部分按照本重整计划
规定的普通债权清偿方案受偿;若担保财产的评估价值超出
所对应的有财产担保债权,则超出部分不属于该有财产担保
债权人享有优先受偿权的范围。
经管理人审查,截至2025年10月20日,景峰医药无
有财产担保债权人。
4.普通债权
为最大限度地保护债权人合法权益、提高债权人的受偿
水平,根据景峰医药实际情况,将按照常德中院裁定批准的
重整计划规定的清偿方案予以清偿。
5.劣后债权
景峰医药在进入重整前产生的民事惩罚性赔偿金、行政
罚款、刑事罚金、加倍支付迟延履行利息等惩罚性债权为劣
后债权,不予清偿。
四、债权清偿方案
(一)偿债资源
本次重整的偿债资源为重整投资者支付的重整投资款
以及公司自有货币资金、未来持续经营收入等。
(二)债权清偿方案
1.职工债权
职工债权由景峰医药在法院裁定批准重整计划且债权
经法院裁定确认后20个工作日内以现金方式一次性清偿完
毕。
2.税款债权
税款债权由景峰医药在法院裁定批准重整计划且债权
经法院裁定确认后20个工作日内以现金方式一次性清偿完
毕。
3.普通债权
普通债权以债权人为单位,每家普通债权人的普通债权
在200万元以下(含200万元)的部分,由景峰医药在后续
重整计划执行期限内以现金方式一次性清偿完毕。
每家普通债权人的普通债权超过200万元的部分,可选
择如下方案进行受偿:
(1)留债方案
针对每家普通债权人的普通债权超过200万元的部分,
以现金方式予以留债清偿,留债最长期限为2年,留债期限
内不计息。留债期限内,不影响债权人根据有关合同约定以
及法律规定,就其债权未获清偿部分(含合同约定或法定的
利息)向其他债务人(包括但不限于主债务人)主张有关权
益,亦不影响其他担保人或主债务人根据有关合同约定以及
法律规定向债权人清偿债务。
(2)现金清偿方案
每家普通债权人按90%的清偿率在法院裁定批准重整计
划之日起90日内获得现金清偿;剩余未获清偿债权部分在重
整计划执行完毕后豁免清偿,景峰医药不再承担清偿责任。
债权金额超过200万元的普通债权人(含暂缓确认债权
人),应自法院裁定批准重整计划之日起10个工作日内按照
重整计划要求的书面格式提供债权清偿方式选择告知书。债
权人逾期未告知的,视为全额选择按以普通债权清偿方式中
的“现金清偿方案”获得清偿。
(三)劣后债权清偿方案
景峰医药涉及的民事惩罚性赔偿金、行政罚款、刑事罚
金等惩罚性债权将劣后于其他普通债权。针对劣后债权的处
理,由于普通债权未获得全额清偿,因此依法不安排偿债资
源。
(四)合并报表范围内关联债权的清偿方案
景峰医药合并范围内关联债权人对景峰医药享有的债
权,按照本重整计划中同类债权的清偿方案予以清偿。
(五)预计债权清偿方案
1.暂缓确认债权
已向管理人申报,但因涉及未决诉讼等原因而尚未经管
理人审查确定的债权,其中普通债权人应当提前就剩余债权
(剔除200万元现金清偿部分)选择何种方案清偿以书面形
式加以明确,在债权经审查确定后10个工作日内按照本重
整计划要求的书面格式提供债权清偿方式选择告知书。债权
人逾期未告知的,视为全额选择按以普通债权清偿方式中的
“现金清偿方案”获得清偿。
除因诉讼仲裁未决等原因而暂未确认的债权外,在重整
计划(草案)提交债权人会议表决前,已依法申报但因各种
原因在后续重整程序中暂未确认的债权,如在后续重整计划
执行期内仍未获得确认,则由景峰医药在后续重整程序终结
后,在不损害其他债权人权益的情况下,通过包括诉讼、仲
裁、协商等适当方式审查确定债权金额。因诉讼仲裁未决而
暂未确认的债权,依法院或仲裁机构的生效法律文书确定债
权金额和性质。上述暂未确认债权获得确认后10个工作日
内按照重整计划要求的书面格式提供债权清偿方式选择告
知书。债权人逾期未告知的,视为全额选择按以普通债权清
偿方式中的“现金清偿方案”获得清偿。
2.未申报债权
对于景峰医药账面有记载但未申报的债权,本重整计划
已预留相应偿债资源。未申报债权在本重整计划执行期间不
得行使权利;在本重整计划执行完毕后,该部分未申报债权
经申报并经债务人审查确定后,可要求景峰医药按照本重整
计划中规定的同类债权受偿方式予以清偿。根据《企业破产
法》第五十六条规定,为审查和确认补充申报债权的费用,
由补充申报人承担。未依法申报债权的债权人,自重整受理
日起三年内或至该等债权的诉讼时效届满之日(以孰早为
准),未向景峰医药主张权利的,景峰医药不再负有清偿义
务。如因客观因素需调整未申报债权受偿方式的,可由公司
与前述债权人另行协商。
五、重整投资人情况
(一)重整投资人的招募情况
2024年8月1日,公司发布《湖南景峰医药股份有限公
司临时管理人关于意向投资者招募的公告》,临时管理人通
过全国企业破产重整案件信息网同步发布公开招募重整投
资人的公告(以下简称“招募公告”),正式启动投资人公开
招募工作。
截至2024年8月15日,共有4家投资人(联合体按1
家计算)提交报名材料。截至2024年8月25日,共有1家
投资人(联合体按1家计算)缴纳尽调保证金。根据《关于
招募重整投资人延期的公告》,若仅有一名意向投资人或投
资人联合体提交重整投资方案,临时管理人将组织通过商业
谈判方式确定重整投资人。2024年8月27日,景峰医药发
布《关于确定预重整投资人暨风险提示的公告》,最终确定
以石药控股作为牵头投资人的联合体为中选重整投资人。
2025年4月29日,景峰医药发布《关于与重整投资人
签署重整投资协议的公告》,披露了景峰医药、临时管理人
与石药控股及作为联合体成员之一的德源招商签署的重整
投资协议,其中包括投资人认购股份数量及价格条款。
2026年1月9日,管理人、景峰医药与石药控股及德源
招商分别签署《重整投资协议之补充协议》,对投资价格及
投资款支付期限予以补充约定。
2026年1月9日,管理人、景峰医药与中国长城资产管
理股份有限公司签署《重整投资协议》。
经牵头投资人石药控股、管理人等各方与意向投资人沟
通磋商,2026年1月9日,管理人、景峰医药与16家财务
投资人签署《重整投资协议》。
债务人已就重整投资人招募及重整投资协议签署情况
履行了信息披露义务。
(二)重整投资人具体情况
经过公开招募和遴选,确定以石药控股作为牵头投资人
的联合体为景峰医药的重整投资人。根据重整投资人的说明,
重整投资人基本情况介绍如下:
1.石药控股集团有限公司
石药控股集团有限公司成立于1998年3月31日,统一
社会信用代码为911301002360444643。石药控股的实际控
制人为蔡东晨。
景峰医药现任董事及总裁刘树林,曾任石药控股集团有
限公司副总裁兼第一制造中心总裁、总计划调度中心总经理
等职务。景峰医药现任董事马学红曾任石药控股集团有限公
司财务部税务主管、税务经理;河北中润制药有限公司财务
部经理、石药集团欧意药业有限公司财务总监、石药控股集
团有限公司资产管理中心高级总监。景峰医药现任董事廉奇
志曾任石药集团有限公司高级总监、资本运营中心总经理等
职务,现任石药控股集团河北省教育科技有限公司董事。除
此之外,石药控股与景峰医药及其董事、高级管理人员、控
股股东、实际控制人等不存在关联关系或者一致行动关系。
重整完成后,石药控股将成为景峰医药的控股股东。石
药控股承诺自受让转增股票之日起36个月内不转让或者委
托他人管理其基于本次重整投资所直接和间接持有的景峰
医药股票。
2.常德瑞健一禾咨询管理合伙企业(有限合伙)
常德瑞健一禾咨询管理合伙企业(有限合伙)成立于
2025年11月,统一社会信用代码为91430700MAK0Y24L1W。
其实际控制人为徐铭泽。常德瑞健一禾咨询管理合伙企业
(有限合伙)与景峰医药及其董事、高级管理人员、控股股
东、实际控制人等不存在关联关系或者一致行动关系。常德
瑞健一禾咨询管理合伙企业(有限合伙)承诺自受让转增股
票之日起12个月内不转让或者委托他人管理其基于本次重
整投资所直接和间接持有的景峰医药股票。
3.常德智恒咨询管理合伙企业(有限合伙)
常德智恒咨询管理合伙企业(有限合伙)成立于2025
年11月,统一社会信用代码为91430700MAK0XCJA9B。其实
际控制人为贾路琦。常德智恒咨询管理合伙企业(有限合伙)
与景峰医药及其董事、高级管理人员、控股股东、实际控制
人等不存在关联关系或者一致行动关系。常德智恒咨询管理
合伙企业(有限合伙)承诺自受让转增股票之日起12个月
内不转让或者委托他人管理其基于本次重整投资所直接和
间接持有的景峰医药股票。
4.常德市德源招商投资有限公司
常德市德源招商投资有限公司成立于2012年11月9
日,统一社会信用代码为91430700055844286Y。德源招商为
常德市德源投资集团有限公司的全资子公司,实际控制人为
常德市人民政府国有资产监督管理委员会。德源招商投资事
务部部长张莉现任景峰医药代行董事长。除此之外,德源招
商与景峰医药及其董事、高级管理人员、控股股东、实际控
制人等不存在关联关系或者一致行动关系。德源招商承诺自
受让转增股票之日起36个月内不转让或者委托他人管理其
基于本次重整投资所直接和间接持有的景峰医药股票。
5.常德金洋洪赢商务服务合伙企业(有限合伙)
常德金洋洪赢商务服务合伙企业(有限合伙)成立于
2024年12月,统一社会信用代码为91430700MAE748723N。
其实际控制人为江西省人民政府。常德金洋洪赢商务服务合
伙企业(有限合伙)与景峰医药及其董事、高级管理人员、
控股股东、实际控制人等不存在关联关系或者一致行动关系。
常德金洋洪赢商务服务合伙企业(有限合伙)承诺自受让转
增股票之日起12个月内不转让或者委托他人管理其基于本
次重整投资所直接和间接持有的景峰医药股票。
6.宁波量利私募基金管理有限公司(代表宁波量利私募
基金管理有限公司-量利元启1号私募证券投资基金、宁波
量利私募基金管理有限公司-量利元启4号私募证券投资基
金、宁波量利私募基金管理有限公司-量利元玺3号私募证
券投资基金)
宁波量利私募基金管理有限公司成立于2015年4月,
统一社会信用代码为9133021234060269X8。其实际控制人为
何振权。宁波量利私募基金管理有限公司为宁波量利私募基
金管理有限公司-量利元启1号私募证券投资基金、宁波量
利私募基金管理有限公司-量利元启4号私募证券投资基金、
宁波量利私募基金管理有限公司-量利元玺3号私募证券投
资基金的管理人。宁波量利私募基金管理有限公司与景峰医
药及其董事、高级管理人员、控股股东、实际控制人等不存
在关联关系或者一致行动关系。宁波量利私募基金管理有限
公司(代表宁波量利私募基金管理有限公司-量利元启1号
私募证券投资基金、宁波量利私募基金管理有限公司-量利
元启4号私募证券投资基金、宁波量利私募基金管理有限公
司-量利元玺3号私募证券投资基金)承诺自受让转增股票
之日起12个月内不转让或者委托他人管理其基于本次重整
投资所直接和间接持有的景峰医药股票。
7.中国长城资产管理股份有限公司
中国长城资产管理股份有限公司成立于1999年11月,
统一社会信用代码为91110000710925489M。其实际控制人
为国务院国资委。中国长城资产管理股份有限公司现为景峰
医药的股东,持股数量为113,680,665股,持股比例为12.92%。
景峰医药现任董事谢树青,现任中国长城资产管理股份有限
公司湖南省分公司资产经营二部一级业务主管;景峰医药原
监事会主席纪纲(离任未满12个月),现任中国长城资产管
理股份有限公司资产经营六部高级经理。除此之外,中国长
城资产管理股份有限公司与景峰医药及其董事、高级管理人
员、控股股东、实际控制人等不存在关联关系或者一致行动
关系。中国长城资产管理股份有限公司承诺自受让转增股票
之日起36个月内不转让或者委托他人管理其基于本次重整
投资所直接和间接持有的景峰医药股票。
8.湖南财鑫资本管理有限公司
湖南财鑫资本管理有限公司成立于2014年1月,统一
社会信用代码为914307000919795759。其实际控制人为常德
市财政局。湖南财鑫资本管理有限公司与景峰医药及其董事、
高级管理人员、控股股东、实际控制人等不存在关联关系或
者一致行动关系。湖南财鑫资本管理有限公司承诺自受让转
增股票之日起12个月内不转让或者委托他人管理其基于本
次重整投资所直接和间接持有的景峰医药股票。
9.安徽明泽投资管理有限公司(代表安徽明泽投资管理
有限公司-明泽远见6号债券私募证券投资基金)
安徽明泽投资管理有限公司成立于2014年6月,统一
社会信用代码为91440300306094812W。其实际控制人为马
科伟。安徽明泽投资管理有限公司为安徽明泽投资管理有限
公司-明泽远见6号债券私募证券投资基金的管理人。安徽
明泽投资管理有限公司与景峰医药及其董事、高级管理人员、
控股股东、实际控制人等不存在关联关系或者一致行动关系。
安徽明泽投资管理有限公司(代表安徽明泽投资管理有限公
司-明泽远见6号债券私募证券投资基金)承诺自受让转增
股票之日起12个月内不转让或者委托他人管理其基于本次
重整投资所直接和间接持有的景峰医药股票。
