预约演示
更新于:2025-05-07

Jumonji University
更新于:2025-05-07
概览
关联
6
项与 Jumonji University 相关的临床试验JPRN-UMIN000054040
Study on metabolism and bioavailability of newly developed saccharide material, 1-deoxymannose, in humans - Study on metabolism and bioavailability of 1-deoxymannose in humans
开始日期2024-04-10 |
申办/合作机构 |
JPRN-UMIN000053367
Effects of simultaneous intake of water-soluble liquid konjac and indigestible dextrin on gut microbiota composition in young women: an open-label, cross-over, before-after comparison, dose-dependent study - Effects of simultaneous intake of water-soluble liquid konjac and indigestible dextrin on gut microbiota composition in young women: an open-label, cross-over, before-after comparison, dose-dependent study
开始日期2024-01-16 |
申办/合作机构 |
JPRN-UMIN000051700
Development of food products aimed at stimulating salivation and modulating salivary viscosity - Development of food products aimed at stimulating salivation and modulating salivary viscosity
开始日期2023-07-25 |
申办/合作机构 |
100 项与 Jumonji University 相关的临床结果
登录后查看更多信息
0 项与 Jumonji University 相关的专利(医药)
登录后查看更多信息
364
项与 Jumonji University 相关的文献(医药)2025-08-01·Microbiological Research
ArcAB system promotes biofilm formation through direct repression of hapR transcription in Vibrio cholerae
Article
作者: Shimamoto, Toshi ; Shimamoto, Tadashi ; Caigoy, Jant Cres ; Yan, Zhiqun ; Nariya, Hirofumi
2024-12-01·Preventive Medicine Reports
Factors associated with acquiring exercise habits through health guidance for metabolic syndrome among middle-aged Japanese workers: A machine learning approach
Article
作者: Wakaba, Kyohsuke ; Onoue, Takeshi ; Tsushita, Kazuyo ; Wan, Jiawei ; Nakata, Yoshio
2024-10-01·Medicine & Science in Sports & Exercise
What Is The Optimal Choice Of Days For Dietary Recording Method For Collegiate Athletes?
作者: Murata, Hiroko ; Taguchi, Motoko ; Goshozono, Mika
100 项与 Jumonji University 相关的药物交易
登录后查看更多信息
100 项与 Jumonji University 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年02月08日管线快照
无数据报导
登录后保持更新
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
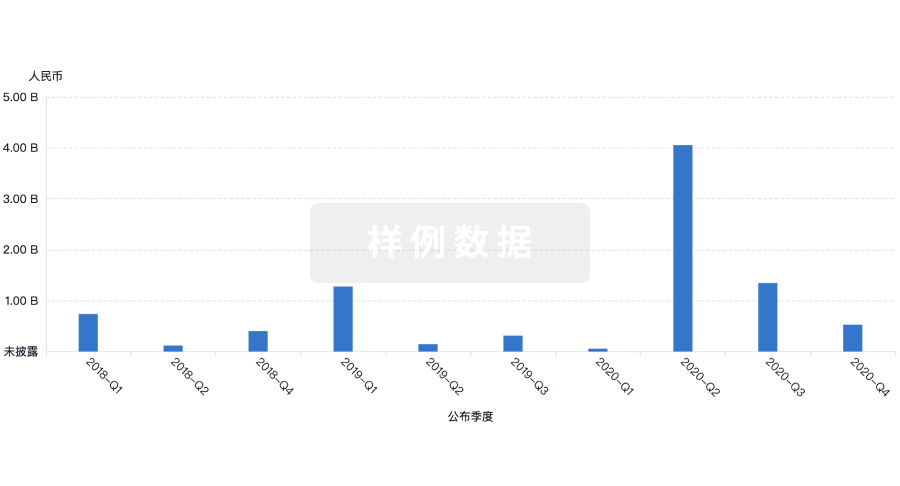
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
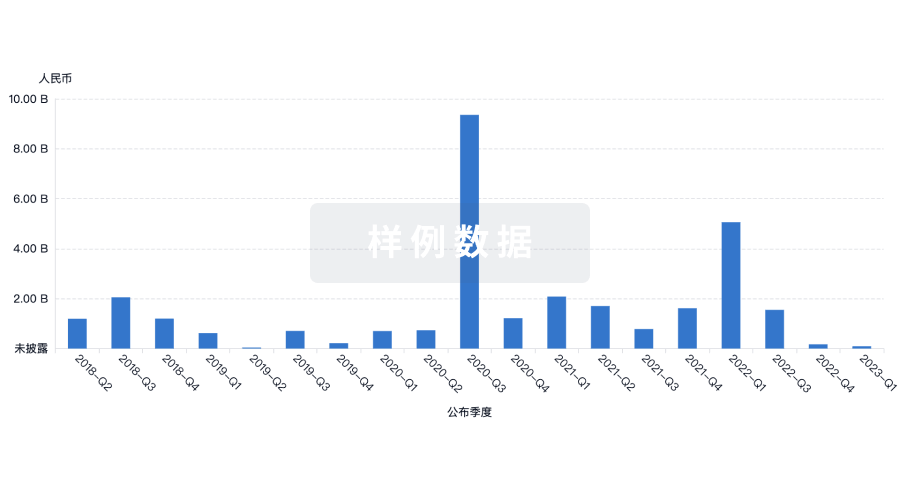
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
