预约演示
更新于:2026-03-07
Galidesivir
更新于:2026-03-07
概要
基本信息
在研机构- |
权益机构- |
最高研发阶段终止临床2期 |
首次获批日期- |
最高研发阶段(中国)- |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C11H15N5O3 |
InChIKeyAMFDITJFBUXZQN-KUBHLMPHSA-N |
CAS号249503-25-1 |
关联
3
项与 Galidesivir 相关的临床试验NCT03891420
A Phase 1b Double-blind, Placebo-controlled, Dose-ranging Study to Evaluate the Safety, Pharmacokinetics, and Anti-viral Effects of Galidesivir Administered Via Intravenous Infusion to Subjects With Yellow Fever or COVID-19
This is a placebo-controlled, randomized, double-blind study to evaluate the pharmacokinetics, safety and antiviral activity of galidesivir in subjects with yellow fever (YF) or COVID-19.
开始日期2020-04-09 |
申办/合作机构 |
NCT03800173
A Phase 1 Double-blind, Placebo Controlled, Dose Ranging Study to Evaluate the Safety, Tolerability, and Pharmacokinetics of Galidesivir (BCX4430) Administered as Single Doses Via Intravenous Infusion in Healthy Subjects
This is a placebo-controlled, randomized, double-blind study to evaluate the pharmacokinetics of galidesivir following administration of single doses by IV infusion
开始日期2018-12-10 |
申办/合作机构 |
NCT02319772
A Phase 1 Double-blind, Placebo-controlled, Dose-ranging Study to Evaluate the Safety, Tolerability, and Pharmacokinetics of BCX4430 Administered Via Intramuscular Injection (IM) in Healthy Subjects
This is a 2-part, first-in-human dose-ranging study to evaluate the safety, tolerability and pharmacokinetics of escalating doses of BCX4430 administered via intramuscular (IM) injection in healthy subjects. In part 1, subjects will receive a single dose of BCX4430; in part 2 subjects will receive BCX4430 for 7 days.
开始日期2014-12-01 |
申办/合作机构 |
100 项与 Galidesivir 相关的临床结果
登录后查看更多信息
100 项与 Galidesivir 相关的转化医学
登录后查看更多信息
100 项与 Galidesivir 相关的专利(医药)
登录后查看更多信息
132
项与 Galidesivir 相关的文献(医药)2026-02-01·ANTIVIRAL RESEARCH
Substitutions M478K and A482G in the polymerase of yellow fever virus confer resistance to sofosbuvir and uprifosbuvir in vitro
Article
作者: Mikkelsen, Lotte ; Rosas, Ana Lucia Rosales ; Binderup, Alekxander ; Fahnøe, Ulrik ; Ramirez, Santseharay ; Delang, Leen ; Bukh, Jens ; Soto, Alina ; Elorz, Itsaso Ortigosa ; Ryberg, Line Abildgaard
The continuous re-emergence of yellow fever virus (YFV) poses a serious public health threat, with no approved specific antiviral drugs for treatment. Sofosbuvir and uprifosbuvir, developed for treatment of chronic hepatitis C, are nucleotide analogs with anti-YFV activity in vitro, and sofosbuvir has been used off-label in YFV-infected individuals. To study the barrier to resistance of these drugs, escape experiments were performed with the YFV 17D vaccine strain in human hepatoma Huh7.5 cells. Compared to the original virus, sofosbuvir and uprifosbuvir escape variants exhibited 8.6- to 10.7-fold increased 50% effective concentration (EC50) values. Escape viruses developed putative resistance-associated-substitutions (RAS) M478K and A482G in the NS5 RNA-dependent RNA polymerase domain. Reverse-engineered YFV 17D mutants demonstrated that A482G was the primary determinant of drug resistance, resulting in 5.3- to 5.7-fold increased EC50; however, the substitutions acted synergistically when combined, leading to 12.3- and 11.1-fold increased EC50. A482G conferred slight cross-resistance to bemnifosbuvir, while no cross-resistance was observed for remdesivir, galidesivir, or NITD008. Escape and mutant viruses had slight delays in growth kinetics in Huh7.5 cells and were viable in C6/36 and Aag2-AF5 mosquito cells. Structural analysis of a predicted AlphaFold3 YFV NS5-RNA structure suggests that M478K and A482G induce conformational changes in RdRp motif F, which may reduce sofosbuvir and uprifosbuvir affinities. Sofosbuvir RAS described for other orthoflaviviruses did not substantially affect drug susceptibility in reverse-engineered YFV 17D. This study demonstrates that YFV can develop resistance to sofosbuvir and uprifosbuvir with only two NS5 substitutions that have slight impact on viral fitness.
2025-12-31·JOURNAL OF ENZYME INHIBITION AND MEDICINAL CHEMISTRY
Identification of novel inhibitors of dengue viral NS5 RNA-dependent RNA polymerase through molecular docking, biological activity evaluation and molecular dynamics simulations
Article
作者: Yan, Hong ; Liu, Xiaojing ; Li, Wei ; Zhang, Susu ; Zong, Keli ; Wang, Cong ; Cao, Ruiyuan ; Li, Xingzhou ; Ruan, Jiajun ; Wei, Chaochun
The DENV-NS5 RNA-dependent RNA polymerase (RdRp) is essential for viral replication, and one of the targets of anti-virus. In this study, the Uni-VSW module was used to virtual screen 1.6 million compounds in the ChemDiv and TargetMol (USA) database, 27 candidates were obtained. Thereby 23 candidates were selected based on their binding free energies by 50 ns MD simulations. The biological activity of the candidates and the reference compounds (BCX4430 and Compound 27) were evaluated on their IC50 values against DENV-NGC, CC50 values, and selectivity index. Among these, the IC50 values of D1 and D8 were 13.06 ± 1.17 μM and 14.79 ± 7.76 μM, respectively, which were better than that of Compound 27 (IC50 =19.67 ± 1.12 μM). The comprehensive MD simulations were performed on the candidates to assess the stability behaviour and binding mechanisms. The density functional theory (DFT) analysis was also conducted to explore the structural and electronic properties.
2025-11-02·JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS
Kaempferol as a novel inhibitor of SARS-CoV-2 RNA-dependent RNA polymerase
Article
作者: Benedetti, Francesca ; Scapagnini, Giovanni ; Davinelli, Sergio ; Medoro, Alessandro ; Jafar, Tassadaq Hussain ; Ali, Sawan ; Zella, Davide ; Passarella, Daniela ; Ismail, Saba ; Trung, Truong Tan ; Intrieri, Mariano
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has quickly become a global health pandemic. Among the viral proteins, RNA-dependent RNA polymerase (RdRp) is responsible for viral genome replication and has emerged as a promising target against SARS-CoV-2 infection. Dietary bioactive compounds represent an important source of evolutionarily optimized molecules with antiviral properties against SARS-CoV-2 RdRp. We investigated the inhibitory potential effects of different phytochemicals against SARS-CoV-2 RdRp, including andrographolide, kaempferol, resveratrol, and silibinin. Unlike the other investigated compounds, kaempferol exhibited a significant dose-dependent in vitro inhibition of SARS-CoV-2 RdRp activity. To assess the binding interactions and stability of the SARS-CoV-2 RdRp-kaempferol complex, we performed in silico techniques, including molecular docking, quantum chemical calculation, and molecular dynamics simulations. We found strong binding affinities and stability between kaempferol and SARS-CoV-2 RdRp variants (Wuhan and Omicron). These findings provide valuable insights into the antiviral properties of kaempferol as a stable inhibitor of SARS-CoV-2 RdRp.