10.常德顺行至通咨询管理合伙企业(有限合伙)
常德顺行至通咨询管理合伙企业(有限合伙)成立于
2025年11月,统一社会信用代码为91430700MAK0MEQ233。
其实际控制人为王立平。常德顺行至通咨询管理合伙企业
(有限合伙)与景峰医药及其董事、高级管理人员、控股股
东、实际控制人等不存在关联关系或者一致行动关系。常德
顺行至通咨询管理合伙企业(有限合伙)承诺自受让转增股
票之日起12个月内不转让或者委托他人管理其基于本次重
整投资所直接和间接持有的景峰医药股票。
11.湖南景一企业管理咨询合伙企业(有限合伙)
湖南景一企业管理咨询合伙企业(有限合伙)成立于2025
年11月,统一社会信用代码为91430700MAK0P7578Q。其实
际控制人为长沙市人民政府国有资产监督管理委员会。湖南
景一企业管理咨询合伙企业(有限合伙)与景峰医药及其董事、
高级管理人员、控股股东、实际控制人等不存在关联关系或
者一致行动关系。湖南景一企业管理咨询合伙企业(有限合伙)
承诺自受让转增股票之日起12个月内不转让或者委托他人
管理其基于本次重整投资所直接和间接持有的景峰医药股
票。
12.青岛祺顺投资管理有限公司(代表青岛祺顺投资管
理有限公司-祺顺扬帆19号私募证券投资基金、青岛祺顺投
资管理有限公司-祺顺扬帆5号私募证券投资基金)
青岛祺顺投资管理有限公司成立于2019年11月,统一
社会信用代码为91370285MA3QYD4TX8。其实际控制人为宋耀。
青岛祺顺投资管理有限公司为青岛祺顺投资管理有限公司-
祺顺扬帆19号私募证券投资基金、青岛祺顺投资管理有限
公司-祺顺扬帆5号私募证券投资基金的管理人。青岛祺顺
投资管理有限公司与景峰医药及其董事、高级管理人员、控
股股东、实际控制人等不存在关联关系或者一致行动关系。
青岛祺顺投资管理有限公司(代表青岛祺顺投资管理有限公
司-祺顺扬帆19号私募证券投资基金、青岛祺顺投资管理有
限公司-祺顺扬帆5号私募证券投资基金)承诺自受让转增
股票之日起12个月内不转让或者委托他人管理其基于本次
重整投资所直接和间接持有的景峰医药股票。
13.杭州源铨投资管理有限公司(代表杭州源铨投资管
理有限公司-源铨东湖6号私募债券投资基金、杭州源铨投
资管理有限公司-源铨东湖9号私募证券投资基金)
杭州源铨投资管理有限公司成立于2017年5月,统一
社会信用代码为91330106MA28T09Q6U。其实际控制人为石
良希。杭州源铨投资管理有限公司为杭州源铨投资管理有限
公司-源铨东湖6号私募债券投资基金、杭州源铨投资管理
有限公司-源铨东湖9号私募证券投资基金的管理人。杭州
源铨投资管理有限公司与景峰医药及其董事、高级管理人员、
控股股东、实际控制人等不存在关联关系或者一致行动关系。
杭州源铨投资管理有限公司(代表杭州源铨投资管理有限公
司-源铨东湖6号私募债券投资基金、杭州源铨投资管理有
限公司-源铨东湖9号私募证券投资基金)承诺自受让转增
股票之日起12个月内不转让或者委托他人管理其基于本次
重整投资所直接和间接持有的景峰医药股票。
14.常德德润产业发展有限公司
常德德润产业发展有限公司成立于2022年5月,统一社
会信用代码为91430700MABNRUDE48。其实际控制人为常德
经济技术开发区财政局。常德德润产业发展有限公司与景峰
医药及其董事、高级管理人员、控股股东、实际控制人等不
存在关联关系或者一致行动关系。常德德润产业发展有限公
司承诺自受让转增股票之日起12个月内不转让或者委托他
人管理其基于本次重整投资所直接和间接持有的景峰医药
股票。
15.上海鑫绰投资管理有限公司(代表上海鑫绰投资管
理有限公司-鑫绰鑫融3号私募证券投资基金)
上海鑫绰投资管理有限公司成立于2014年1月,统一
社会信用代码为913101150900697645。其实际控制人为周
俊彦。上海鑫绰投资管理有限公司为上海鑫绰投资管理有限
公司-鑫绰鑫融3号私募证券投资基金的管理人。上海鑫绰
投资管理有限公司与景峰医药及其董事、高级管理人员、控
股股东、实际控制人等不存在关联关系或者一致行动关系。
上海鑫绰投资管理有限公司(代表上海鑫绰投资管理有限公
司-鑫绰鑫融3号私募证券投资基金)承诺自受让转增股票
之日起12个月内不转让或者委托他人管理其基于本次重整
投资所直接和间接持有的景峰医药股票。
(三)重整投资人受让转增股份情况
依据管理人、景峰医药分别与重整投资人签订的协议,
重整投资人合计受让879,774,351股转增股票,总投资额约
为20.61亿元。重整投资人支付的投资款按照本重整计划之
规定,用于包含但不限于支付破产费用、共益债务、破产债
权以及相关支出。
重整投资人受让股份情况如下:
投资人名称认购价格
(元/股)石药控股集团有
限公司1.45常德瑞健一禾咨
询管理合伙企业
(有限合伙)3.57常德智恒咨询管
理合伙企业(有
限合伙)3.57常德市德源招商
投资有限公司1.88常德金洋洪赢商
务服务合伙企业
(有限合伙)3.57宁波量利私募基
金管理有限公司
-量利元启1号
私募证券投资基
金3.57中国长城资产管
理股份有限公司3.57湖南财鑫资本管
理有限公司3.57投资人名称认购价格
(元/股)安徽明泽投资管
理有限公司-明
泽远见6号债券
私募证券投资基
金3.57常德顺行至通咨
询管理合伙企业
(有限合伙)3.57湖南景一企业管
理咨询合伙企业
(有限合伙)3.57宁波量利私募基
金管理有限公司
-量利元启4号私
募证券投资基金3.57青岛祺顺投资管
理有限公司-祺
顺扬帆5号私募
证券投资基金3.57杭州源铨投资管
理有限公司-源
铨东湖6号私募
债券投资基金3.57杭州源铨投资管
理有限公司-源
铨东湖9号私募
证券投资基金3.57常德德润产业发
展有限公司3.57宁波量利私募基
金管理有限公司3.57投资人名称认购价格
(元/股)-量利元玺3号私
募证券投资基金 青岛祺顺投资管
理有限公司-祺
顺扬帆19号私募
证券投资基金3.57上海鑫绰投资管
理有限公司-鑫
绰鑫融3号私募
证券投资基金3.57合计-六、经营方案
基于“优化存量,培育增量”的未来发展思路,景峰医
药未来业务板块将包括中成药为主、化药为辅的存量业务,
以及生物医药为主的增量业务,积极拓展合成生物产业。具
体发展举措如下:
(一)优化存量,培育增量,创新引领、双轮驱动
1.聚焦存量优势产品,提升营销能力,创新创研新产品
重整完成后,景峰医药将启动生产设备和生产过程的革
新、研发方向与产业趋势的融合,提升管理水平,重塑品牌
优势,打造具有国际先进水平的重点产业化项目。具体举措
如下:
(1)优化产品管线,聚焦存量核心产品
公司产品种类丰富,未来将逐步淘汰长期缺乏竞争力的
品种,按照潜力和目前的产品结构,公司将聚焦心血管领域
产品,并持续不断优化产品结构。具体如下:
一是公司将成立心血管销售部,聚焦心血管产品在等级
医院的终端拓展、零售终端拓展及学术推广工作。心脑宁胶
囊、参芎葡萄糖注射液、乐脉丸的等级医院渠道划归心血管
销售部管理和销售;心脑宁和参芎除了稳定现有代理商的终
端和纯销以外,要重点提升全国top100医院的学术推广覆
盖和纯销提升速度,另一方面要积极拓展空白市场和代理商
广度,设立终端开发目标进行考核,在精细化招商的指引下
逐步提升终端数量和纯销数量,为未来销售夯实基础。公司
将推动参芎葡萄糖注射液纳入国家医保目录作为景峰医药
重点工作任务。
二是对于重点产品心脑宁胶囊(全国独家),目前为医
保乙类药品,未来拟研发增加其适应症(针对精准干预急性
冠状动脉综合征后状态合并颈动脉斑块残余风险研究),加
速完成药品再评价、逐步提升药品价值并申请进入基药目录。
三是重启参芎葡萄糖注射液三期临床相关研究,扩大产
品适应症重新申报国家医保药品目录,同时加速金鸡丸、乐
脉丸等中药产品的多元发展;对于通过一致性评价的品种,
深度挖掘销售价值,增加产品的中标概率。
四是完善升级生产线,拓展玻璃酸钠制剂产品(包括玻
璃酸钠美容填充项目和水光针项目)产能。淘汰生产端陈旧
设备,改善生产自动化水平,通过委托加工等多种方式,高
效解决工厂产能不足现状,保证旺季销售供应。
五是围绕榄香烯开展系统性研究,开展质量标准提升研
究,并推进提取后副产物的资源化利用;在此基础上,开发
榄香烯系列衍生物及相关制剂,拓展其在抗肿瘤及其他疾病
治疗领域的应用范围;同时,通过优化提取工艺,降低生产
成本,提高资源利用效率,最终构建从原料到制剂的全产业
链价值提升路径。
(2)拓展销售渠道,提升营销能力
一是调整营销模式。将由各子公司独立经营销售转变为
母公司“统一规划,统一销售,统一管理,统一团队”的四
统一销售模式,促进存量产品发展。按照产品治疗领域和销
售渠道的属性进行划分形成条线管理,通过组织架构的调整,
使销售队伍在执行力、专业度上快速提升。打造心脑血管、
抗肿瘤等领域专业化的营销团队。
二是继续拓展基层及民营医疗机构等三终端,提升产品
销售和市场覆盖比例。将参芎葡萄糖注射液、乐脉丸、金鸡
丸、通迪胶囊、妇平胶囊、儿童回春颗粒、注射用克林霉素
磷酸酯、阿糖腺苷、奥美拉唑等列为发力品种,参芎重点拓
展代理商数量,填补空白市场,加强代理商管理,要求更多
市场覆盖和纯销提升;梳理、整合其他产品的三终端渠道和
代理商现状,积极开发新的市场机会,做好代理商选择及优
化,尽快完成三终端销售的布局和价值体现;普药部分做好
现有市场覆盖的维护及提升,利用积极有效的市场调研和价
格杠杆,努力培育5-10个战略合作伙伴,加快提升普药的
市场份额和销售金额。
三是规划集采药品销售方案。成立集采销售部,规划仿
制药布局,积极探索和参加国家集采及省际联盟集采,力争
更多产品进入国家集采通道和中选更多省份的集采招标。其
中,重点推进海南锦瑞公司仿制药一致性评价的过审,以及
玻璃酸钠注射液重点开发上量。另外,市场准入团队积极学
习和解读集采政策,结合公司产品结构,响应国家集采号召,
积极参与、力争中选,统一配送、跟进销售。后续公司将更
加关注多家中选省份的销售策略制定和备选省份的销售拓
展计划。
四是建立和壮大零售队伍,打造公司销售业务新增长点。
零售连锁市场逐年增长,公司将建立专门的零售队伍,通过
外部引进和内部培养的方式快速建立零售专业销售团队,产
品聚焦心脑宁胶囊、镇痛活络酊、冰栀伤痛气雾剂、乐脉丸、
金鸡丸等产品;全国top10连锁力争直接合作或者贴牌销售,
其他连锁选择专业的代理商合作,快速上架,专业推广,迅
速提升销售,为后续销售增长带来强劲动力。
(3)创新创研新产品,实现产业升级
借鉴产业投资人的研发创新优势,公司未来医药研发将
以“建立完整、高效、国内领先的研究开发系统,制定并持
续更新科学、稳健的产品线规划”为目标,建立健全与国际
接轨的研发质量管理体系,加强建设研发项目管理体系。
一是加快推进化药新药项目落地,NMPA1类新药交联玻
璃酸钠注射液根据III期临床CED沟通会的要求进行补充研
究,推动完成III期临床试验样品的生产与工艺验证,并启
动III期临床研究。
二是推动普通注射剂项目完成一致性评价及工艺验证,
推进玻璃酸钠注射液、注射用氯诺昔康开展一致性评价工作。
三是持续推进抗肿瘤产品研发,以多种方式解决抗肿瘤
药物伊立替康注射液、注射用吉西他滨、注射用异环磷酰胺
等产品的产能不足及产销问题。
四是公司在产品立项上将继续保持谨慎态度,依据自身
研发优势及核心竞争力,从多维度进行产品评估,结合未来
产品市场竞争态势的预测,以及未来销售量等市场数据,确
定研发方向,推进临床研究以及里程碑的逐步实现,以实现
产品上市后的收益。
2.培育生物医药第二增长曲线
(1)常德市政策支持生物医药产业,已形成产业集聚
生物制药产业在常德市拥有深厚的产业基础。常德在生
物制造领域底蕴深厚,拥有亚洲最大的甾体原料药和中间体
生产出口基地、全国最大的酶制剂生产出口基地、全省第一
方阵的生物医药企业、全省规模领先的食品加工产业。目前,
常德市已建成国家级合成生物制造产业创新平台1个、省级
创新平台5个、省级科技成果转化中试基地1个。常德经开
区和津市市的合成生物中试转化基地即将投入使用,安乡县
合成生物产业应用研发中心已经建成。常德市和湖南文理学
院共建合成生物学公共技术平台,一体布局生物合成、分子
检测、生物提取、生物发酵和生物医学等高能级实验室,争
创合成生物省级、国家级重点实验室。
生物制药产业在常德市拥有完备的园区配套支撑。按照
“一中心三基地”产业布局,津市市、常德经开区产业承载
优势,安乡县原料资源优势,全力打造全省引领、全国有名
的常德“生物制造谷”,推动合成生物制造产业高端化、绿
色化、融合化、协同化发展。
(2)石药控股及景峰医药在生物制药领域拥有深厚积
累
石药控股在生物制药领域拥有深厚积累,研发实力雄厚。
石药控股及其下属企业依托纳米制剂药物、细胞治疗、长效
注射剂等八大技术平台,聚焦肿瘤、精神神经、心血管等领
域,并在中药和合成生物学等领域有所布局。
景峰医药研发中心已分别打造生物药开发技术平台、大
分子交联技术平台、纳米分散系DDS平台、MUPS(MultipleUnit
PelletSystem)OSD平台、预灌封制剂产业化平台等研发平台,
致力于建立高技术壁垒的生物药品研发与生产平台,为拓展
生物制药提供研发基础保障。
(3)依托区位优势及企业积累,培育第二增长曲线
重整完成后,景峰医药将依托常德市对生物医药行业的
政策支持、资金保障、产业集聚,并凭借石药控股和公司在
生物医药行业的积累与研发实力,着力培育生物医药作为第
二增长曲线,拓展合成生物产品及关联赛道,逐步成长为双
轮驱动的优质公司。具体举措包括但不限于:
一是聚焦合成生物学与创新药械,推动具体项目落地。
公司将重点引入与现有业务具有战略协同效应的优质合成
生物学项目,在常德逐步构建从研发、中试到产业化的合成
生物制造产业链,为业绩贡献新增量。
二是联合设立专项产业基金,构建项目孵化与引入平台。
该基金将借助石药控股的产业资源和基金管理经验,主要投
向生物医药及相关联的高成长性领域,在追求财务回报的同
时,积极引导被投企业将研发成果和生产基地落户常德,形
成“基金投资—项目孵化—产业落地”的良性循环。
三是整合内外部生物医药资源,打造“研发-生产-销售”
一体化生态。推动石药控股与景峰医药的生物医药协同研发
项目在常德优先转化及生产落地,并借助石药控股整合两湖
两广销售资源的优势,在常德设立销售总部统一管理,从而
全面强化常德总部的职能,打造“研发-生产-销售”的生物
医药全链路,实现第二曲线扎实增长。
(二)优化组织架构,提升管理水平,降本增效
借鉴产业投资人丰富的企业管理经验,景峰医药将进一
步优化组织运营管理架构,强化企业管理,加强景峰医药及
子公司在战略、财务、人才、业务等方面的规范运作管理,
不断提升内控管理体系运行质量,通过建立和运行常态化的
管理提升机制,降低管理成本及费用,推进和保障公司可持
续发展。具体如下:
1.优化组织架构,强化总部管控
一是公司将继续严格执行《公司法》、《证券法》及相
关规定,严格履行股东会及董事会的监督指导作用,充分发
挥独立董事的风险把控作用。
二是重构组织架构,通过销售集中、采购集中、财务集
中三集中模式,实现扁平化管理。强化母公司管控力度,梳
理人员配置,定岗定编,先推动实现财务专业和营销专业纳
入母公司的集中体系化管理,再逐步拓展至生产、研发、采
购等专业管理;发挥母公司计划、调度、考核等管理职能,
合理科学调配母公司内部资源,实现公司整体利益最大化。
2.完善管理制度,提升管理水平
一是优化业务流程。加强制度和流程的执行管理,提高
组织运营效率;完善预算管理体系与制度,实现管理的横向
到边、纵向到底的全覆盖;完善健全投资管理制度,以理性
的、科学的投资决策和管理制度引领企业稳健增长;建立高
效管理机制,搭建周报、半月会和经营分析会等方式,确保
重点工作、核心指标的达成。
二是强化财务风险管控。包括重点加强资金审批控制、
完善内部会计稽核制度、严格执行预算和收支管理等;公司
将合理运用法规及制度筹措资金,并提高现金周转率和回款
速度、降低回款周期;将安排工作组加大对应收账款的清欠
回收力度,进一步加强应收账款管理,加速资金回笼以缓解
资金压力。
三是制定人才发展战略。公司将全面梳理人员配置,完
善薪酬考核体系,在培养人才的同时留住人才。公司将围绕
经营目标,落地预算管理,加强指标与绩效考核的管理,分
配机制重点向业务线、市场线倾斜,激发员工的工作热情,
强化以结果为导向的考核机制,实行内部竞争上岗制和轮岗
制,激发内部活力。
3.加强成本管控,盘活低效资产
一是落地成本领先战略。公司将持续抓好生产组织和原
料采购,降低运营成本并认真做好开源节流、增收节支工作。
二是盘活低效资产,提升资产运营效率,降低资产运营
压力。公司将继续对低效业务、低效资产与闲置资产进行重
整与盘活,充分提高资产利用率、提高存货周转率,不断强
化主营业务,提升经营现金流。
七、重整计划(草案)的表决和批准
(一)重整计划(草案)的表决
1.分组表决
景峰医药债权人会议设普通债权组对重整计划(草案)
进行正式表决。根据《企业破产法》第八十二条和第八十三
条、《最高人民法院关于适用
若干问题的规定(三)》第十一条,职工债权、税款债权组
因权益未受调整而不参与表决,依法不设表决组。
因重整计划(草案)涉及出资人权益调整事项,设出资
人组对出资人权益调整事项进行表决。
2.表决机制
(1)债权人组的表决机制
根据《企业破产法》第八十四条第二款的规定,出席会
议的同一表决组的债权人过半数同意重整计划(草案),并
且其所代表的债权额占该组债权总额的三分之二以上的,即
为该组通过重整计划(草案)。
(2)出资人组的表决机制
根据最高人民法院《关于切实审理好上市公司破产重整
案件工作座谈会纪要》第18条第三款的规定,经参与表决
的出资人所持表决权三分之二以上通过的,即为该组通过重
整计划(草案)。
(二)重整计划(草案)的批准
普通债权组表决通过重整计划(草案),且出资人组表
决通过出资人权益调整事项时,重整计划(草案)即为通过。
债务人将依法向常德中院提出批准重整计划的申请。
部分表决组未通过重整计划(草案)的,债务人或者管
理人可以同未通过重整计划(草案)的表决组协商。该表决
组可以在协商后再表决一次。双方协商的结果不得损害其他
表决组的利益。未通过重整计划(草案)的表决组拒绝再次
表决或者再次表决仍未通过重整计划(草案),但重整计划
(草案)符合《企业破产法》第八十七条规定的,债务人或
者管理人可以申请人民法院批准重整计划(草案)。
(三)重整计划批准生效
重整计划(草案)在依据《企业破产法》第八十四条至
第八十七条的相关规定,由各表决组表决通过并经常德中院
裁定批准后生效,或部分表决组表决虽未通过但经常德中院
裁定批准后生效。
(四)未获批准的后果
重整计划(草案)未获得债权人会议通过并且未依照《企
业破产法》第八十七条的规定获得常德中院批准;或已经表
决通过的重整计划未获得法院裁定批准的,管理人将依法申
请常德中院裁定终止重整程序,并宣告景峰医药破产。
八、重整计划的执行
本重整计划由景峰医药负责执行。
(一)执行期限
本重整计划的执行期限为常德中院裁定批准重整计划
(草案)之日起6个月。在此期间,景峰医药应当严格依照
重整计划的规定清偿债务,并随时支付破产费用。
(二)执行期限的延长与提前
如非景峰医药自身原因,致使本重整计划无法在上述期
限内执行完毕,景峰医药应于执行期限届满前,向常德中院
提交延长重整计划执行期限的申请,并根据常德中院批准的
执行期限继续执行。
(三)执行完毕的标准
自下列条件全部满足之日起,重整计划视为执行完毕:
1.管理人收到重整投资人受让转增股票的全部现金对价
款;
2.用于引入重整投资人的转增股票已登记至管理人开立
的证券账户或已经登记至相关投资人或其指定主体名下的
证券账户;
3.破产费用已经支付完毕或已经提存至管理人账户;各
类债权已经按照债权调整和清偿方案获得清偿,或用于清偿
的现金已提存至管理人账户。
(四)司法协助执行事项
重整计划执行过程中,涉及需要有关单位协助执行的,
景峰医药及/或管理人可向常德中院提出申请,请求常德中院
向有关单位出具要求其协助执行的司法文书。
(五)执行完毕的效力
根据重整计划予以豁免的债权,景峰医药不再承担清偿
责任。
(六)重整计划的变更
本重整计划执行过程中,因遇国家政策调整、法律修改
变化等特殊情况或发生意外事件或不可抗力致使本重整计
划无法继续执行的,可由管理人召集债权人会议,或由债务
人向人民法院申请召开债权人会议,就是否同意变更重整计
划进行表决。债权人会议决议同意变更重整计划的,债务人
应自决议通过之日起十日内提请常德中院批准。常德中院裁
定批准变更重整计划的,景峰医药或者管理人应当在六个月
内提出新的重整计划。变更后的重整计划应提交给因重整计
划变更而遭受不利影响的债权人组和出资人组进行表决。变
更后的重整计划在权益受到调整或影响的债权人组及/或出
资人组表决通过并获得法院裁定批准后,由景峰医药执行变
更后的重整计划,管理人予以监督。
虽有上述约定,若重整计划(草案)在经债权人会议表
决通过后常德中院裁定批准前或重整计划在执行过程中,需
变更重整财务投资人、上调受让股票价格,或调整清偿方案
的,在调整后的方案不减损债权人利益或对债权人更有利的
情况下,债务人可直接予以变更,无需通过债权人会议进行
表决。调整后的重整计划(草案)或重整计划由管理人向常
德中院报备。
重整计划确系无法通过修改等方式实现继续执行的,常
德中院可应管理人或利害关系人的请求裁定终止重整计划
的执行,并宣告景峰医药破产。
(七)有关重整计划执行的重大不确定事项
重整计划执行过程中,可能会出现投资人违约、无法转
增股票等风险。
九、重整计划执行的监督
管理人负责监督重整计划的执行。
(一)监督期限
本重整计划执行的监督期限与重整计划的执行期限相
同,自常德中院裁定批准重整计划之日起计算。
(二)监督期限的延长与提前
如根据重整计划执行的实际情况,需要延长管理人监督
重整计划的执行期限的,则管理人将向常德中院提交延长重
整计划执行监督期限的申请,并根据常德中院批准的期限继
续履行监督职责。
重整计划执行期限提前到期的,执行监督期限相应提前
到期。
(三)监督期限的终止
监督期限届满或公司执行完毕本重整计划时,管理人将
向常德中院提交监督报告,自监督报告提交之日起,管理人(未完)
2026-02-03
·新浪财经
湖南景峰医药股份有限公司重整计划湖南景峰医药股份有限公司二〇二六年一月目录释义 ...... 5前言 ...... 10摘要 ...... 12正文 ...... 14一、景峰医药基本情况 ...... 14(一)基本信息 ...... 14(二)股本结构 ...... 14(三)预重整及重整的申请及受理情况 ...... 15(四)资产情况 ...... 16(五)负债情况 ...... 17(六)偿债能力分析 ...... 18二、出资人权益调整方案 ...... 19(一)出资人权益调整的必要性、合理性 ...... 19(二)出资人权益调整的范围 ...... 20(三)出资人权益调整的内容 ...... 21(四)除权与除息 ...... 21(五)出资人权益调整方案的执行效果 ...... 22三、债权分类及调整方案 ...... 23(一)债权分类方案 ...... 23(二)债权调整方案 ...... 23四、债权清偿方案 ...... 25(一)偿债资源 ...... 25(二)债权清偿方案 ...... 25(三)劣后债权清偿方案 ...... 26(四)合并报表范围内关联债权的清偿方案 ...... 26(五)预计债权清偿方案 ...... 26五、重整投资人情况 ...... 28(一)重整投资人的招募情况 ...... 28(二)重整投资人具体情况 ...... 29(三)重整投资人受让转增股份情况 ...... 37六、经营方案 ...... 39(一)优化存量,培育增量,创新引领、双轮驱动 ...... 39(二)优化组织架构,提升管理水平,降本增效 ...... 45七、重整计划(草案)的表决和批准 ...... 47(一)重整计划(草案)的表决 ...... 47(二)重整计划(草案)的批准 ...... 48(三)重整计划批准生效 ...... 48(四)未获批准的后果 ...... 48八、重整计划的执行 ...... 49(一)执行期限 ...... 49(二)执行期限的延长与提前 ...... 49(三)执行完毕的标准 ...... 49(四)司法协助执行事项 ...... 49(五)执行完毕的效力 ...... 50(六)重整计划的变更 ...... 50(七)有关重整计划执行的重大不确定事项 ...... 51九、重整计划执行的监督 ...... 51(一)监督期限 ...... 51(二)监督期限的延长与提前 ...... 51(三)监督期限的终止 ...... 51十、其他说明情况 ...... 52(一)重整计划的生效 ...... 52(二)偿债资源的提存及预留、处理 ...... 52(三)偿债资源的分配 ...... 53(四)财产保全措施的解除及信用等级的修复 ...... 54(五)债权人对其他还款义务人的权利 ...... 54(六)对实控人原有股权调整的方案 ...... 55(七)破产费用的支付及共益债务的清偿 ...... 55(八)转让债权的清偿 ...... 56(九)重整计划的解释 ...... 