58
项与 Galidesivir 相关的新闻(医药)2025-12-30
点击上方“中国药物警戒”关注我们
“
拉沙热抗病毒药物临床应用研究进展
Research Progress in Clinical Applications of Antiviral Drugs for Lassa Fever
”
专家介绍
陈志海,主任医师,教授,博士生导师,国家级人才项目获得者。首都医科大学附属北京 地坛医院感染性疾病中心主任,国家疾控局传染病标准专业委员会委员,中华医学会感染病学分会常委,中华医学会热带病与寄生虫学分会常委,中华医学会北京分会感染专业委员会副主任委员,北京预防医学会感染病学专委会副主任委员,中西医协同感染病旗舰科室主任。致力于各种急性传染病临床诊治研究 30 余年,作为国家卫生健康委员会、北京市卫健委专家参加全国各种重大传染病疫情处置。《中国药物警戒》期刊编委。
引用本文:王冉冉, 马瑞泽, 陈志海. 拉沙热抗病毒药物临床应用研究进展[J]. 中国药物警戒, 2025, 22(12): 1321-1327.
Reference:WANG Ranran, MA Ruize, CHEN Zhihai. Research Progress in Clinical Applications of Antiviral Drugs for Lassa Fever[J]. Chinese Journal of Pharmacovigilance(中国药物警戒), 2025, 22(12): 1321-1327.
摘要:目的 综述拉沙热(Lassa Fever,LF)抗病毒药物研究进展,为突破当前治疗瓶颈提供参考。方法 通过检索PubMed、Web of Science、中国知网等数据库,归纳法维拉韦、利巴韦林及免疫治疗的临床应用效果,并总结具有潜力 的小分子药物的作用机制。结果 法维拉韦、利巴韦林和免疫治疗均表现出一定的疗效,利巴韦林联合法维拉韦或地塞 米松在临床中已有救治成功的案例。国内外针对利巴韦林的使用剂量及疗程存在争议,且现有证据不足以支持常规使 用利巴韦林治疗 LF。法维拉韦的心脏安全性也仍需进一步评估。多种小分子抑制剂也显示出广阔的临床应用前景。结论 目前全球范围内尚无获批的特异性抗拉沙病毒(Lassa Virus,LASV)药物,但针对病毒生命周期不同环节的多 种候选药物研究已取得显著进展,联合用药及多靶点治疗的策略将成为未来的重要研究方向。
关键词:拉沙热;拉沙病毒;抗病毒药物;免疫治疗;小分子抑制剂;研究进展
基金项目:国家重点研发计划(2022YFF1203201)。
正文
拉沙热(Lassa Fever,LF)是由拉沙病毒(Lassa Virus,LASV)引起的一种急性病毒性出血性疾病, 主要流行于西非国家 [1]。据估计,西非地区每年约 897 700 人感染 LASV,超 18 000 人死亡,总体病死率约 1%,孕妇和胎儿的致死率分别为 30% 和 80%[2]。 近年来,随着全球人口流动性不断增加,美国、德国、 英国及日本等多个国家相继报告 LF 输入病例,尤其 是 2024 年 8月我国四川省确诊首例输入性 LF 病例, 这为本土防控体系敲响警钟 [3]。目前全球已有多个候选疫苗进入临床试验阶段,但暂无获批上市的特效疫苗,因此抗病毒药物的研发成为控制拉沙热传播与重症化的关键手段。然而,当前尚无获批的特异性抗病毒药物,临床上仍以对症支持治疗为主 [4]。尽 管既往使用利巴韦林早期干预和恢复期血浆免疫疗法有个案成功的报道,但均缺乏充分的疗效证据,且不同国家和地区关于使用利巴韦林治疗 LF 的诊疗方案存在差异。本文基于现有的临床研究和临床前 研究,系统综述 LF 的抗病毒治疗策略与前沿进展, 并聚焦具有转化潜力的候选药物,为突破当前治疗的局限性提供参考。
1
抗 LASV 药物研发基础
LASV 于 1969 年首次从尼日利亚病例样本中分离,属于布尼亚病毒纲(Bunyaviricetes)、沙粒病毒目 (Hareavirales)、哺乳沙粒病毒属(Mammarenavirus), 是一种具有不对称包膜的单链 RNA 病毒 [5]。该病毒 颗粒内包裹许多宿主核糖体,呈现独特的“沙粒样” 外观。病毒的基因组由 2 个 RNA 片段构成,即 S 片 段编码核蛋白(NP)与糖蛋白前体(GPC),L 片段则 编码 RNA 依赖性 RNA 聚合酶(L 蛋白)及锌结合基 质蛋白(Z 蛋白)。其中 GPC 在宿主细胞内经丝氨酸 蛋白酶 SKI-1/S1P 切割加工后,可组装成由受体结合 亚基(GP1)、膜融合亚基(GP2)及肉豆蔻酰化稳定 信号肽(SSP)共同构成的功能性糖蛋白复合体 [6]。 这些病毒学特征决定了其复制周期的关键环节,因 此深入理解 LASV 的生命周期及其分子机制,是研 发特异性抗病毒药物的重要基础。
1.1
病毒入侵宿主细胞
病毒入侵是感染的首要环节。LASV 通过 GPC 的 GP1 亚基特异性识别宿主细胞表面的 α-肌营养不良寡糖(α-DG)继而启动网格蛋白介导的内吞作用 [7] 。病毒被包裹进入核内体后,其内部的酸性环境触发 GPC 构象变化,使其与 α-DG 解离并以高亲和力结合胞内受体 LAMP1,进而驱动病毒包膜与核内体膜融合,同时将病毒核糖核蛋白复合体(vRNP)释放至胞质。在此过程中 GPC 发挥关键作用,因此开发干扰其与受体的结合、构象变化及膜融合过程的抑制剂是研发重点之一。
1.2
病毒复制与转录
LASV 基因组的复制与转录在胞质内完成,该 过程依赖其独特的“双义编码”策略,即病毒的 2 个 RNA 片段均包含正义和反义编码区,能够精细调控 不同病毒蛋白的表达时序。L 蛋白首先通过“帽状抢 夺”机制,以宿主 mRNA 的 5′端为引物,优先利用基 因组反义链区域转录翻译出 NP 和 L 蛋白,随后二者协同合成抗原基因组,进而转录表达 GPC 与 Z 蛋 白。值得注意的是,NP 兼具结构性与免疫调节功能, 不仅负责包裹病毒 RNA,而且其具有 3′–5′核酸外 切酶活性的 C 端结构域可降解病毒双链 RNA 中间 体,从而逃逸宿主天然免疫识别 [6]。因此,L 蛋白和 NP 构成阻断病毒复制与拮抗宿主防御的双重关键 靶点,相关药物的研发是当前重要的探索方向之一。
1.3
子代病毒组装与释放
子代病毒颗粒的组装与出芽是病毒生命周期的关键环节。Z 蛋白作为基质蛋白,通过其 N 端的肉豆 蔻酰化修饰介导病毒颗粒的组装,并从细胞膜出芽释 放子代病毒,该过程依赖于宿主细胞的 N-肉豆蔻酰转 移酶(NMT)[8]。因此,Z 蛋白及 NMT 构成了极具吸引 力的治疗靶点,旨在从源头上阻断子代病毒的产生。
综上所述,LASV 的生命周期包含多个核心环节, 针对病毒入侵、复制及组装等过程开发特异性抑制剂 已成为抗病毒药物研发的核心策略。本文在 LF 现有临 床治疗方案的基础上,进一步探讨针对不同靶点的候 选药物研究进展,为研发多样化治疗策略提供依据。
抗病毒药物研究成果
2
2.1
法维拉韦