56释义景峰医药/公司/债务人指湖南景峰医药股份有限公司常德中院指湖南省常德市中级人民法院深交所指深圳证券交易所中证登深圳分公司指中国证券登记结算有限责任公司深圳分公司临时管理人、管理人指由常德中院指定的景峰医药临时管理人、管理人债权人指符合《企业破产法》第四十四条之规定的,景峰医药的某个、部分或全体债权人出资人指出资人组会议召开通知中所载明的股权登记日在中证登深圳分公司登记在册的景峰医药股东重整投资人指经公开招募后确定的,与景峰医药管理人、景峰医药签署《重整投资协议》和/或《重整投资协议之补充协议》的主体石药控股指石药控股集团有限公司德源招商指常德市德源招商投资有限公司重整计划(草案)指依据《企业破产法》第七十九条之规定,债务人制作并提交的重整计划(草案)重整计划指重整计划(草案)获债权人会议及出资人组表决通过,或虽未获表决通过但经常德中院裁定批准后的重整计划破产费用指《企业破产法》第四十一条规定的,人民法院受理破产申请后发生的相关费用共益债务指《企业破产法》第四十二条规定的,人民法院受理破产申请后发生的相关债务有财产担保债权指依据《企业破产法》第八十二条第一款第(一)项之规定,对景峰医药特定财产享有担保权的债权职工债权指依据《企业破产法》第八十二条第一款第(二)项等相关规定,包括景峰医药所欠职工的工资和医疗、伤残补助、抚恤费用,所欠的应当划入职工个人账户的基本养老保险、基本医疗保险费用,法律、行政法规规定应当支付给职工的补偿金普通债权指依据《企业破产法》第八十二条第一款第(四)项之规定,债权人对景峰医药享有的债权审查确定的债权指债权申报期限内经债权人申报并经管理人依法审查确定的债权未申报债权指与债务人构成债权债务关系,未在重整计划(草案)提交债权人会议表决前向管理人依法申报但可能受法律保护的债权暂缓确定债权指已向管理人申报但截至重整计划(草案)提交之日,因诉讼未决、需要补充证据材料、债权人提出异议等原因尚未经管理人审查确定的债权转增股票指根据本重整计划规定的出资人权益调整方案,以景峰医药股本879,774,351股为基数,实施资本公积转增股本形成的股票提存指对应偿债资源按照本重整计划规定暂支付至管理人或管理人指定方名下,视为提存完毕评估机构指为景峰医药重整提供资产评估和偿债能力分析服务的北京中企华资产评估有限责任公司审计机构指为景峰医药重整提供审计服务的大信会计师事务所(特殊普通合伙)《偿债能力分析报告》指评估机构以2025年9月30日为基准日出具的《湖南景峰医药股份有限公司重整项目偿债能力分析报告》(中企华评咨字(2025)第5769号)《资产清算价值报告》指评估机构以2025年9月30日为基准日出具的《湖南景峰医药股份有限公司重整涉及的资产清算价值项目评估咨询报告》(中企华评咨字(2025)第5768号)《资产市场价值报告》指评估机构以2025年9月30日为基准日出具的《湖南景峰医药股份有限公司重整涉及的资产市场价值项目资产评估报告》(中企华评报字(2025)第5766号)《审计报告》指审计机构以2025年9月30日为基准日出具的《湖南景峰医药股份有限公司审计报告》(大信专审字[2025]第39-00008号)重整计划的通过指依据《企业破产法》第八十六条第一款之规定,债权人会议各表决组及出资人组会议均通过重整计划(草案)时,重整计划即为通过重整计划的批准指依据《企业破产法》第八十六条第二款或第八十七条第三款之规定,重整计划获得常德中院裁定批准重整计划的执行期限指根据《企业破产法》第八十一条第(五)项之规定,在景峰医药重整计划中载明的执行期限及法院裁定延长的重整计划执行期限重整计划执行监督期限指依据《企业破产法》第九十条之规定,本重整计划规定的管理人监督重整计划执行的期限《企业破产法》指自2007年6月1日起施行的《中华人民共和国企业破产法》元指本重整计划中除特别注明外,均为人民币元前言景峰医药为深交所上市公司,主营业务为以自有资产进行医药、医疗项目投资;生物制药技术项目的研发与投资;商品进出口贸易;企业管理咨询、医疗医药研发技术咨询(依法须经批准的项目,经相关部门批准后方可开展经营活动)。由于经营不善,景峰医药目前生产经营不佳,无法清偿到期负债,且已不具备债务清偿能力。常德中院根据债权人申请,于2024年7月2日对景峰医药启动预重整并指定北京市中伦律师事务所作为公司预重整期间的临时管理人,组织开展预重整各项工作。预重整期间,经过公开招募和遴选程序,临时管理人、景峰医药分别与石药控股、德源招商签署了《重整投资协议》。2025年10月21日,常德中院作出(2024)湘07破申7号《民事裁定书》,裁定受理彭东钜、上海鑫绰投资管理有限公司对景峰医药的重整申请,并同日作出(2025)湘07破15号《决定书》,指定北京市中伦律师事务所担任湖南景峰医药股份有限公司重整管理人,具体开展重整各项工作。景峰医药的重整工作得到了政府及常德中院的高度重视和大力支持。为保证重整成功,避免景峰医药破产清算,管理人在常德中院的监督和指导下,严格遵照《企业破产法》的规定全面履行相关职责。预重整期间,临时管理人启动债权、资产的调查工作,进行了第一轮债权审核,完成了重整投资人招募工作,为进入法院正式重整做好准备工作,节省正式重整时间、提高成功率。重整期间,管理人监督公司自行管理财产和营业事务,确保公司生产经营及职工稳定,保障重整工作的顺利推进;同时,景峰医药及管理人全力以赴做好与重整相关的各项具体工作,包括债权申报受理与审查、财产调查、职工债权调查、重整投资人协商谈判、重整计划(草案)的论证和制作等。在景峰医药预重整及重整各项工作推进过程中,常德中院均依法监督,确保重整程序依法合规开展,切实保障各方主体的合法权益。截至目前,管理人已经完成重整所需各项基础工作,对景峰医药的整体现状已有全面了解。在充分听取债权人、重整投资人、股东等各方意见和建议的基础上,在充分尊重评估机构出具的《资产市场价值报告》《资产清算价值报告》及《偿债能力分析报告》结论的前提下,在充分进行法律上的风险评估和论证、可行性预判和分析的条件下,景峰医药严格依据《企业破产法》第七十九条、第八十条、第八十一条之规定,在法律、法规及司法解释允许的范围内,结合景峰医药实际情况,制作重整计划(草案),供债权人会议审议、表决,并由出资人组会议对重整计划(草案)中涉及的出资人权益调整事项进行表决。摘要一、重整完成后,景峰医药的企业法人性质及市场主体资格不变,并将最大可能消除债务危机及其他风险,提升上市公司质量。二、本重整计划将以景峰医药现有股本879,774,351股为基数,按照每10股转增10股的比例实施资本公积金转增股本,共计可转增879,774,351股股票(最终实际转增的股票数量以中证登深圳分公司实际登记确认的数量为准)。转增后,景峰医药总股本将增至1,759,548,702股(最终转增的准确股票数量以中证登深圳分公司实际登记确认的数量为准)。前述转增的879,774,351股股票不再向原股东分配,全部用于引入重整投资人,并由重整投资人提供资金认购,相应资金用于根据重整计划的规定支付破产费用、清偿各类债务、补充公司流动资金等。三、职工债权、税款债权不作调整,在法院裁定批准重整计划且债权经法院裁定确认后20个工作日内以现金方式一次性清偿完毕。四、普通债权的清偿方案如下:普通债权以债权人为单位,每家普通债权人的普通债权在200万元以下(含200万元)的部分,由景峰医药在后续重整计划执行期限内以现金方式一次性清偿完毕。每家普通债权人的普通债权超过200万元的部分,可选择如下方案进行受偿:(1)留债方案针对每家普通债权人的普通债权超过200万元的部分,以现金方式予以留债清偿,留债最长期限为2年,留债期限内不计息。留债期限内,不影响债权人根据有关合同约定以及法律规定,就其债权未获清偿部分(含合同约定或法定的利息)向其他债务人(包括但不限于主债务人)主张有关权益,亦不影响其他担保人或主债务人根据有关合同约定以及法律规定向债权人清偿债务。(2)现金清偿方案每家普通债权人按90%的清偿率在法院裁定批准重整计划后90日内获得现金清偿;剩余未获清偿债权部分在重整计划执行完毕后豁免清偿,景峰医药不再承担清偿责任。债权金额超过200万元的普通债权人(含暂缓确认债权人),应自法院裁定批准重整计划之日起10个工作日内按照重整计划要求的书面格式提供债权清偿方式选择告知书。债权人逾期未告知的,视为全额选择按以普通债权清偿方式中的“现金清偿方案”获得清偿。本重整计划执行完毕后,全体债权人的债权将得到较高清偿,公司的财务状况将得到根本改善,在减轻债务负担的同时,公司可以有效提升经营效率和盈利能力,全体投资者的利益将有望得到有效保护。上述为本重整计划核心内容的摘录或总结,具体内容文义以正文表述为准。正文一、景峰医药基本情况(一)基本信息景峰医药注册地址为湖南省常德经济技术开发区樟木桥街道双岗社区桃林路661号(双创大厦1703室),营业期限为1998年12月18日至无固定期限,统一社会信用代码为914306007121062680,法定代表人为刘树林,注册资本为87,977.4351万元人民币,总股本879,774,351股。景峰医药是一家在深圳证券交易所A股公开发行股票的上市公司,证券代码为000908。景峰医药经营范围为:以自有资产进行医药、医疗项目投资;生物制药技术项目的研发与投资;商品进出口贸易;企业管理咨询、医疗医药研发技术咨询(依法须经批准的项目,经相关部门批准后方可开展经营活动)。(二)股本结构1.前十大股东截至2025年9月30日,景峰医药总股本为879,774,351股,期末股东总数为27,854户,前10大股东持股情况如下:股东名称/姓名股数(股)持股比例中国长城资产管理股份有限公司113,680,66512.92%叶湘武112,252,28612.76%平江县国有资产事务中心11,083,3691.26%武阳7,419,8140.84%赵建锋5,675,0000.65%股东名称/姓名股数(股)持股比例陈永华5,268,1100.60%陆琦5,000,0000.57%侯健4,951,6000.56%孙江波4,601,0000.52%杜永生3,953,4870.45%2.实际控制人及其持股情况截至目前,叶湘武及其一致行动人合计持有公司股份数量为114,684,310股,占公司股份总数的13.04%,景峰医药的实际控制人为叶湘武。(三)预重整及重整的申请及受理情况2024年6月26日,申请人彭东钜、上海鑫绰投资管理有限公司向常德中院申请对景峰医药进行重整,并同时申请对景峰医药启动预重整。2024年7月2日,常德中院作出(2024)湘07破申7号《决定书》,常德中院决定对景峰医药启动预重整。2024年7月30日,常德中院作出(2024)湘07破申7号之一《决定书》,指定北京市中伦律师事务所担任景峰医药预重整期间临时管理人。2025年10月21日,常德中院作出(2024)湘07破申7号《民事裁定书》,裁定受理债权人对景峰医药的重整申请,并于同日作出(2025)湘07破15号《决定书》,指定北京市中伦律师事务所为景峰医药重整管理人。(四)资产情况1.账面资产情况截至2025年9月30日,景峰医药(母公司单体,下同)的资产总额为11.41亿元。景峰医药主要资产由长期股权投资、其他应收款等构成。景峰医药截至2025年9月30日的账面资产明细情况具体如下:类别账面价值(万元)占比流动资产合计11,198.939.82%其中:其他应收款10,658.339.34%其他流动资产482.170.42%货币资金58.430.05%非流动资产合计102,867.8490.18%其中:长期股权投资100,126.2087.78%无形资产2,735.782.40%固定资产5.860.01%资产合计114,066.77100.00%2.资产评估情况根据评估机构出具的《资产市场价值报告》,截至2025年9月30日,基于重整状态下的持续经营假设为前提,景峰医药总资产的评估价值为59,180.50万元,具体如下:类别市场价值(万元)流动资产合计10,930.94非流动资产合计48,249.56其中:长期股权投资45,297.68固定资产16.96无形资产2,934.92类别市场价值(万元)资产合计59,180.50(五)负债情况结合审计机构出具的《审计报告》等,并经管理人对景峰医药债权申报的审查,对于已申报债权,管理人已进行审查认定,对于未申报债权,管理人已作预留相应偿债资源。景峰医药负债情况具体如下:1.债权申报情况截至目前,共有60家债权人向管理人申报了债权(不含职工债权),申报债权总额为233,295.05万元,其中,2家债权人申报债权性质为税款及社保债权,对应申报金额为137.94万元,4家债权人申报债权性质为有财产担保的债权(含一家债权人申报债权性质为建设工程优先权债权),对应申报金额为92,467.05万元,54家债权人申报债权性质为普通债权,对应申报金额为140,690.06万元。2.债权审查情况(1)审查确定的债权债权人已申报债权中,截至重整计划(草案)披露日,经管理人审查确定的债权总额为97,750.52万元,涉及债权人43家。其中包括1家税款债权,债权金额为131.77万元;42家普通债权,债权金额为97,618.75万元。(2)暂缓确认债权截至重整计划(草案)披露日,不存在暂缓确认债权。(3)不予确认的债权债权人已申报债权中,截至重整计划(草案)披露日,不予确认的债权对应41家债权人申报的债权(包含部分不予确认的情形),涉及债权申报金额为135,521.13万元。需要说明的是,对于在景峰医药预重整阶段申报的附利息债权,管理人已将相应利息追加计算至重整受理日前一日。债权最终审查结论以经常德中院裁定确认的债权表中记载的为准。3.职工债权调查情况经管理人调查,截至2025年10月20日,景峰医药的职工债权总额为272.81万元。4.税款债权情况经管理人调查,截至2025年10月20日,景峰医药的税款债权为131.77万元,涉及1家债权人。5.未申报债权情况截至2025年10月20日,根据审计机构出具的《审计报告》、公司的说明及管理人调查梳理,未在重整计划(草案)提交债权人会议表决前申报但可能受法律保护的债权总额约8,088万元。(六)偿债能力分析为给债权人表决重整计划(草案)提供必要参考,管理人委托评估机构对景峰医药在假定破产清算条件下的偿债能力进行了分析。根据评估机构出具的《偿债能力分析报告》,截至评估基准日,在模拟破产清算状态下,假定景峰医药全部资产能够按清算价值实际变现,按照《企业破产法》规定的清偿顺序,破产财产的变现所得在支付必要的破产费用、共益债务、职工债权、有财产担保的债权后,普通债权清偿率约为21.61%。考虑到景峰医药如实施破产清算,能够达到《偿债能力分析报告》中普通债权清偿率的前提,一是资产处置时能够按照评估价值变现,二是处置税费等破产费用能够控制在管理人和评估机构预测的范围内。同时,司法实践中破产清算程序耗时可能较长,将进一步产生超过预期的各项费用。基于以上因素,景峰医药实际在破产清算状态下可用于偿债的资金将可能比《偿债能力分析报告》预计的更低,导致债权人的利益进一步受损。而在本次重整中,对于普通债权的200万元以下(含200万元)部分,将全额现金清偿,因此该部分普通债权,清偿率为100%。对于普通债权超过200万元的部分,在“现金清偿方案”中债权人超过200万元的部分为现金清偿,清偿率为90%;在“留债方案”中债权人超过200万元的债权,留债最长期限为2年,留债清偿后预计清偿率为100%。二、出资人权益调整方案(一)出资人权益调整的必要性、合理性根据最高人民法院、中国证监会《关于切实审理好上市公司破产重整案件工作座谈会纪要》“五、关于上市公司破产重整计划草案的制定及表决”之“16.出资人权益调整”中规定,“上市公司资产不足以清偿全部债务且普通债权人不能在重整计划中全额获得清偿的,原则上应对出资人权益进行调整。”鉴于景峰医药已不能清偿到期债务,且明显缺乏清偿能力,生产经营和财务状况均已陷入困境。同时,普通债权人不能在重整计划中全额获得清偿。因此,本重整计划安排对出资人权益进行调整。景峰医药在确定出资人权益调整方案时主要考虑了如下因素:1.本重整计划偿债方案中所需的偿债资源。2.景峰医药经营方案中涉及未来经营恢复、业务升级、内部管理系统改造升级等所需的资金。3.充分考虑中小投资者利益,避免过度转增导致对中小投资者权益的稀释。综合上述因素,本次出资人权益调整以景峰医药现有总股本为基础,按照每10股转增10股的比例实施资本公积转增股本。该安排符合《上市公司监管指引第11号——上市公司破产重整相关事项》的规定。(二)出资人权益调整的范围根据《企业破产法》第八十五条第二款之规定,重整计划(草案)涉及出资人权益调整事项的,应当设出资人组,对该事项进行表决。景峰医药出资人组由截至出资人组会议召开公告所载明确定的股权登记日在中证登深圳分公司登记在册的景峰医药股东组成。上述股东在出资人组会议之股权登记日后至本出资人权益调整方案实施完毕前,由于交易或非交易原因导致持股情况发生变动的,本重整计划规定的出资人权益调整方案的效力及于其股票的受让方及/或承继方。(三)出资人权益调整的内容本次权益调整以景峰医药现有总股本879,774,351股为基数,按照每10股转增10股的比例实施资本公积金转增股本,共计可转增879,774,351股股票(最终实际转增的股票数量以中证登深圳分公司实际登记确认的数量为准)。转增后,景峰医药总股本将增至1,759,548,702股。上述转增股票不向原股东分配,全部由重整投资人支付现金对价受让,具体受让股票数以后续重整计划执行阶段的司法协助执行通知书载明的内容及中证登深圳分公司实际登记确认的数量为准。(四)除权与除息重整计划(草案)经法院裁定批准后执行,根据本重整计划规定实施的资本公积转增股本将全部用于引进重整投资人;本次转增后,公司在总股本扩大的同时债务规模明显减少、所有者权益明显增加;公司原每股股票所代表的企业实际价值(以每股净资产计算)较重整前显著提升。这与转增前后公司所有者权益不变、需要按照《深圳证券交易所交易规则(2023修订)》第4.4.2条中载明的除权(息)参考价计算公式对股票价格进行调整的一般情形存在本质差别。因此,重整计划实施后,为反映出资人权益调整事项对景峰医药股票价值的影响,需要对本次资本公积金转增股票除权计算公式进行调整,并可能对本次资本公积转增股权登记日次一交易日的股票开盘参考价进行调整。根据《深圳证券交易所交易规则(2023修订)》第4.4.2条的规定:“除权(息)参考价计算公式为:除权(息)参考价=[(前收盘价-现金红利)+配股价格×股份变动比例]÷(1+股份变动比例),证券发行人认为有必要调整上述计算公式时,可以向本所提出调整申请并说明理由。经本所同意的,证券发行人应当向市场公布该次除权(息)适用的除权(息)参考价计算公式。”公司已聘请财务顾问对本次重整中拟实施资本公积转增股本除权参考价格的计算公式进行论证,最终以财务顾问出具的专项意见为准。如果股权登记日公司股票收盘价高于财务顾问专项意见中确认的本次重整资本公积转增股本的平均价的,公司股票将按除权后价格于股权登记日次一交易日调整开盘参考价。如果股权登记日公司股票收盘价格低于或等于财务顾问专项意见中确认的本次重整资本公积转增股本的平均价的,公司股权登记日次一交易日的股票开盘参考价无需调整。后续若上述拟调整的除权参考价格的计算公式或相关计算参数因裁定批准的重整计划或根据监管规则要求需要另行调整的,公司将按照前述要求进行调整。(五)出资人权益调整方案的执行效果根据上述出资人权益调整方案,景峰医药出资人所持有的公司股票绝对数量不会因本次重整而减少。重整完成后,景峰医药的基本面将发生根本性改善,并将逐步提升持续盈利能力,公司价值也将进一步提升,有利于保护公司、债权人及广大出资人的合法权益。三、债权分类及调整方案根据《企业破产法》规定,景峰医药债权将按照职工债权、税款债权、有财产担保债权、普通债权以及劣后债权进行分类,具体分类及调整原则如下:(一)债权分类方案1.职工债权经管理人调查,截至2025年10月20日,景峰医药的职工债权总额为272.81万元。2.税款债权经管理人审查,截至2025年10月20日,景峰医药的税款债权为131.77万元,涉及1家债权人。3.有财产担保债权经管理人审查,截至2025年10月20日,景峰医药无有财产担保债权人。4.普通债权经管理人审查确定,截至2025年10月20日,普通债权总额为97,618.75万元,共计42家债权人。5.劣后债权截至2025年10月20日,经管理人审查,景峰医药无劣后债权。(二)债权调整方案1.职工债权职工债权不作调整,在法院裁定批准重整计划且债权经法院裁定确认后20个工作日内以现金方式一次性清偿完毕。2.税款债权根据《企业破产法》第八十二条第一款第(三)项规定的景峰医药欠付的税款本金,在本次重整中不作调整,在法院裁定批准重整计划且债权经法院裁定确认后20个工作日内以现金方式一次性清偿完毕。3.有财产担保的债权有财产担保债权人通过设定财产担保或依据相关法律规定而对景峰医药特定财产享有优先受偿的权利,以担保财产持续经营状态下的评估价值确定优先受偿范围。若担保财产的评估价值不足以清偿所对应的有财产担保债权,则有财产担保债权超出担保财产评估价值的部分按照本重整计划规定的普通债权清偿方案受偿;若担保财产的评估价值超出所对应的有财产担保债权,则超出部分不属于该有财产担保债权人享有优先受偿权的范围。经管理人审查,截至2025年10月20日,景峰医药无有财产担保债权人。4.普通债权为最大限度地保护债权人合法权益、提高债权人的受偿水平,根据景峰医药实际情况,将按照常德中院裁定批准的重整计划规定的清偿方案予以清偿。5.劣后债权景峰医药在进入重整前产生的民事惩罚性赔偿金、行政罚款、刑事罚金、加倍支付迟延履行利息等惩罚性债权为劣后债权,不予清偿。四、债权清偿方案(一)偿债资源本次重整的偿债资源为重整投资者支付的重整投资款以及公司自有货币资金、未来持续经营收入等。(二)债权清偿方案1.职工债权职工债权由景峰医药在法院裁定批准重整计划且债权经法院裁定确认后20个工作日内以现金方式一次性清偿完毕。2.税款债权税款债权由景峰医药在法院裁定批准重整计划且债权经法院裁定确认后20个工作日内以现金方式一次性清偿完毕。3.普通债权普通债权以债权人为单位,每家普通债权人的普通债权在200万元以下(含200万元)的部分,由景峰医药在后续重整计划执行期限内以现金方式一次性清偿完毕。每家普通债权人的普通债权超过200万元的部分,可选择如下方案进行受偿:(1)留债方案针对每家普通债权人的普通债权超过200万元的部分,以现金方式予以留债清偿,留债最长期限为2年,留债期限内不计息。留债期限内,不影响债权人根据有关合同约定以及法律规定,就其债权未获清偿部分(含合同约定或法定的利息)向其他债务人(包括但不限于主债务人)主张有关权益,亦不影响其他担保人或主债务人根据有关合同约定以及法律规定向债权人清偿债务。(2)现金清偿方案每家普通债权人按90%的清偿率在法院裁定批准重整计划之日起90日内获得现金清偿;剩余未获清偿债权部分在重整计划执行完毕后豁免清偿,景峰医药不再承担清偿责任。债权金额超过200万元的普通债权人(含暂缓确认债权人),应自法院裁定批准重整计划之日起10个工作日内按照重整计划要求的书面格式提供债权清偿方式选择告知书。债权人逾期未告知的,视为全额选择按以普通债权清偿方式中的“现金清偿方案”获得清偿。(三)劣后债权清偿方案景峰医药涉及的民事惩罚性赔偿金、行政罚款、刑事罚金等惩罚性债权将劣后于其他普通债权。针对劣后债权的处理,由于普通债权未获得全额清偿,因此依法不安排偿债资源。(四)合并报表范围内关联债权的清偿方案景峰医药合并范围内关联债权人对景峰医药享有的债权,按照本重整计划中同类债权的清偿方案予以清偿。(五)预计债权清偿方案1.暂缓确认债权已向管理人申报,但因涉及未决诉讼等原因而尚未经管理人审查确定的债权,其中普通债权人应当提前就剩余债权(剔除200万元现金清偿部分)选择何种方案清偿以书面形式加以明确,在债权经审查确定后10个工作日内按照本重整计划要求的书面格式提供债权清偿方式选择告知书。债权人逾期未告知的,视为全额选择按以普通债权清偿方式中的“现金清偿方案”获得清偿。除因诉讼仲裁未决等原因而暂未确认的债权外,在重整计划(草案)提交债权人会议表决前,已依法申报但因各种原因在后续重整程序中暂未确认的债权,如在后续重整计划执行期内仍未获得确认,则由景峰医药在后续重整程序终结后,在不损害其他债权人权益的情况下,通过包括诉讼、仲裁、协商等适当方式审查确定债权金额。因诉讼仲裁未决而暂未确认的债权,依法院或仲裁机构的生效法律文书确定债权金额和性质。上述暂未确认债权获得确认后10个工作日内按照重整计划要求的书面格式提供债权清偿方式选择告知书。债权人逾期未告知的,视为全额选择按以普通债权清偿方式中的“现金清偿方案”获得清偿。2.未申报债权对于景峰医药账面有记载但未申报的债权,本重整计划已预留相应偿债资源。未申报债权在本重整计划执行期间不得行使权利;在本重整计划执行完毕后,该部分未申报债权经申报并经债务人审查确定后,可要求景峰医药按照本重整计划中规定的同类债权受偿方式予以清偿。根据《企业破产法》第五十六条规定,为审查和确认补充申报债权的费用,由补充申报人承担。未依法申报债权的债权人,自重整受理日起三年内或至该等债权的诉讼时效届满之日(以孰早为准),未向景峰医药主张权利的,景峰医药不再负有清偿义务。如因客观因素需调整未申报债权受偿方式的,可由公司与前述债权人另行协商。五、重整投资人情况(一)重整投资人的招募情况2024年8月1日,公司发布《湖南景峰医药股份有限公司临时管理人关于意向投资者招募的公告》,临时管理人通过全国企业破产重整案件信息网同步发布公开招募重整投资人的公告(以下简称“招募公告”),正式启动投资人公开招募工作。截至2024年8月15日,共有4家投资人(联合体按1家计算)提交报名材料。截至2024年8月25日,共有1家投资人(联合体按1家计算)缴纳尽调保证金。根据《关于招募重整投资人延期的公告》,若仅有一名意向投资人或投资人联合体提交重整投资方案,临时管理人将组织通过商业谈判方式确定重整投资人。2024年8月27日,景峰医药发布《关于确定预重整投资人暨风险提示的公告》,最终确定以石药控股作为牵头投资人的联合体为中选重整投资人。2025年4月29日,景峰医药发布《关于与重整投资人签署重整投资协议的公告》,披露了景峰医药、临时管理人与石药控股及作为联合体成员之一的德源招商签署的重整投资协议,其中包括投资人认购股份数量及价格条款。