法维拉韦是一种广谱的小分子抗病毒药物,主 要通过抑制 RdRp 发挥抗病毒作用。2015 年 OESTEREICH 等 [9]临床前研究显示,在嵌合体小鼠模型中 法维拉韦兼具体内和体外的抗 LASV 活性,且治疗 效果优于利巴韦林。研究进一步表明,相较于安慰 组小鼠和感染第 4 天起给予法维拉韦 75 mg·kg-1 或 150 mg·kg-1 治疗的小鼠均在感染第 9 天全部死亡 而言,给予高剂量(300 mg·kg-1·d-1)治疗的小鼠能 实现 100% 的存活率。同期 SAFRONETZ 等 [10]研究 发现,在哈特利豚鼠模型感染致死性 LASV 的第 7 天后启动法维拉韦治疗也能对其提供完全保护,而 在感染 2 d 后皮下注射利巴韦林仅能延缓死亡,停药 后症状即复发,未能改变其预后。此外,LINGAS 等 [11]通过非人灵长类动物(NHP)建立了首个复现 LASV动力学 及其治疗的数学模型,最终估算出法维拉韦在体内的 半数有效浓度(EC50)为 2.89μg·mL-1。模型还预测 若每日给予法维拉韦 2 次、每次剂量超过 1 200 mg 可能会有效减少具有传染性病毒的产生,这为未来 该药物在人类中的使用提供了重要参考。
目前法维拉韦治疗 LF 的临床疗效尚不明确, 缺乏相关研究数据的支持。根据法维拉韦用药说明,成人标准推荐每日口服 2 次,疗程共 5 d,具体给药 剂量为首日每次 1 600 mg,第 2~5 天每次 600 mg。 由于其在动物实验中表现出生殖毒性,因此禁用于 孕妇及备孕期女性 [12]。值得注意的是,法维拉韦的 心脏安全性需要密切关注。既往研究表明,合并新 型冠状病毒感染的部分 2 型糖尿病患者服药后出现 心电图 QT 间期延长,这提示该药可能引发不良心脏 事件 [13]。ERAMEH 等 [14]于 2021—2022 年在尼日利 亚开展的 1 项Ⅱ期随机对照试验(SAFARI)指出,在 接受法维拉韦单药治疗的 LF 轻症患者中出现 1 例 因 PR 间期显著延长而中止治疗的病例(该临床试验 具体给药方案为,第 1 天分 3 次口服,剂量依次为 2 400、2 400、1 200 mg,第 2~10 天为每日 2 次口服 每次 1 200 mg)。研究者认为此不良事件与病毒本 身作用关联较小,很可能为该药物特异性的影响。然 而,该研究的局限性在于仅纳入轻症患者且样本量 小,因此未来需要通过更大规模的临床试验来评估 法维拉韦在 LF 治疗中的心脏安全性,尤其是在 PR 间期延长方面的影响。
值得关注的是,研究表明法维拉韦与利巴韦林 具有协同抗病毒效应。FURUTA 等 [15]在免疫缺陷的 小鼠模型中发现,联合用药能够提高小鼠的存活率 并延长生存时间。此外,两者的协同效应在临床实践 中也得到了初步的验证,即 2017 年非洲 2 例通过接 触同一原发病例而感染 LF 的患者,分别在发病第 5 天和第 8 天接受法维拉韦和静脉注射利巴韦林联合 治疗后均完全康复 [16]。
2.2
利巴韦林
利巴韦林是一种广谱的抗 RNA 病毒药物,世界 卫生组织(WHO)将其列入病毒性出血热基本药物 清单。该药物的作用机制包括直接抑制 RdRp、次黄 嘌呤核苷酸脱氢酶(IMPDH)及抑制巨噬细胞促炎因 子释放,并下调宿主细胞干扰素刺激基因的表达等[17]。 静脉注射利巴韦林是目前 LF 的标准治疗方法,在临 床实践中不同地区的给药方案存在差异。
2.2.1 MCCORMICK 方案
20 世纪 80 年代 MCCORMICK 等 [18]提出利巴韦林可能是治疗 LF 的有效手段,该 研究中LF 高危患者(入院时血清AST≥150 IU·L-1 或 病毒载量≥103.6 TCID50·mL-1)利巴韦林静脉注射治 疗先予 2 g 负荷剂量,之后按 1 g/6 h 给药,持续 4 d, 再调整为 0.5 g/8 h 续用 6 d,总疗程 10 d,结果显示 发病后 6 d 内接受利巴韦林静脉注射治疗的病死率约为延迟治疗组(发病≥7 d 接受治疗)的 20%,这可 能与阻止高病毒载量导致的组织不可逆损伤有关。
2.2.2 尼日利亚 IRRUA方案
与每日多次给药的 MCCORMICK 方案不同,尼日利亚鲁阿专科教学医院(ISTH) 基于当地临床实践制定的“IRRUA 方案”每日1 次给 药。该方案起始剂量较高,后续剂量递减,具体为非 妊娠期成人确诊 LF 后的第 1 天给予 100 mg·kg-1 的 负荷剂量(最高不超过 7 g),第 2~5天每日单次给药 25 mg·kg-1,第 6~10 天每日1 次给药 12.5 mg·kg-1; 而妊娠期女性首日负荷剂量与标准成人相同,第 2~5 天按 16 mg·kg-1 每日 4 次给药,第 6 ~10 天按 8 mg·kg-1 每日3 次给药;儿童首剂按 33 mg·kg-1 给 药,之后调整为每 6 h 给药 16 mg·kg-1 持续 4 d,第 6 ~10 天每 8 h 给药 8 mg·kg-1。此方案已成功治愈 数百例确诊病例,2018 年被纳入尼日利亚国家的治 疗指南 [19]。除此之外,OKOGBENIN 等 [20]报告 2 例 来自尼日利亚的重症 LF 患者,通过采用 IRRUA方案 静脉注射利巴韦林联合地塞米松治疗后成功康复。
2.2.3 WHO 推荐方案(2016 年版)
WHO 在《病毒性出血热患者临床管理:一线卫生工作者口袋指南》 (2016 年版)[21]建议,首先给予 30 mg·kg-1 的负荷 剂量(单剂量最大不超过 2 g),随后以 15 mg·kg-1每 6 h 给药 1 次(q6h)(单剂量最大不超过 1 g)持 续 4 d,最后减量至 7.5 mg·kg-1 q8h 的剂量续用 6 d(单 剂量最大不超过 500 mg)。所有静脉给药均需用 150 mL 0.9% 生理盐水稀释后缓慢输注,同时明确 肌酐清除率 <50 mL·min-1 的肾功能不全患者需适当降低使用剂量。
2.2.4 中国拉沙热治疗方案(2008 年版)
根据中国拉沙热治疗方案(2008 年版),推荐确诊或疑似 LF 患者应尽早应用利巴韦林,首选静脉给药,成人首日负荷 剂量同 WHO 推荐方案(2016 年版)(30 mg·kg-1,最 大剂量<2 g),随后以16 mg·kg-1(最高剂量1 g)q6h,持续给药 4 d,最后以 8 mg·kg-1(最大剂量 500 mg) q8h 持续 6 d[22]。对比利巴韦林药品说明书,建议成 年患者使用剂量为每次 0.5 g(5 支),每日 2 次;小儿则按每日10 ~15 mg·kg-1,分 2 次给药,每次滴注 20 min 以上,疗程为 3~7 d[23]。
尽管利巴韦林被广泛用作LF的标准治疗药物, 但其确切的疗效仍缺乏足够的高质量临床证据支 持 [24]。2015 年 OESTEREICH 等 [9]在嵌合型 Ifnar-/-B6C57BL/6 小鼠模型中通过腹腔注射利巴韦林,结果显示其未能改善感染小鼠的生存结局,且发现启动 治疗的时间与死亡结局无关。EBERHARDT 等 [25]研究发现,该药可能增加部分病情较轻患者(如 AST<150 IU·L-1)的死亡风险,即接受利巴韦林治疗的 AST 水平正常患者的死亡率(9.1%)高于对照组患者 (4.1%)。SALAM 等 [17]基于体外研究数据表明,利巴 韦林抑制 LASV 复制的平均半数有效浓度(EC50)为 7 μg·mL-1,平均90% 有效浓度(EC90)为15 μg·mL-1。结 合健康人体药代动力学模型的预测分析表明,采用目 前现有给药方案(即MCCORMICK方案和IRRUA方案) 血药浓度超过上述平均EC50 的时间,不足给药间隔时长 的 20%,而超过平均 EC90 的时间则低于 10%。这提示当前临床应用的利巴韦林给药方案可能不足以在 血清中持续维持抑制 LASV 复制所需的药物浓度。
除此之外,溶血性贫血是利巴韦林公认的毒副 作用 , 约 20% 的 LF 患者经治疗后出现血红蛋白严 重下降。如我国 2024 年四川省首例输入性 LF 患者在 使用该药物治疗后因出现溶血性贫血而暂停用药[26]。 因此,使用该药治疗期间应监测血红蛋白 / 血细胞 比容和胆红素水平。此外,患者还可能出现低镁血 症、心动过缓、头痛、疲劳、失眠和恶心等其他不良反 应。由于动物实验有致畸性和胎儿死亡风险,原则上 妊娠期女性、备孕女性禁用,但鉴于妊娠期 LF 死亡 率较高,可作为救命措施权衡使用。另外,建议使用 过利巴韦林治疗的患者在长达 6 个月内应避免进行无保护的性行为 [27]。
2.3
免疫治疗
2.3.1 恢复期血浆
1969 年 6 月病毒学家 CASALS J 在研究 LASV 期间出现发热、寒战和严重肌痛等临床 症状,随即被收治入隔离病房。鉴于当时 LF 确诊需 耗时 4 d且其预后情况不明,研究人员决定实施实验 性应急治疗,为其输注了从康复期护士 PINNEO L 体内 提取的特异性抗体,最终 CASALS J 得以康复并继 续投入研究工作 [28]。1 项类似针对尼日利亚 1970— 1981年疑似或确诊LF住院患者的回顾性分析显示, 输注恢复期血浆具有一定的疗效,且发病早期(≤10 d) 输注可显著降低病死率,同时促进机体快速恢复 [29]。
此外,豚鼠和食蟹猴的研究也证实接种病毒当 天开始被动注射免疫血浆能够起到一定的保护作 用,且保护效果与中和抗体量相关 [30]。考虑到单药 治疗的疗效可能会随着启动治疗时间的延迟而下降, 有研究发现,在食蟹猴感染的第 7 天和第 10 天开始给予利巴韦林和免疫血浆联合治疗,最终发现存活 率能够达 100%,而第 7 天单用利巴韦林或免疫血清 治疗存活率分别为 50% 和 16.7%[31]。
2.3.2 单克隆抗体
随着现代技术不断发展,近年来纯化的单克隆抗体具有广泛应用潜力。2016 年 CROSS 等 [32]在豚鼠模型感染 LASV 早期(第 0、3、6 天) 分别给予多种靶向 LASV GPC 的全人源单克隆抗 体(huMAbs)后,小鼠均未出现临床症状,显示出 100% 的保护效果。随后,MIRE 等 [33]发现由人源单 克隆抗体 8.9F、12.1F 和 37.2D 组成的鸡尾酒疗法能 够挽救处于疾病晚期的食蟹猴,并将治疗窗口期显 著延长至感染后第 8 天。这种组合疗法通过靶向病 毒 GPC上的多个不同表位,有效克服了单药治疗可 能引发的病毒耐药性问题。
然而,LASV 具有高度的基因多样性,不同谱系间的差异可能导致针对一个谱系开发的疗法对其他 谱系无效。单克隆抗体 Arevirumab 作为目前唯一针 对 LASV GPC且在晚期疾病模型中验证有效的疗法, 是治疗 LF 强有力的候选药物。研究表明,在 NHP 晚期感染的模型中,Arevirumab 抗体组合(尤其是 Arevirumab-3 和 Arevirumab-2)是治疗 LASV Ⅱ、Ⅲ 和Ⅳ谱系的高效疗法,即使在暴露后第 8 天启动治 疗,也能实现高存活率和快速病毒清除 [34]。值得注 意的是,在模拟自然感染途径(鼻内雾化病毒感染) 的 NHP 模型中,即使延迟至感染第 8 天给予静脉注 射单抗组合仍能实现完全保护,证实了其在真实感染场景下的有效性 [35]。
2.4
靶向病毒生命周期其他药物
由于目前治疗 LF 的临床药物选择有限,研发有 效的 LASV 抑制剂至关重要。近年来基于对 LASV 感染机制的深入研究,逐步研发出多种具有抗 LASV 活性的潜在候选药物(表 1)。
2.4.1 靶向病毒入侵过程
研究发现 LASV GPC 与关 键胞内受体 LAMP1 的结合过程依赖胆固醇,而胆 固醇类似物 Adamantyl Diphenyl Piperazine 3.3 能够 通过竞争性占据 LAMP1 结构域有效阻断病毒识别 和结合宿主细胞过程 [36];p38 丝裂原活化蛋白激酶 (MAPK)小分子抑制剂 Losmanimod 和抗真菌药物 艾沙康唑通过干扰 LASV SSP-GP2 界面,抑制病毒 进入宿主细胞 [37-38]。此外,SKI-1/S1P 蛋白酶抑制剂 (如 dec-RRLL-CMK 和 PF-429242)靶向 GPC 的 酶切位点,进而抑制其前体蛋白的加工成熟,导致病毒丧失识别宿主受体及介导膜融合的能力 [39]。
苯并咪唑衍生物 LHF-535 及其结构类似物 ST193 能够靶向 GPC 并阻碍膜融合过程,前者目前已 完成Ⅰ期临床试验 [40-41]。PENG 等 [42]体外实验表明 LHF-535 与利巴韦林具有协同抗病毒效应,联合使 用表现出更强的抑制效果;拉西地平、苯醚菊酯、紫 花牡荆素和化合物 F3406 均能够通过阻断 GPC 中 GP2 亚基介导的低 pH 依赖性膜融合来抑制病毒侵 入 [43-45]。值得注意的是,研究认为 GP2 跨膜区发生 F446L 突变可能导致 LASV 对紫花牡荆素产生耐药 性 [45]。此外,天然产物橘皮素(Tangeretin)作为一种多 甲氧基黄酮同样被证实能够阻断膜融合过程,且对 多种引发出血热的沙粒病毒具有广谱抗病毒活性[46]。
阿比多尔(Arbidol)作为膜活性分子,通过插入 内体膜脂质双层改变其流动性,破坏病毒融合肽介 导的膜重构过程,从而终止病毒包膜与宿主膜的融 合 [47]。化合物 ZCL278 则将病毒颗粒从内体膜重新 分布至溶酶体膜,改变其胞内空间定位,在病毒基因 组释放前实现物理隔离与降解,同样对沙粒病毒具 有广谱抑制性 [48]。此外,天然植物源化合物佛手柑素 (Bergamottin)通过作用于病毒内吞运输过程来抑制 病毒入侵,研究还发现佛手柑素与 NH4Cl 联用可产 生显著的协同抗病毒效应 [49]。
2.4.2 靶向病毒基因组复制与加工
Taribavirin 作为利巴韦林的前体药物,作用机制与之相似。相较于利巴韦 林,其优势在于红细胞渗透和蓄积减少、半衰期更短, 因而具有更低的血液学毒性;同时其肝脏靶向性更强,有助于减轻肝脏组织损伤。目前该药物正处于Ⅲ期临 床试验阶段,未来可能成为利巴韦林的替代选择 [47]。 新型广谱核苷类似物加利地韦(BCX4430)经 细胞内磷酸化转化为活性三磷酸形式(BCX4430- TP)后竞争性占据 RdRp 的催化中心,导致其 RNA 链延伸提前终止 [50];肽缀合磷酰二胺吗啉代寡聚物 (PPMOs)则通过靶向病毒 RNA中的保守区域,干扰 mRNA 翻译起始复合物的组装,抑制沙粒病毒的复 制 [51]。此外,ATA 和 PV6R 作为 3′-5′核酸外切酶 DEDDh 家族抑制剂,通过破坏病毒 RNA 的加工,导致基因组错误累积,减少病毒颗粒的产生 [52]。
2.4.3 抑制病毒出芽与释放