2026年1月9日,管理人、景峰医药与石药控股及德源招商分别签署《重整投资协议之补充协议》,对投资价格及投资款支付期限予以补充约定。2026年1月9日,管理人、景峰医药与中国长城资产管理股份有限公司签署《重整投资协议》。经牵头投资人石药控股、管理人等各方与意向投资人沟通磋商,2026年1月9日,管理人、景峰医药与16家财务投资人签署《重整投资协议》。债务人已就重整投资人招募及重整投资协议签署情况履行了信息披露义务。(二)重整投资人具体情况经过公开招募和遴选,确定以石药控股作为牵头投资人的联合体为景峰医药的重整投资人。根据重整投资人的说明,重整投资人基本情况介绍如下:1.石药控股集团有限公司石药控股集团有限公司成立于1998年3月31日,统一社会信用代码为911301002360444643。石药控股的实际控制人为蔡东晨。景峰医药现任董事及总裁刘树林,曾任石药控股集团有限公司副总裁兼第一制造中心总裁、总计划调度中心总经理等职务。景峰医药现任董事马学红曾任石药控股集团有限公司财务部税务主管、税务经理;河北中润制药有限公司财务部经理、石药集团欧意药业有限公司财务总监、石药控股集团有限公司资产管理中心高级总监。景峰医药现任董事廉奇志曾任石药集团有限公司高级总监、资本运营中心总经理等职务,现任石药控股集团河北省教育科技有限公司董事。除此之外,石药控股与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。重整完成后,石药控股将成为景峰医药的控股股东。石药控股承诺自受让转增股票之日起36个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。2.常德瑞健一禾咨询管理合伙企业(有限合伙)常德瑞健一禾咨询管理合伙企业(有限合伙)成立于2025年11月,统一社会信用代码为91430700MAK0Y24L1W。其实际控制人为徐铭泽。常德瑞健一禾咨询管理合伙企业(有限合伙)与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。常德瑞健一禾咨询管理合伙企业(有限合伙)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。3.常德智恒咨询管理合伙企业(有限合伙)常德智恒咨询管理合伙企业(有限合伙)成立于2025年11月,统一社会信用代码为91430700MAK0XCJA9B。其实际控制人为贾路琦。常德智恒咨询管理合伙企业(有限合伙)与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。常德智恒咨询管理合伙企业(有限合伙)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。4.常德市德源招商投资有限公司常德市德源招商投资有限公司成立于2012年11月9日,统一社会信用代码为91430700055844286Y。德源招商为常德市德源投资集团有限公司的全资子公司,实际控制人为常德市人民政府国有资产监督管理委员会。德源招商投资事务部部长张莉现任景峰医药代行董事长。除此之外,德源招商与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。德源招商承诺自受让转增股票之日起36个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。5.常德金洋洪赢商务服务合伙企业(有限合伙)常德金洋洪赢商务服务合伙企业(有限合伙)成立于2024年12月,统一社会信用代码为91430700MAE748723N。其实际控制人为江西省人民政府。常德金洋洪赢商务服务合伙企业(有限合伙)与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。常德金洋洪赢商务服务合伙企业(有限合伙)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。6.宁波量利私募基金管理有限公司(代表宁波量利私募基金管理有限公司-量利元启1号私募证券投资基金、宁波量利私募基金管理有限公司-量利元启4号私募证券投资基金、宁波量利私募基金管理有限公司-量利元玺3号私募证券投资基金)宁波量利私募基金管理有限公司成立于2015年4月,统一社会信用代码为9133021234060269X8。其实际控制人为何振权。宁波量利私募基金管理有限公司为宁波量利私募基金管理有限公司-量利元启1号私募证券投资基金、宁波量利私募基金管理有限公司-量利元启4号私募证券投资基金、宁波量利私募基金管理有限公司-量利元玺3号私募证券投资基金的管理人。宁波量利私募基金管理有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。宁波量利私募基金管理有限公司(代表宁波量利私募基金管理有限公司-量利元启1号私募证券投资基金、宁波量利私募基金管理有限公司-量利元启4号私募证券投资基金、宁波量利私募基金管理有限公司-量利元玺3号私募证券投资基金)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。7.中国长城资产管理股份有限公司中国长城资产管理股份有限公司成立于1999年11月,统一社会信用代码为91110000710925489M。其实际控制人为国务院国资委。中国长城资产管理股份有限公司现为景峰医药的股东,持股数量为113,680,665股,持股比例为12.92%。景峰医药现任董事谢树青,现任中国长城资产管理股份有限公司湖南省分公司资产经营二部一级业务主管;景峰医药原监事会主席纪纲(离任未满12个月),现任中国长城资产管理股份有限公司资产经营六部高级经理。除此之外,中国长城资产管理股份有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。中国长城资产管理股份有限公司承诺自受让转增股票之日起36个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。8.湖南财鑫资本管理有限公司湖南财鑫资本管理有限公司成立于2014年1月,统一社会信用代码为914307000919795759。其实际控制人为常德市财政局。湖南财鑫资本管理有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。湖南财鑫资本管理有限公司承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。9.安徽明泽投资管理有限公司(代表安徽明泽投资管理有限公司-明泽远见6号债券私募证券投资基金)安徽明泽投资管理有限公司成立于2014年6月,统一社会信用代码为91440300306094812W。其实际控制人为马科伟。安徽明泽投资管理有限公司为安徽明泽投资管理有限公司-明泽远见6号债券私募证券投资基金的管理人。安徽明泽投资管理有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。安徽明泽投资管理有限公司(代表安徽明泽投资管理有限公司-明泽远见6号债券私募证券投资基金)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。10.常德顺行至通咨询管理合伙企业(有限合伙)常德顺行至通咨询管理合伙企业(有限合伙)成立于2025年11月,统一社会信用代码为91430700MAK0MEQ233。其实际控制人为王立平。常德顺行至通咨询管理合伙企业(有限合伙)与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。常德顺行至通咨询管理合伙企业(有限合伙)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。11.湖南景一企业管理咨询合伙企业(有限合伙)湖南景一企业管理咨询合伙企业(有限合伙)成立于2025年11月,统一社会信用代码为91430700MAK0P7578Q。其实际控制人为长沙市人民政府国有资产监督管理委员会。湖南景一企业管理咨询合伙企业(有限合伙)与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。湖南景一企业管理咨询合伙企业(有限合伙)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。12.青岛祺顺投资管理有限公司(代表青岛祺顺投资管理有限公司-祺顺扬帆19号私募证券投资基金、青岛祺顺投资管理有限公司-祺顺扬帆5号私募证券投资基金)青岛祺顺投资管理有限公司成立于2019年11月,统一社会信用代码为91370285MA3QYD4TX8。其实际控制人为宋耀。青岛祺顺投资管理有限公司为青岛祺顺投资管理有限公司-祺顺扬帆19号私募证券投资基金、青岛祺顺投资管理有限公司-祺顺扬帆5号私募证券投资基金的管理人。青岛祺顺投资管理有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。青岛祺顺投资管理有限公司(代表青岛祺顺投资管理有限公司-祺顺扬帆19号私募证券投资基金、青岛祺顺投资管理有限公司-祺顺扬帆5号私募证券投资基金)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。13.杭州源铨投资管理有限公司(代表杭州源铨投资管理有限公司-源铨东湖6号私募债券投资基金、杭州源铨投资管理有限公司-源铨东湖9号私募证券投资基金)杭州源铨投资管理有限公司成立于2017年5月,统一社会信用代码为91330106MA28T09Q6U。其实际控制人为石良希。杭州源铨投资管理有限公司为杭州源铨投资管理有限公司-源铨东湖6号私募债券投资基金、杭州源铨投资管理有限公司-源铨东湖9号私募证券投资基金的管理人。杭州源铨投资管理有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。杭州源铨投资管理有限公司(代表杭州源铨投资管理有限公司-源铨东湖6号私募债券投资基金、杭州源铨投资管理有限公司-源铨东湖9号私募证券投资基金)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。14.常德德润产业发展有限公司常德德润产业发展有限公司成立于2022年5月,统一社会信用代码为91430700MABNRUDE48。其实际控制人为常德经济技术开发区财政局。常德德润产业发展有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。常德德润产业发展有限公司承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。15.上海鑫绰投资管理有限公司(代表上海鑫绰投资管理有限公司-鑫绰鑫融3号私募证券投资基金)上海鑫绰投资管理有限公司成立于2014年1月,统一社会信用代码为913101150900697645。其实际控制人为周俊彦。上海鑫绰投资管理有限公司为上海鑫绰投资管理有限公司-鑫绰鑫融3号私募证券投资基金的管理人。上海鑫绰投资管理有限公司与景峰医药及其董事、高级管理人员、控股股东、实际控制人等不存在关联关系或者一致行动关系。上海鑫绰投资管理有限公司(代表上海鑫绰投资管理有限公司-鑫绰鑫融3号私募证券投资基金)承诺自受让转增股票之日起12个月内不转让或者委托他人管理其基于本次重整投资所直接和间接持有的景峰医药股票。(三)重整投资人受让转增股份情况依据管理人、景峰医药分别与重整投资人签订的协议,重整投资人合计受让879,774,351股转增股票,总投资额约为20.61亿元。重整投资人支付的投资款按照本重整计划之规定,用于包含但不限于支付破产费用、共益债务、破产债权以及相关支出。