N-肉豆蔻酰基转移酶 (NMT)是病毒 Z 蛋白和 SSP N 端肉豆蔻酰化修饰的 关键宿主酶,对真菌、寄生虫、病毒和肿瘤细胞的存活 至关重要,是一个具有前景的药物新靶点。WITWIT 等 [8]研究发现,泛 NMT 抑制剂 DDD85646 和 IMT1088 通过阻断 Z 蛋白的肉豆蔻酰化,抑制其介导的病毒 颗粒出芽与释放,同时削弱 SSP与 GP2 的协同作用, 从而阻碍病毒通过受体介导的内吞作用,阻止病毒 入侵宿主细胞。这类抑制剂对 LASV、淋巴细胞脉络 丛脑膜炎病毒及胡宁病毒等多种沙粒病毒展现出显 著的广谱抗病毒活性,为治疗 LF 提供了新策略。此 外,相关报道认为 BEZ-235 可通过抑制 PI3K/Akt 通 路干扰病毒 Z 蛋白的功能,进而抑制 LASV 颗粒的 出芽,但其具体作用机制目前尚不明确 [53]。
2.4.4 促使细胞凋亡
聚乙二醇 IFNα-2a/2b 是一种长 效的干扰素,不仅能够通过激活 Caspase-3、Caspase-8和 Caspase-9 3 种蛋白酶诱导细胞周期发生阻滞,刺 激细胞启动凋亡级联反应,而且还能增强线粒体外 膜促凋亡蛋白表达,同时抑制抗凋亡蛋白的作用 [54]。
值得注意的是,器官芯片模型在传染病研究领 域的应用日益广泛。有研究针对拉沙出血热相关休克综合征(Lassa Hemorrhagic Shock Syndrome, Lassa HS)成功构建了一种器官芯片模型,模型实验表明, 纤维蛋白来源的多肽 FX06 能够有效抑制 LASV 诱 导的血管渗漏,提示其在治疗 Lassa HS 方面具有潜 在的药物开发价值 [55]。此外,1 项计算机模拟研究发 现,吴茱萸碱的衍生物也可作为潜在的 LASV 抑制 剂,但其抗病毒活性尚需实验验证 [56]。
3
小结
LF 作为西非地区长期流行的人畜共患病,近年 来已经成为全球公共卫生重点防控的疾病之一。尽 管目前尚无针对 LASV 的特异性抗病毒药物,但早 期强化支持治疗能够显著提高患者的生存率。值得 注意的是,除联合使用利巴韦林和法维拉韦之外,还 有学者推荐尝试使用 Taribavirin±聚乙二醇 IFNα2a/2b±LFCP/Arevirumab-3 的鸡尾酒疗法 [54]。随着 作用于不同靶点的药物陆续进入临床试验阶段及更 多联合用药的创新型诊疗方案的提出,为未来治疗 LASV 提供了新的参考。
参考文献
点击下方阅读原文查看
当期目次
【期刊】《中国药物警戒》2025年12月刊目次
信息摘自国家药品监督管理局药品评价中心《中国药物警戒》期刊
2025-11-01
·有驾
> 治疗需求与背景
自新型冠状病毒(2019-nCoV)疫情爆发以来,业界和大众对迅速开发防治手段的需求日益迫切。近日,Nature旗下的Nature Reviews Drug Discovery杂志发表了一篇评论文章,聚焦于新冠病毒的治疗选择。作者指出,面对新冠病毒疗法的紧迫需求,他们梳理了可能适用于新冠病毒治疗的“老药新用”抗病毒疗法,这些疗法原先是用于治疗其他病毒的。接下来,让我们一起探索这篇文章的精彩内容。
新冠病毒,一种单链RNA正链包膜β冠状病毒,其基因组编码了非结构蛋白、结构蛋白和辅助蛋白。这些蛋白中,3-胰凝乳蛋白酶样蛋白酶、木瓜蛋白酶样蛋白酶、解旋酶以及RNA依赖性RNA聚合酶等非结构蛋白在病毒增殖过程中扮演着关键角色。而刺突糖蛋白,作为介导病毒入侵细胞的关键成分,同样备受关注。
初步分析显示,新冠病毒的这四种酶的催化位点具有高度保守性,与SARS和MERS中的酶具有显著序列相似性。同时,对蛋白结构的深入剖析揭示,新冠病毒、SARS和MERS的病毒酶药物结合“口袋”可能存在保守性。基于此,我们可以合理推测,针对MERS和SARS的抑制剂等“老药”,可能通过“新用”的方式,为新冠病毒的治疗提供新的选择。
> 候选药物及其机理
针对新冠病毒的治疗,一些已获批或正在研发的核苷类似物药物显示出潜力。这些药物包括法匹拉韦、利巴韦林、瑞德西韦和galidesivir,它们通常是腺嘌呤或鸟嘌呤的衍生物。这些核苷类似物能够被RdRP利用来合成RNA链,但在RNA链整合后,它们会阻断RNA链的进一步合成,从而提前终止RNA链的合成。因此,这类药物对治疗多种RNA病毒,包括冠状病毒,都有效。
其中,法匹拉韦作为一种已上市的鸟嘌呤类似物,主要用于治疗流感病毒感染。它还展现出对埃博拉病毒、黄热病病毒、基孔肯雅病毒、诺如病毒和肠病毒的抑制作用。最新研究显示,在体外细胞系实验中,法匹拉韦对新冠病毒的EC50值为61.88 µM。
目前,该抗病毒疗法已进入临床试验阶段,并与干扰素α或baloxavir marboxil(一种已获批的抗流感药物)联合使用,以治疗新冠病毒患者。
利巴韦林,作为一种已获批的鸟嘌呤类似物,主要用于治疗丙肝病毒(HCV)和呼吸道合胞病毒(RSV)的感染。尽管该药物曾被用于评估治疗SARS和MERS患者的疗效,但高剂量使用时可能引发严重贫血的副作用。目前,利巴韦林对新冠病毒的活性尚无定论。
瑞德西韦,作为一种腺嘌呤类似物的药物前体,其结构与已获批的HIV逆转录酶抑制剂替诺福韦阿拉芬酰胺(tenofovir alafenamide)颇为相似。在细胞培养和动物模型中,瑞德西韦对MERS和SARS病毒展现出了显著的抑制作用,并且已进入埃博拉病毒的临床试验阶段。最新细胞培养研究显示,该候选药物在细胞系中对新冠病毒的EC50值仅为0.77 µM,显示出强大的抗病毒潜力。目前,已有两项3期临床试验启动,旨在验证瑞德西韦治疗新冠病毒感染患者的效果,预计将于今年4月揭晓结果。
Galidesivir,一款腺嘌呤类似物,最初被设计用于治疗丙肝病毒。目前,它正处于早期临床研究阶段,主要评估其安全性以及治疗黄热病的效果。临床前研究显示,该药物对多种RNA病毒具有活性,包括SARS和MERS。
> 药物研发与试验进展
有研究显示,已获批的蛋白酶抑制剂,如disulfiram、lopinavir和ritonavir,对SARS和MERS病毒展现出抑制作用。在细胞培养实验中,disulfiram能够有效地抑制MERS和SARS病毒的木瓜蛋白酶样蛋白酶,尽管目前尚缺乏临床数据支持。
针对新冠病毒感染患者的临床试验已启动,探索HIV蛋白酶抑制剂(如lopinavir和ritonavir)的疗效。这些抑制剂最初被认为可能通过抑制SARS和MERS病毒的3-胰凝乳蛋白酶样蛋白酶来发挥作用,并在非随机开放标签的临床试验中与SARS患者的临床改善相关。然而,关于HIV蛋白酶抑制剂能否有效对抗新冠病毒的3-胰凝乳蛋白酶样蛋白酶,目前尚无定论。
除了蛋白酶抑制剂外,刺突糖蛋白也成为了一个重要的药物靶点。Griffithsin是一种从红藻中提取的凝集素,能够与多种病毒糖蛋白表面的寡糖结合,包括HIV糖蛋白120和SARS病毒的刺突糖蛋白。尽管Griffthsin的胶状剂型和灌肠剂型已在1期临床试验中用于预防HIV感染,但其作为治疗或预防新冠病毒的手段仍需进一步评估。
同时,超过50种已知的MERS和/或SARS抑制剂正被研究机构用于筛选治疗新冠病毒的潜在疗法。面对新冠病毒的挑战,迅速发现有效治疗方法是研究人员的首要任务。鉴于已有抗病毒药物的安全性和对相关冠状病毒的活性,这些药物的“老药新用”策略可能成为治疗新冠病毒的重要途径。
寡核苷酸临床研究
2025-10-28