重整投资人受让股份情况如下:序号投资人名称受让股票数量(股)认购价格(元/股)受让股票总对价(元)1石药控股集团有限公司457,482,6621.45663,349,859.902常德瑞健一禾咨询管理合伙企业(有限合伙)70,558,0003.57251,892,060.003常德智恒咨询管理合伙企业(有限合伙)70,206,0003.57250,635,420.004常德市德源招商投资有限公司65,132,3051.88122,448,733.405常德金洋洪赢商务服务合伙企业(有限合伙)30,000,0003.57107,100,000.006宁波量利私募基金管理有限公司-量利元启1号私募证券投资基金25,810,7793.5792,144,481.037中国长城资产管理股份有限公司20,000,0003.5771,400,000.008湖南财鑫资本管理有限公司20,000,0003.5771,400,000.00序号投资人名称受让股票数量(股)认购价格(元/股)受让股票总对价(元)9安徽明泽投资管理有限公司-明泽远见6号债券私募证券投资基金18,747,2533.5766,927,693.2110常德顺行至通咨询管理合伙企业(有限合伙)17,595,3843.5762,815,520.8811湖南景一企业管理咨询合伙企业(有限合伙)15,000,0003.5753,550,000.0012宁波量利私募基金管理有限公司-量利元启4号私募证券投资基金14,568,6023.5752,009,909.1413青岛祺顺投资管理有限公司-祺顺扬帆5号私募证券投资基金14,110,7483.5750,375,370.3614杭州源铨投资管理有限公司-源铨东湖6号私募债券投资基金10,237,4703.5736,547,767.9015杭州源铨投资管理有限公司-源铨东湖9号私募证券投资基金9,868,8573.5735,231,819.4916常德德润产业发展有限公司8,800,0003.5731,416,000.0017宁波量利私募基金管理有限公司7,258,3963.5725,912,473.72序号投资人名称受让股票数量(股)认购价格(元/股)受让股票总对价(元)-量利元玺3号私募证券投资基金18青岛祺顺投资管理有限公司-祺顺扬帆19号私募证券投资基金3,097,8643.5711,059,374.4819上海鑫绰投资管理有限公司-鑫绰鑫融3号私募证券投资基金1,300,0313.574,641,110.67-合计879,774,351-2,060,857,594.18六、经营方案基于“优化存量,培育增量”的未来发展思路,景峰医药未来业务板块将包括中成药为主、化药为辅的存量业务,以及生物医药为主的增量业务,积极拓展合成生物产业。具体发展举措如下:(一)优化存量,培育增量,创新引领、双轮驱动1.聚焦存量优势产品,提升营销能力,创新创研新产品重整完成后,景峰医药将启动生产设备和生产过程的革新、研发方向与产业趋势的融合,提升管理水平,重塑品牌优势,打造具有国际先进水平的重点产业化项目。具体举措如下:(1)优化产品管线,聚焦存量核心产品公司产品种类丰富,未来将逐步淘汰长期缺乏竞争力的品种,按照潜力和目前的产品结构,公司将聚焦心血管领域产品,并持续不断优化产品结构。具体如下:一是公司将成立心血管销售部,聚焦心血管产品在等级医院的终端拓展、零售终端拓展及学术推广工作。心脑宁胶囊、参芎葡萄糖注射液、乐脉丸的等级医院渠道划归心血管销售部管理和销售;心脑宁和参芎除了稳定现有代理商的终端和纯销以外,要重点提升全国top100医院的学术推广覆盖和纯销提升速度,另一方面要积极拓展空白市场和代理商广度,设立终端开发目标进行考核,在精细化招商的指引下逐步提升终端数量和纯销数量,为未来销售夯实基础。公司将推动参芎葡萄糖注射液纳入国家医保目录作为景峰医药重点工作任务。二是对于重点产品心脑宁胶囊(全国独家),目前为医保乙类药品,未来拟研发增加其适应症(针对精准干预急性冠状动脉综合征后状态合并颈动脉斑块残余风险研究),加速完成药品再评价、逐步提升药品价值并申请进入基药目录。三是重启参芎葡萄糖注射液三期临床相关研究,扩大产品适应症重新申报国家医保药品目录,同时加速金鸡丸、乐脉丸等中药产品的多元发展;对于通过一致性评价的品种,深度挖掘销售价值,增加产品的中标概率。四是完善升级生产线,拓展玻璃酸钠制剂产品(包括玻璃酸钠美容填充项目和水光针项目)产能。淘汰生产端陈旧设备,改善生产自动化水平,通过委托加工等多种方式,高效解决工厂产能不足现状,保证旺季销售供应。五是围绕榄香烯开展系统性研究,开展质量标准提升研究,并推进提取后副产物的资源化利用;在此基础上,开发榄香烯系列衍生物及相关制剂,拓展其在抗肿瘤及其他疾病治疗领域的应用范围;同时,通过优化提取工艺,降低生产成本,提高资源利用效率,最终构建从原料到制剂的全产业链价值提升路径。(2)拓展销售渠道,提升营销能力一是调整营销模式。将由各子公司独立经营销售转变为母公司“统一规划,统一销售,统一管理,统一团队”的四统一销售模式,促进存量产品发展。按照产品治疗领域和销售渠道的属性进行划分形成条线管理,通过组织架构的调整,使销售队伍在执行力、专业度上快速提升。打造心脑血管、抗肿瘤等领域专业化的营销团队。二是继续拓展基层及民营医疗机构等三终端,提升产品销售和市场覆盖比例。将参芎葡萄糖注射液、乐脉丸、金鸡丸、通迪胶囊、妇平胶囊、儿童回春颗粒、注射用克林霉素磷酸酯、阿糖腺苷、奥美拉唑等列为发力品种,参芎重点拓展代理商数量,填补空白市场,加强代理商管理,要求更多市场覆盖和纯销提升;梳理、整合其他产品的三终端渠道和代理商现状,积极开发新的市场机会,做好代理商选择及优化,尽快完成三终端销售的布局和价值体现;普药部分做好现有市场覆盖的维护及提升,利用积极有效的市场调研和价格杠杆,努力培育5-10个战略合作伙伴,加快提升普药的市场份额和销售金额。三是规划集采药品销售方案。成立集采销售部,规划仿制药布局,积极探索和参加国家集采及省际联盟集采,力争更多产品进入国家集采通道和中选更多省份的集采招标。其中,重点推进海南锦瑞公司仿制药一致性评价的过审,以及玻璃酸钠注射液重点开发上量。另外,市场准入团队积极学习和解读集采政策,结合公司产品结构,响应国家集采号召,积极参与、力争中选,统一配送、跟进销售。后续公司将更加关注多家中选省份的销售策略制定和备选省份的销售拓展计划。四是建立和壮大零售队伍,打造公司销售业务新增长点。零售连锁市场逐年增长,公司将建立专门的零售队伍,通过外部引进和内部培养的方式快速建立零售专业销售团队,产品聚焦心脑宁胶囊、镇痛活络酊、冰栀伤痛气雾剂、乐脉丸、金鸡丸等产品;全国top10连锁力争直接合作或者贴牌销售,其他连锁选择专业的代理商合作,快速上架,专业推广,迅速提升销售,为后续销售增长带来强劲动力。(3)创新创研新产品,实现产业升级借鉴产业投资人的研发创新优势,公司未来医药研发将以“建立完整、高效、国内领先的研究开发系统,制定并持续更新科学、稳健的产品线规划”为目标,建立健全与国际接轨的研发质量管理体系,加强建设研发项目管理体系。一是加快推进化药新药项目落地,NMPA1类新药交联玻璃酸钠注射液根据III期临床CED沟通会的要求进行补充研究,推动完成III期临床试验样品的生产与工艺验证,并启动III期临床研究。二是推动普通注射剂项目完成一致性评价及工艺验证,推进玻璃酸钠注射液、注射用氯诺昔康开展一致性评价工作。三是持续推进抗肿瘤产品研发,以多种方式解决抗肿瘤药物伊立替康注射液、注射用吉西他滨、注射用异环磷酰胺等产品的产能不足及产销问题。四是公司在产品立项上将继续保持谨慎态度,依据自身研发优势及核心竞争力,从多维度进行产品评估,结合未来产品市场竞争态势的预测,以及未来销售量等市场数据,确定研发方向,推进临床研究以及里程碑的逐步实现,以实现产品上市后的收益。2.培育生物医药第二增长曲线(1)常德市政策支持生物医药产业,已形成产业集聚生物制药产业在常德市拥有深厚的产业基础。常德在生物制造领域底蕴深厚,拥有亚洲最大的甾体原料药和中间体生产出口基地、全国最大的酶制剂生产出口基地、全省第一方阵的生物医药企业、全省规模领先的食品加工产业。目前,常德市已建成国家级合成生物制造产业创新平台1个、省级创新平台5个、省级科技成果转化中试基地1个。常德经开区和津市市的合成生物中试转化基地即将投入使用,安乡县合成生物产业应用研发中心已经建成。常德市和湖南文理学院共建合成生物学公共技术平台,一体布局生物合成、分子检测、生物提取、生物发酵和生物医学等高能级实验室,争创合成生物省级、国家级重点实验室。生物制药产业在常德市拥有完备的园区配套支撑。按照“一中心三基地”产业布局,津市市、常德经开区产业承载优势,安乡县原料资源优势,全力打造全省引领、全国有名的常德“生物制造谷”,推动合成生物制造产业高端化、绿色化、融合化、协同化发展。(2)石药控股及景峰医药在生物制药领域拥有深厚积累石药控股在生物制药领域拥有深厚积累,研发实力雄厚。石药控股及其下属企业依托纳米制剂药物、细胞治疗、长效注射剂等八大技术平台,聚焦肿瘤、精神神经、心血管等领域,并在中药和合成生物学等领域有所布局。景峰医药研发中心已分别打造生物药开发技术平台、大分子交联技术平台、纳米分散系DDS平台、MUPS(MultipleUnitPelletSystem)OSD平台、预灌封制剂产业化平台等研发平台,致力于建立高技术壁垒的生物药品研发与生产平台,为拓展生物制药提供研发基础保障。(3)依托区位优势及企业积累,培育第二增长曲线重整完成后,景峰医药将依托常德市对生物医药行业的政策支持、资金保障、产业集聚,并凭借石药控股和公司在生物医药行业的积累与研发实力,着力培育生物医药作为第二增长曲线,拓展合成生物产品及关联赛道,逐步成长为双轮驱动的优质公司。具体举措包括但不限于:一是聚焦合成生物学与创新药械,推动具体项目落地。公司将重点引入与现有业务具有战略协同效应的优质合成生物学项目,在常德逐步构建从研发、中试到产业化的合成生物制造产业链,为业绩贡献新增量。二是联合设立专项产业基金,构建项目孵化与引入平台。该基金将借助石药控股的产业资源和基金管理经验,主要投向生物医药及相关联的高成长性领域,在追求财务回报的同时,积极引导被投企业将研发成果和生产基地落户常德,形成“基金投资—项目孵化—产业落地”的良性循环。三是整合内外部生物医药资源,打造“研发-生产-销售”一体化生态。推动石药控股与景峰医药的生物医药协同研发项目在常德优先转化及生产落地,并借助石药控股整合两湖两广销售资源的优势,在常德设立销售总部统一管理,从而全面强化常德总部的职能,打造“研发-生产-销售”的生物医药全链路,实现第二曲线扎实增长。(二)优化组织架构,提升管理水平,降本增效借鉴产业投资人丰富的企业管理经验,景峰医药将进一步优化组织运营管理架构,强化企业管理,加强景峰医药及子公司在战略、财务、人才、业务等方面的规范运作管理,不断提升内控管理体系运行质量,通过建立和运行常态化的管理提升机制,降低管理成本及费用,推进和保障公司可持续发展。具体如下:1.优化组织架构,强化总部管控一是公司将继续严格执行《公司法》、《证券法》及相关规定,严格履行股东会及董事会的监督指导作用,充分发挥独立董事的风险把控作用。二是重构组织架构,通过销售集中、采购集中、财务集中三集中模式,实现扁平化管理。强化母公司管控力度,梳理人员配置,定岗定编,先推动实现财务专业和营销专业纳入母公司的集中体系化管理,再逐步拓展至生产、研发、采购等专业管理;发挥母公司计划、调度、考核等管理职能,合理科学调配母公司内部资源,实现公司整体利益最大化。2.完善管理制度,提升管理水平一是优化业务流程。加强制度和流程的执行管理,提高组织运营效率;完善预算管理体系与制度,实现管理的横向到边、纵向到底的全覆盖;完善健全投资管理制度,以理性的、科学的投资决策和管理制度引领企业稳健增长;建立高效管理机制,搭建周报、半月会和经营分析会等方式,确保重点工作、核心指标的达成。二是强化财务风险管控。包括重点加强资金审批控制、完善内部会计稽核制度、严格执行预算和收支管理等;公司将合理运用法规及制度筹措资金,并提高现金周转率和回款速度、降低回款周期;将安排工作组加大对应收账款的清欠回收力度,进一步加强应收账款管理,加速资金回笼以缓解资金压力。三是制定人才发展战略。公司将全面梳理人员配置,完善薪酬考核体系,在培养人才的同时留住人才。公司将围绕经营目标,落地预算管理,加强指标与绩效考核的管理,分配机制重点向业务线、市场线倾斜,激发员工的工作热情,强化以结果为导向的考核机制,实行内部竞争上岗制和轮岗制,激发内部活力。3.加强成本管控,盘活低效资产一是落地成本领先战略。公司将持续抓好生产组织和原料采购,降低运营成本并认真做好开源节流、增收节支工作。二是盘活低效资产,提升资产运营效率,降低资产运营压力。公司将继续对低效业务、低效资产与闲置资产进行重整与盘活,充分提高资产利用率、提高存货周转率,不断强化主营业务,提升经营现金流。七、重整计划(草案)的表决和批准(一)重整计划(草案)的表决1.分组表决景峰医药债权人会议设普通债权组对重整计划(草案)进行正式表决。根据《企业破产法》第八十二条和第八十三条、《最高人民法院关于适用<中华人民共和国企业破产法>若干问题的规定(三)》第十一条,职工债权、税款债权组因权益未受调整而不参与表决,依法不设表决组。因重整计划(草案)涉及出资人权益调整事项,设出资人组对出资人权益调整事项进行表决。2.表决机制(1)债权人组的表决机制根据《企业破产法》第八十四条第二款的规定,出席会议的同一表决组的债权人过半数同意重整计划(草案),并且其所代表的债权额占该组债权总额的三分之二以上的,即为该组通过重整计划(草案)。