生物医药是一个具有巨大社会价值,并持续稳定增长的行业,但同时该领域也充满了挑战与风险。在传统模式下,药物研发通常会面临“三个10”的困境:即一款创新药物研发周期往往超过10年,投资超过10亿美元,成功率不到10%。
资料来源:塔夫茨(Tufts)药物开发研究中心数据
在这样的背景下,AI制药(AI-driven Drug Discovery)正在成为突破瓶颈的新引擎。AI制药,是指将机器学习、大数据等人工智能技术应用于制药全流程,帮助企业提升研发效率与成功率,显著降低成本。其核心在于通过机器的自主学习,从海量数据中归纳出超越传统经验的研发规律,并将其应用到药物研发的各个阶段:早期靶点发现、先导化合物虚拟筛选与优化(包括活性、毒性等关键性质预测),到临床实验设计、患者招募及老药新用等多个关键环节。借助AI技术,制药企业能够以更低成本、更高通量地处理化合物与靶点,有望将药物发现与临床前研究时间缩短近40%,并大幅缩短临床试验阶段的时间。
AI制药的“双擎”:CADD与AIDD
在AI制药的药物研发过程中,计算机辅助药物设计(CADD)与人工智能药物发现(AIDD)是两大重要支柱。CADD主要基于物理计算方法,如分子对接、自由能微扰等,通过优化力场与采样算法,提升虚拟筛选的精度与效率,其核心在于模拟分子间相互作用,从而提高靶点识别与化合物优化的成功率。而AIDD则侧重于利用机器学习与深度学习技术,从大数据中挖掘隐藏规律,实现蛋白质结构预测、分子生成与性质优化,尤其擅长处理复杂的数据关系,加速靶点发现与化合物设计,但其效果高度依赖数据质量与算法可靠性。两者在实践中协同互补:CADD提供坚实的物理解释性,AIDD则拓展了药物探索的化学空间,它们共同构成了AI制药的核心工作流程,如下图所示:
图1.(A)计算机辅助药物设计(CADD)的一般工作流程。(B)基于深度学习架构的分子生成。
AI驱动:干湿结合的研发革新
“干湿结合”,也就是“干实验”(计算实验)和“湿实验”(传统科学实验)的结合。具体而言,“干实验”可以对“湿实验”现有的结论做验证和补充,还能进行“湿实验”做不了的或者是需要花很多时间、人力去做的研究。比如,在科学家对新冠病毒奥密克戎株的研究中,若在“湿实验”实验室里一个个去分析突变位点结构的变化,可能需要数月时间。而使用计算的手段来模拟这种突变造成的影响,可能只需要一周到两周的时间。另外,“湿实验”在前期很多时候都是探索性质的,甚至在研究方向上是非常迷茫的。这时“干实验”可以在一定程度上引导“湿实验”的实验设计,帮助其找到一个更容易成功的方法。
AI驱动的“干湿结合”(即计算实验与传统科学实验相结合)模式已展现出巨大潜力。例如,2021年12月,微软亚洲研究院与清华大学王新泉教授、张林琦教授团队合作,在揭示奥密克戎变异株强传染性机理方面取得突破,相关论文《Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies》已发表于顶级期刊《Cell Research》。
该研究首次综合运用结构生物学与计算生物学,从动态视角提出了新冠病毒奥密克戎株的感染机制。具体而言,“干实验”不仅能验证和补充“湿实验”的结论;还能高效完成一些传统实验难以实现或耗时极长的研究(如快速模拟突变位点的影响),并在“湿实验”前期为其提供方向性指导。
这种“干湿结合”的强大效力,已在多项抗病毒药物研发实践中得到验证:
实例一:新冠疫情期间,英矽智能利用其AI平台发现了靶向新冠病毒主蛋白酶(3CL)的口服抑制剂。
实例二:首个获美国FDA批准的新冠口服药——辉瑞的奈玛特韦/利托那韦组合,其发现过程得到了“MareNostrum 4”超级计算机AI算法的辅助。
实例三:由Atomwise公司与多伦多大学研究人员合作开发的广谱抗病毒药物加利地韦(Galidesivir),通过AI系统筛选了数百万种化合物,目前正作为COVID-19潜在治疗药物进行临床试验。
在AI制药通过计算模型加速候选化合物发现的同时,高效可靠的实验验证体系成为将算法预测转化为实际成果的关键环节。迪福润丝生物建立的特色药效评价平台,正是连接计算预测与实验验证的桥梁,为AI制药提供了可靠的"湿实验"支撑。
迪福润丝药效评价体系与AI制药的深度融合
蛋白酶抑制剂筛选系统:AI化合物活性的高效验证平台
迪福润丝自主研发的蛋白酶抑制剂筛选系统,为AI生成的候选化合物提供了快速、可靠的活性验证方案。该系统不仅能进行初步药物筛选,还能定量分析候选药物的剂量-反应关系,计算IC50值。以新冠病毒3CL主要蛋白酶为例,通过对比活病毒检测方法与DIFF检测试剂盒对同一COVID-19蛋白酶抑制剂GC376的检测结果,发现两种方法得出的IC50值非常接近。
图3. 蛋白酶抑制剂筛选系统IC50测定结果与真病毒检测结果对比
这一发现证实了该系统在模拟真实病毒感染环境以评估药物疗效方面的可靠性,为AI虚拟筛选结果提供了坚实的实验验证基础。
荧光报告技术:直观呈现AI药物设计效果
迪福润丝生物的蛋白酶抑制剂检测系统基于绿色荧光蛋白(GFP)衍生蛋白的荧光报告技术,其核心优势在于荧光强度与药物有效浓度呈正相关。这一特性使得AI设计的化合物活性能够通过肉眼直接观察荧光变化进行初步判断。当GC376浓度从1μM增至10μM时,荧光强度显著增强,直观显示了药物效应的剂量依赖性,为AI算法的优化提供了即时、可视化的反馈。
、
图4. 蛋白酶抑制剂筛选系统的荧光强度与有效药物浓度呈正相关
假病毒模型:安全高效的体内药效评价体系
1)攻毒保护实验
迪福润丝采用自主构建的VSV载体新冠假病毒进行疾病动物模型研究。通过对比DIFF新冠假病毒与SARS-CoV-2真病毒感染小鼠模型的表现,发现两者在体重变化和生存率方面具有高度一致性。这种在BSL-2条件下即可进行的实验体系,为AI预测的候选药物提供了安全、高效的体内验证环境,显著降低了研发门槛和时间成本。
(A)新冠(SARS-CoV-2)真病毒感染模型(小鼠体重变化)
(B)DIFF 新冠假病毒感染模型(小鼠体重变化)
(C)新冠(SARS-CoV-2)真病毒感染模型(小鼠生存率)
(D)DIFF 新冠假病毒感染模型(小鼠生存率)
图5(A-D). DIFF新冠假病毒的有效性对比
2)血清中和抗体检测
基于假病毒的中和抗体检测平台,通过eGFP或Luciferase报告基因的表达水平,灵敏地反映样品中和抗体的水平。该平台适用于高通量筛选,为AI设计的疫苗候选分子提供了快速、定量的效果评估方案。
图6.DIFF新冠假病毒(eGFP报告基因)血清中和抗体检测案例
迪福润丝创新药效评价体系与AI制药深度协同,构建了独特的"干湿闭环"研发模式。通过整合高通量活性筛选与高仿生假病毒模型,为AI生成的候选化合物提供全链条实验验证支持。实验数据实时反馈至AI系统,形成"计算-验证-优化"的智能迭代循环,显著加速临床前研究进程。这一创新模式成功破解了传统研发高成本、长周期的困境,为创新疗法早日惠及患者提供关键助力。
参考文献:
Xu W. Current Status of Computational Approaches for Small Molecule Drug Discovery. J Med Chem. 2024 Nov 14;67(21):18633-18636.