(2)出资人组的表决机制根据最高人民法院《关于切实审理好上市公司破产重整案件工作座谈会纪要》第18条第三款的规定,经参与表决的出资人所持表决权三分之二以上通过的,即为该组通过重整计划(草案)。(二)重整计划(草案)的批准普通债权组表决通过重整计划(草案),且出资人组表决通过出资人权益调整事项时,重整计划(草案)即为通过。债务人将依法向常德中院提出批准重整计划的申请。部分表决组未通过重整计划(草案)的,债务人或者管理人可以同未通过重整计划(草案)的表决组协商。该表决组可以在协商后再表决一次。双方协商的结果不得损害其他表决组的利益。未通过重整计划(草案)的表决组拒绝再次表决或者再次表决仍未通过重整计划(草案),但重整计划(草案)符合《企业破产法》第八十七条规定的,债务人或者管理人可以申请人民法院批准重整计划(草案)。(三)重整计划批准生效重整计划(草案)在依据《企业破产法》第八十四条至第八十七条的相关规定,由各表决组表决通过并经常德中院裁定批准后生效,或部分表决组表决虽未通过但经常德中院裁定批准后生效。(四)未获批准的后果重整计划(草案)未获得债权人会议通过并且未依照《企业破产法》第八十七条的规定获得常德中院批准;或已经表决通过的重整计划未获得法院裁定批准的,管理人将依法申请常德中院裁定终止重整程序,并宣告景峰医药破产。八、重整计划的执行本重整计划由景峰医药负责执行。(一)执行期限本重整计划的执行期限为常德中院裁定批准重整计划(草案)之日起6个月。在此期间,景峰医药应当严格依照重整计划的规定清偿债务,并随时支付破产费用。(二)执行期限的延长与提前如非景峰医药自身原因,致使本重整计划无法在上述期限内执行完毕,景峰医药应于执行期限届满前,向常德中院提交延长重整计划执行期限的申请,并根据常德中院批准的执行期限继续执行。(三)执行完毕的标准自下列条件全部满足之日起,重整计划视为执行完毕:1.管理人收到重整投资人受让转增股票的全部现金对价款;2.用于引入重整投资人的转增股票已登记至管理人开立的证券账户或已经登记至相关投资人或其指定主体名下的证券账户;3.破产费用已经支付完毕或已经提存至管理人账户;各类债权已经按照债权调整和清偿方案获得清偿,或用于清偿的现金已提存至管理人账户。(四)司法协助执行事项重整计划执行过程中,涉及需要有关单位协助执行的,景峰医药及/或管理人可向常德中院提出申请,请求常德中院向有关单位出具要求其协助执行的司法文书。(五)执行完毕的效力根据重整计划予以豁免的债权,景峰医药不再承担清偿责任。(六)重整计划的变更本重整计划执行过程中,因遇国家政策调整、法律修改变化等特殊情况或发生意外事件或不可抗力致使本重整计划无法继续执行的,可由管理人召集债权人会议,或由债务人向人民法院申请召开债权人会议,就是否同意变更重整计划进行表决。债权人会议决议同意变更重整计划的,债务人应自决议通过之日起十日内提请常德中院批准。常德中院裁定批准变更重整计划的,景峰医药或者管理人应当在六个月内提出新的重整计划。变更后的重整计划应提交给因重整计划变更而遭受不利影响的债权人组和出资人组进行表决。变更后的重整计划在权益受到调整或影响的债权人组及/或出资人组表决通过并获得法院裁定批准后,由景峰医药执行变更后的重整计划,管理人予以监督。虽有上述约定,若重整计划(草案)在经债权人会议表决通过后常德中院裁定批准前或重整计划在执行过程中,需变更重整财务投资人、上调受让股票价格,或调整清偿方案的,在调整后的方案不减损债权人利益或对债权人更有利的情况下,债务人可直接予以变更,无需通过债权人会议进行表决。调整后的重整计划(草案)或重整计划由管理人向常德中院报备。重整计划确系无法通过修改等方式实现继续执行的,常德中院可应管理人或利害关系人的请求裁定终止重整计划的执行,并宣告景峰医药破产。(七)有关重整计划执行的重大不确定事项重整计划执行过程中,可能会出现投资人违约、无法转增股票等风险。九、重整计划执行的监督管理人负责监督重整计划的执行。(一)监督期限本重整计划执行的监督期限与重整计划的执行期限相同,自常德中院裁定批准重整计划之日起计算。(二)监督期限的延长与提前如根据重整计划执行的实际情况,需要延长管理人监督重整计划的执行期限的,则管理人将向常德中院提交延长重整计划执行监督期限的申请,并根据常德中院批准的期限继续履行监督职责。重整计划执行期限提前到期的,执行监督期限相应提前到期。(三)监督期限的终止监督期限届满或公司执行完毕本重整计划时,管理人将向常德中院提交监督报告,自监督报告提交之日起,管理人的监督职责终止。十、其他说明情况(一)重整计划的生效依据《企业破产法》第八十四条至第八十七条之相关规定,重整计划(草案)在景峰医药债权人会议、出资人组会议表决通过并经常德中院裁定批准,或债权人会议、出资人组会议表决虽未通过但经申请常德中院裁定批准后生效。重整计划生效后,对债务人、全体债权人、重整投资人和出资人具有法律约束力。本重整计划对相关方权利义务的规定效力及于该项权利义务的承继方或受让方。(二)偿债资源的提存及预留、处理1.偿债资源的提存债权已经确定的债权人未按照重整计划的规定领受偿债资源的,根据重整计划应向其分配的资金将提存至管理人指定的银行账户。提存后视为景峰医药已经根据本重整计划履行了清偿义务,不再就所涉债权承担清偿责任。前述提存的偿债资金自债务人发布重整计划执行完毕公告之日起满三年,债权人仍不领取的,视为放弃领受偿债资金的权利,已提存的偿债资金将归还景峰医药,作为景峰医药的流动资金。2.偿债资源的预留对于因诉讼、仲裁未决、债权人异议等原因导致管理人暂时无法做出审查结论的债权,将按照重整计划应向其分配的资金预留偿债资源,以最终确定的债权金额为准,按照本重整计划规定的同类债权清偿方案受偿。未申报债权在本重整计划执行期间不得行使权利;在本重整计划执行完毕后,该部分未申报债权经申报并经债务人审查确定后,可按本重整计划的规定要求景峰医药按同类债权受偿方式予以清偿。按照本重整计划已预留的偿债资金在清偿该等债权后仍有剩余的,剩余的偿债资金将用于补充公司流动资金。如最终确认债权对应应受领的偿债资源超过预留的偿债资源,超过部分仍由景峰医药承担清偿责任,预留现金不足以清偿债权的部分,以届时景峰医药自有资金予以清偿。上述提存/预留的偿债现金,在提存/预留期间均不计息。(三)偿债资源的分配每家债权人以现金方式受偿的债权部分,偿债资金原则上以银行转账方式向债权人进行分配。债权人应自常德中院裁定批准重整计划之日起10个工作日内,按照本重整计划附件要求的书面格式(见附件一、附件二)提供接受偿债资金的银行账户信息。逾期不提供相关信息、因债权人自身和/或其关联方的原因,导致偿债资金不能到账,或账户被冻结、扣划,产生的法律后果和风险由相关债权人自行承担,且不影响重整计划的执行完毕。景峰医药按照本重整计划的规定向债权人指定的银行账户完成偿债资金的分配后,视为景峰医药已经根据本重整计划履行了清偿义务,不再就所涉债权承担清偿责任。(四)财产保全措施的解除及信用等级的修复1.财产保全措施的解除根据《企业破产法》第十九条的规定,人民法院受理破产申请后,有关债务人财产的保全措施应当解除。尚未解除对景峰医药财产保全措施的债权人,应当在本重整计划获得法院裁定批准后30日内协助办理完毕解除财产保全措施的手续。如债权人未能在前述规定期限内向有关法院提交解除措施申请的,管理人及/或景峰医药有权申请常德中院依照本重整计划的规定予以强制解除。景峰医药有权根据债权人配合解除财产保全措施的情况向该债权人支付偿债资金,因相关债权人不配合导致无法按期受领偿债资金的,不视为重整计划未能执行完毕。2.信用等级的修复申请强制执行并将景峰医药纳入失信被执行人名单的各债权人,应当在本重整计划获得法院裁定批准之日起30日内向相关法院申请删除景峰医药的失信信息,并解除对债务人法定代表人、主要负责人及其他相关人员的限制消费令及其他信用惩戒措施。在本重整计划经常德中院裁定批准后,景峰医药可向相关债权银行提出信用记录修复申请,相关债权银行应及时调整企业信贷分类,并上报中国人民银行征信系统调整企业征信记录。(五)债权人对其他还款义务人的权利根据《企业破产法》第九十二条第三款的规定,债权人对债务人的保证人和其他连带债务人所享有的权利,不受重整计划的影响;债权人依本重整计划未被清偿的债权部分,债权人有权依照相关担保合同约定要求担保人承担相应清偿责任。如保证人或其他连带债务人清偿上述债权的,其是否可向景峰医药追偿,将依照《民法典》担保制度、《最高人民法院关于适用<中华人民共和国企业破产法>若干问题的规定(三)》等法律规定处理,若依法可追偿的,其将替代原债权人对景峰医药预留偿债资源享有受偿权利。(六)对实控人原有股权调整的方案根据最高人民法院、中国证监会《关于切实审理好上市公司破产重整案件工作座谈会纪要》第16条有关规定,“控股股东、实际控制人及其关联方因违法违规行为对上市公司造成损害的,制定重整计划草案时应当根据其过错程度对控股股东及实际控制人支配的原有股权作相应调整。”参考上述规定,本重整计划对景峰医药控股股东、实际控制人权益的调整方案为:叶湘武无偿、无条件且不可撤销的让渡本人所持300万股景峰医药股票,所让渡的股票由公司自行处置,变现所得归公司所有。(七)破产费用的支付及共益债务的清偿1.破产费用景峰医药破产费用包括重整案件受理费、管理人报酬、聘请专业机构的费用、转增股票登记税费、股票过户税费及管理人执行职务的费用等,暂合计约2,200万元。其中,重整案件受理费依据《诉讼费用交纳办法》支付;债务人转增股票登记税费及股票过户税费、管理人执行职务的费用根据重整计划执行实际情况随时支付;管理人报酬依据《最高人民法院关于审理企业破产案件确定管理人报酬的规定》规定的比例上限计算,并以现金支付给管理人;管理人聘请中介机构的费用依据相关合同的约定支付。2.共益债务景峰医药重整期间的共益债务,包括继续营业以及由此产生的其他债务,约合计900万元(具体以实际支付金额为准),由景峰医药按照《企业破产法》相关规定随时清偿。(八)转让债权的清偿债权人依法对外转让债权的,受让人按照原债权人根据本重整计划就该笔债权可以获得的偿债资源受偿;债权人将一笔债权向两个以上的受让人转让债权的,债权按照一笔计算,偿债资源按照受让人各自债权比例分配。债权转让方应当将债权转让相关信息在法院出具协助执行通知书前通知管理人,同时债权受让方应当向管理人提供受领信息,管理人将按照债权受让方的指示进行偿债资源的分配,相关债权人逾期不提供相关信息,产生的法律后果和风险由相关债权人自行承担。如转让通知到达管理人前已经分配的,由转让双方自行协商或依法解决。(九)重整计划的解释在本重整计划执行过程中,若债权人或利益相关方对重整计划部分内容存在不同理解,且该理解将导致利益相关方的权益受到影响的,则债权人或利益相关方可以向景峰医药或管理人申请对重整计划相关内容进行解释。管理人在收到该申请之后,应基于公平、公正的原则对相关内容进行解释。(以下无正文)附件一:关于领受偿债资金的账户信息告知书(债权金额200万元以下(含200万元)的普通债权人)湖南景峰医药股份有限公司:按照湖南景峰医药股份有限公司重整计划的规定,请将本公司/本人受领的应分配款项转入如下账户:开户银行:账户名称:账号:债权人名称(签章/签字):年月日附件二:关于领受偿债资金的账户信息告知书(债权金额200万元以上的普通债权人)湖南景峰医药股份有限公司:按照湖南景峰医药股份有限公司重整计划的规定,本公司/本人债权清偿方式选择见以下信息,请将本公司/本人受领的应分配款项转入如下账户:1.关于普通债权中超过200万元的部分之清偿方式选择?现金清偿?留债2.银行账户信息开户银行:账户名称:账号:债权人名称(签章/签字):年月日公告原文
100 项与 阿糖腺苷 相关的药物交易
登录后查看更多信息
研发状态
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| 单纯疱疹病毒性脑炎 | 日本 | 1984-02-15 | |
| 带状疱疹 | 美国 | 1976-11-26 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
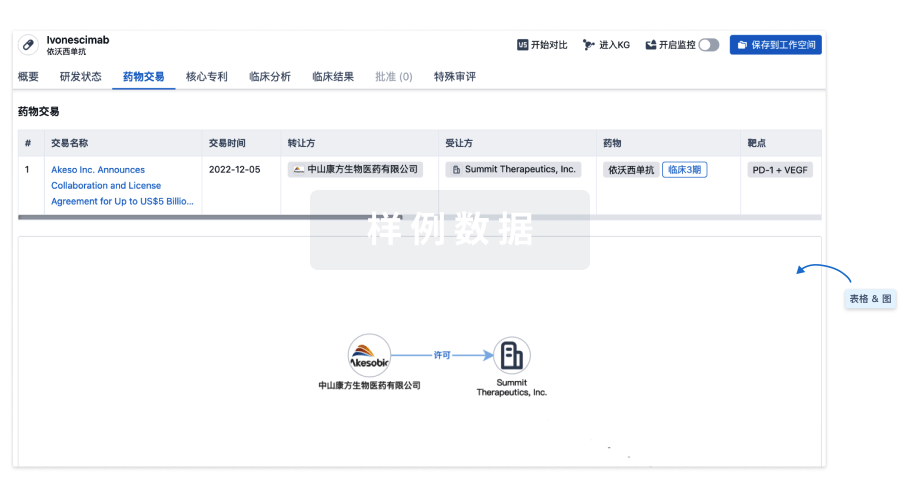
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
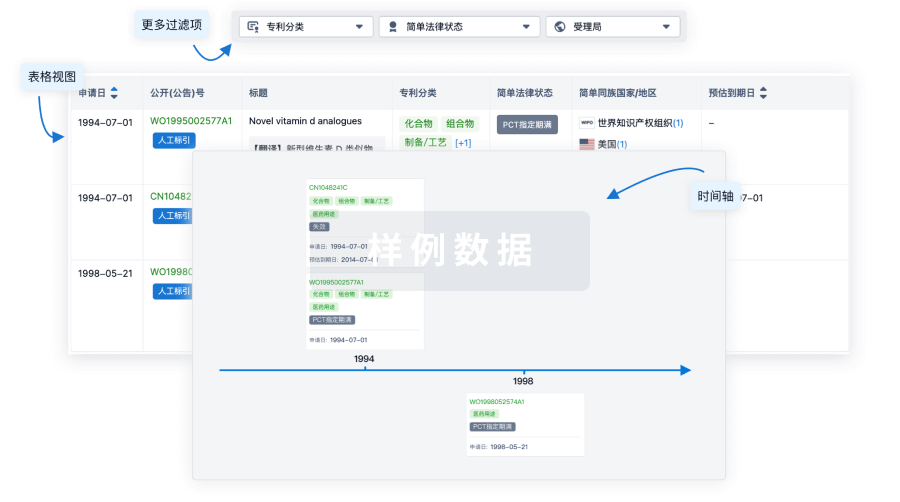
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
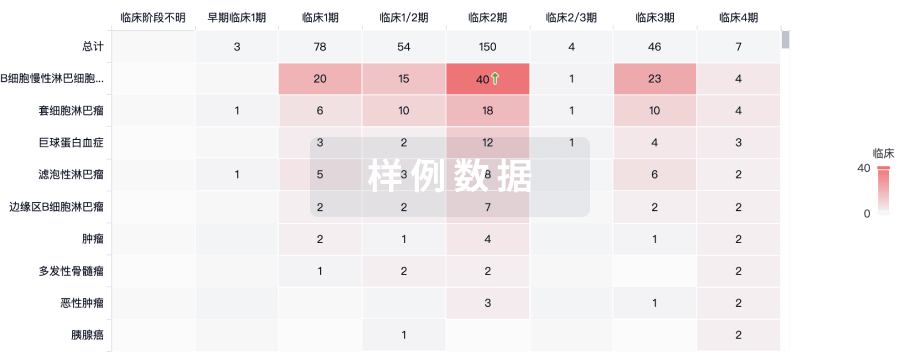
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
批准
利用最新的监管批准信息加速您的研究。
登录
或
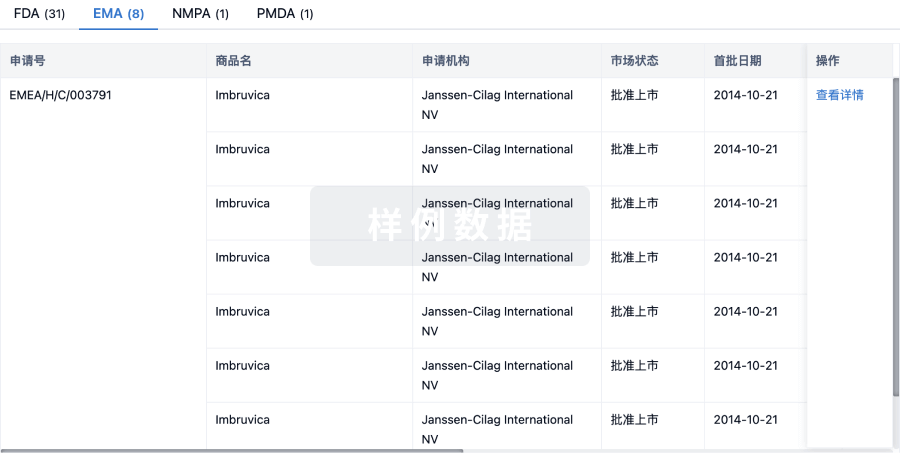
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用