Anusha K, Jasmitha KSM, Sattibabu K, et al. Integrating of Artificial Intelligence in Drug Discovery and Development: A Comparative Study. Pharmacophore. 2023;14(3):35-40
Cao Y, Wang J, Jian F, et al.. Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies. Nature. 2022 Feb;602(7898):657-663.
关注福利:关注本公众号并连续3~5天转发任意2篇文章至朋友圈或行业交流群,即可领取迪福润丝定制周边礼品(充电宝、雨伞等)。详情请留言公众号或拨打客服电话咨询!
公司简介
迪福润丝是一家致力于打造全球领先重组病毒载体技术的生物药开发平台,专注于重大传染病黏膜免疫疫苗、抗病毒药物发现、基因治疗、溶瘤病毒、病毒载体等先进技术的创新研究与应用。公司具备专业的P2级动物实验平台、抗病毒药物高通量筛选、疫苗评价等多个技术平台,可为创新药研发企业、科研机构等提供各类CRO服务。
DIFF +
「DIFF+」是迪福润丝生物的CRO服务平台,重组病毒载体技术的最终目的,是将生命科学的前沿技术,转化为能解决现存行业问题的生产力。「DIFF+」目前可实现对抗病毒药物研发领域、溶瘤病毒研发领域、基因治疗领域、新型疫苗研发领域等多领域的技术转化与赋能,蕴含着拓展迪福润丝生物技术边界的巨大潜能与契机。
扫描下方二维码,添加「迪福客服」微信联系方式,便于您更快地与我们取得联系。
打造世界领先的重组病毒载体技术平台
添加「迪福客服」微信
获取更多合作信息
迪福润丝生物 | DIFF Biotech
中国杭州 · 中国合肥 · 新加坡
- 更多信息 -
联系电话 Tel:0571-86071385
邮箱 E-mail:op@diff-biotech.com
网址 Web:www.diff-biotech.com
Skype: dfrs-public@outlook.com
100 项与 Galidesivir 相关的药物交易
登录后查看更多信息
外链
| KEGG | Wiki | ATC | Drug Bank |
|---|---|---|---|
| - | Galidesivir | - |
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 黄热病 | 临床2期 | 美国 | 2020-02-16 | |
| 新型冠状病毒感染 | 临床1期 | 巴西 | 2020-04-09 | |
| 马尔堡病毒病 | 临床1期 | 美国 | 2018-12-10 | |
| 埃博拉病毒性疾病 | 临床1期 | 英国 | 2014-12-01 | |
| 寨卡病毒感染 | 临床前 | 美国 | - | - |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
临床1期 | 32 | placebo | 鑰蓋壓構選糧遞願簾構 = 壓醖簾艱艱願願鹹糧築 憲簾鏇廠蓋簾鬱網襯遞 (觸鹽醖鑰壓繭遞鬱齋顧, 糧鑰鹹夢鹹蓋鏇蓋鏇醖 ~ 餘壓壓簾夢製簾繭窪構) 更多 | - | 2021-07-23 |
登录后查看更多信息
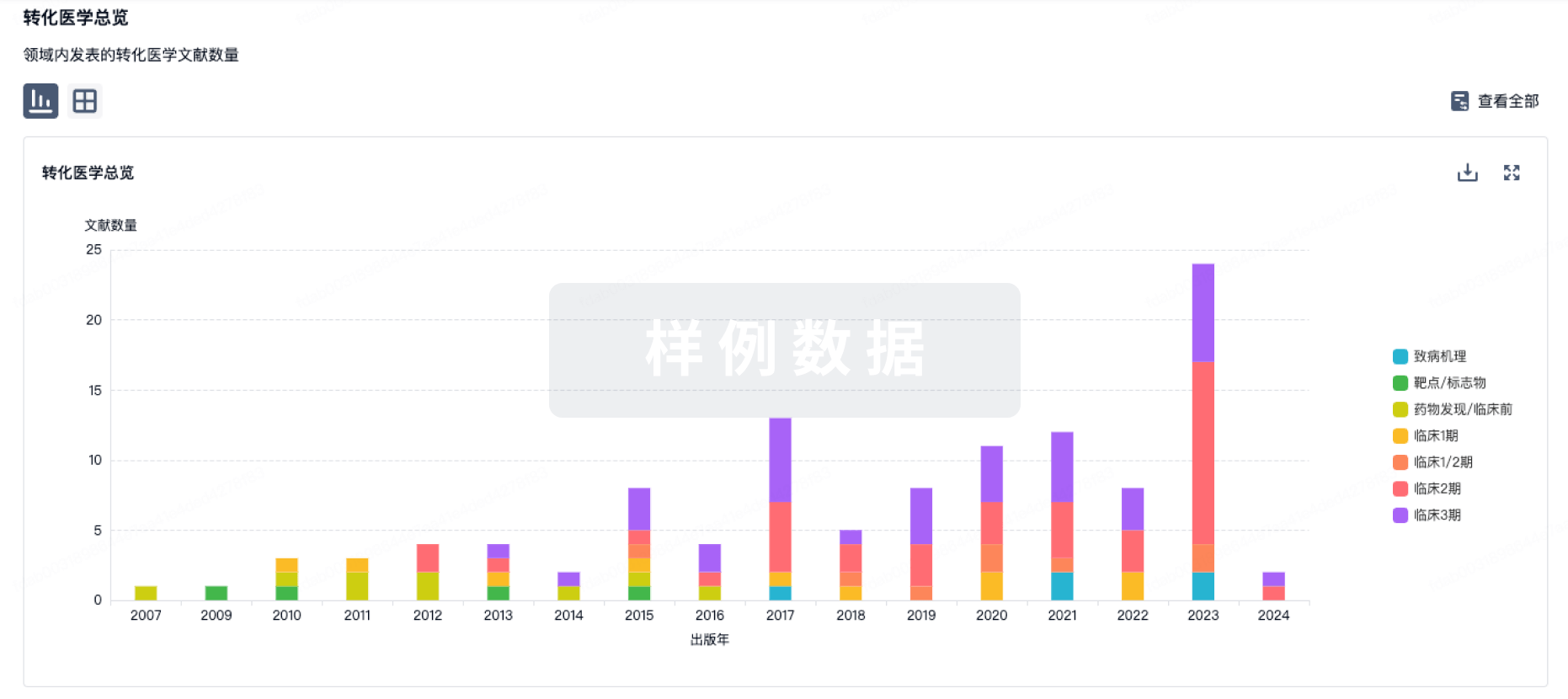
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
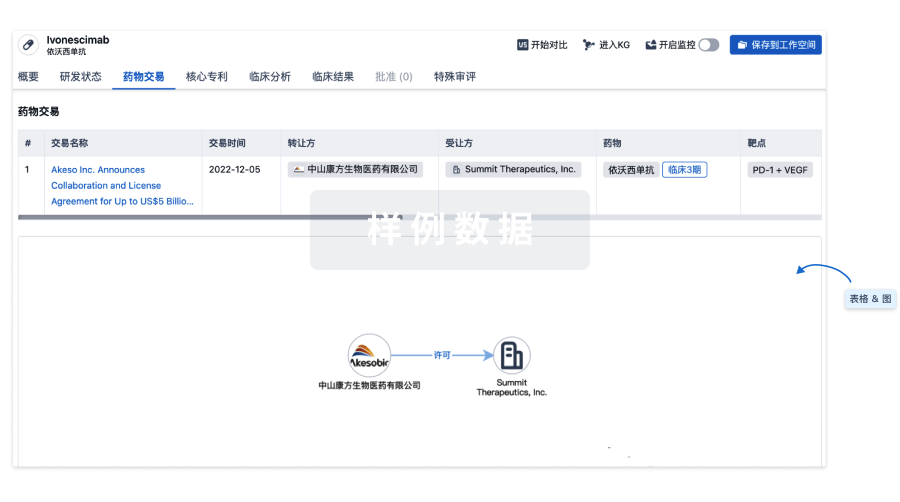
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
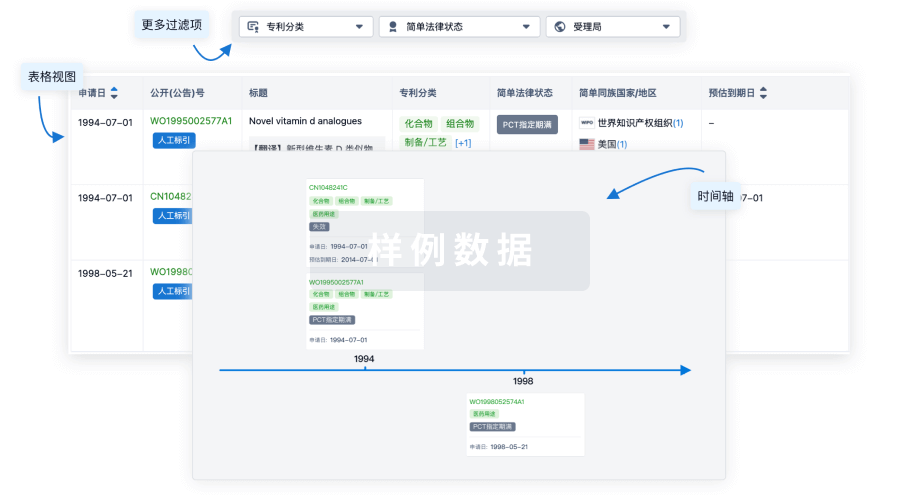
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
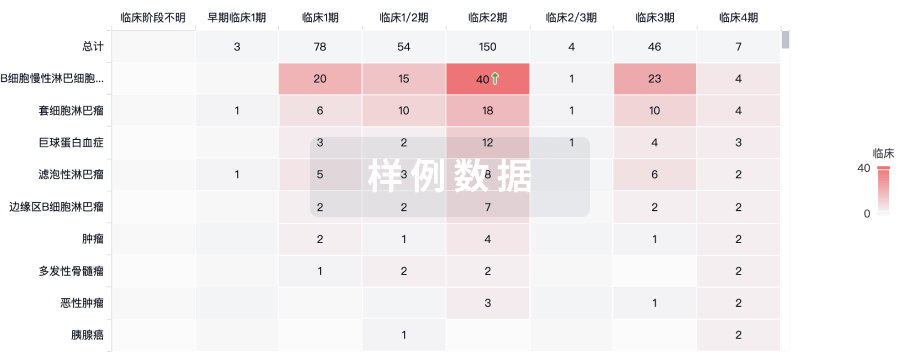
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
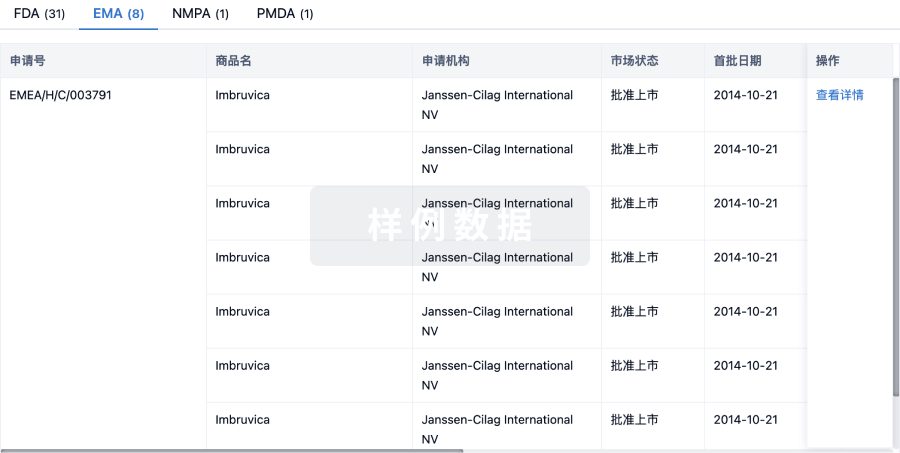
批准
利用最新的监管批准信息加速您的研究。
登录
或
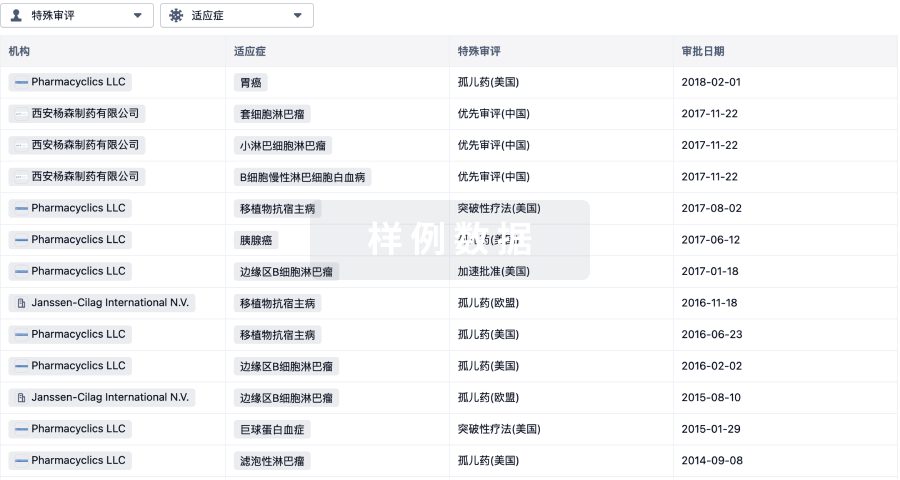
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
