预约演示
更新于:2025-05-07
ADRA2 x D2 receptor x Histamine receptor
更新于:2025-05-07
关联
1
项与 ADRA2 x D2 receptor x Histamine receptor 相关的药物作用机制 ADRA2拮抗剂 [+2] |
非在研适应症- |
最高研发阶段批准上市 |
首次获批国家/地区 日本 |
首次获批日期1977-01-01 |
5
项与 ADRA2 x D2 receptor x Histamine receptor 相关的临床试验IRCT20200913048708N1
Evaluation of efficacy and safety of adding Chlorpromazine to Atazanavir/Ritonavir regimen in the treatment of COVID-19 patients, a randomized double-blind clinical trial
开始日期2020-10-31 |
IRCT20090726002232N4
Comparison of Haloperidol, Promethazine, Triflupeazine and Chlorpromazine in terms of velocity and durability of the sedation among acute aggressive patients
开始日期2017-12-22 |
100 项与 ADRA2 x D2 receptor x Histamine receptor 相关的临床结果
登录后查看更多信息
100 项与 ADRA2 x D2 receptor x Histamine receptor 相关的转化医学
登录后查看更多信息
0 项与 ADRA2 x D2 receptor x Histamine receptor 相关的专利(医药)
登录后查看更多信息
5
项与 ADRA2 x D2 receptor x Histamine receptor 相关的文献(医药)2011-12-01·Therapeutic Advances in Psychopharmacology3区 · 医学
Receptor mechanisms of antipsychotic drug action in bipolar disorder – focus on asenapine
3区 · 医学
ArticleOA
作者: Reynolds, Gavin P
2011-05-01·Pain1区 · 医学
Influence from genetic variability on opioid use for cancer pain: A European genetic association study of 2294 cancer pain patients
1区 · 医学
Article
作者: R. Sabatowski ; O. Dale ; S. Kaasa ; Kristin Bjordal ; V. Sigurdardottir ; S. Lundström ; L. Radbruch ; A. Caraceni ; F. Strasser ; F. Skorpen ; A. Davies ; M. Kloke ; M. Maltoni ; T. Fladvad ; P. M. Fayers ; P. Klepstad
2008-12-01·Neuropsychopharmacologia Hungarica : a Magyar Pszichofarmakologiai Egyesulet lapja = official journal of the Hungarian Association of Psychopharmacology
Quetiapin alkalmazása bipoláris zavarban.
Review
作者: Sümegi, András
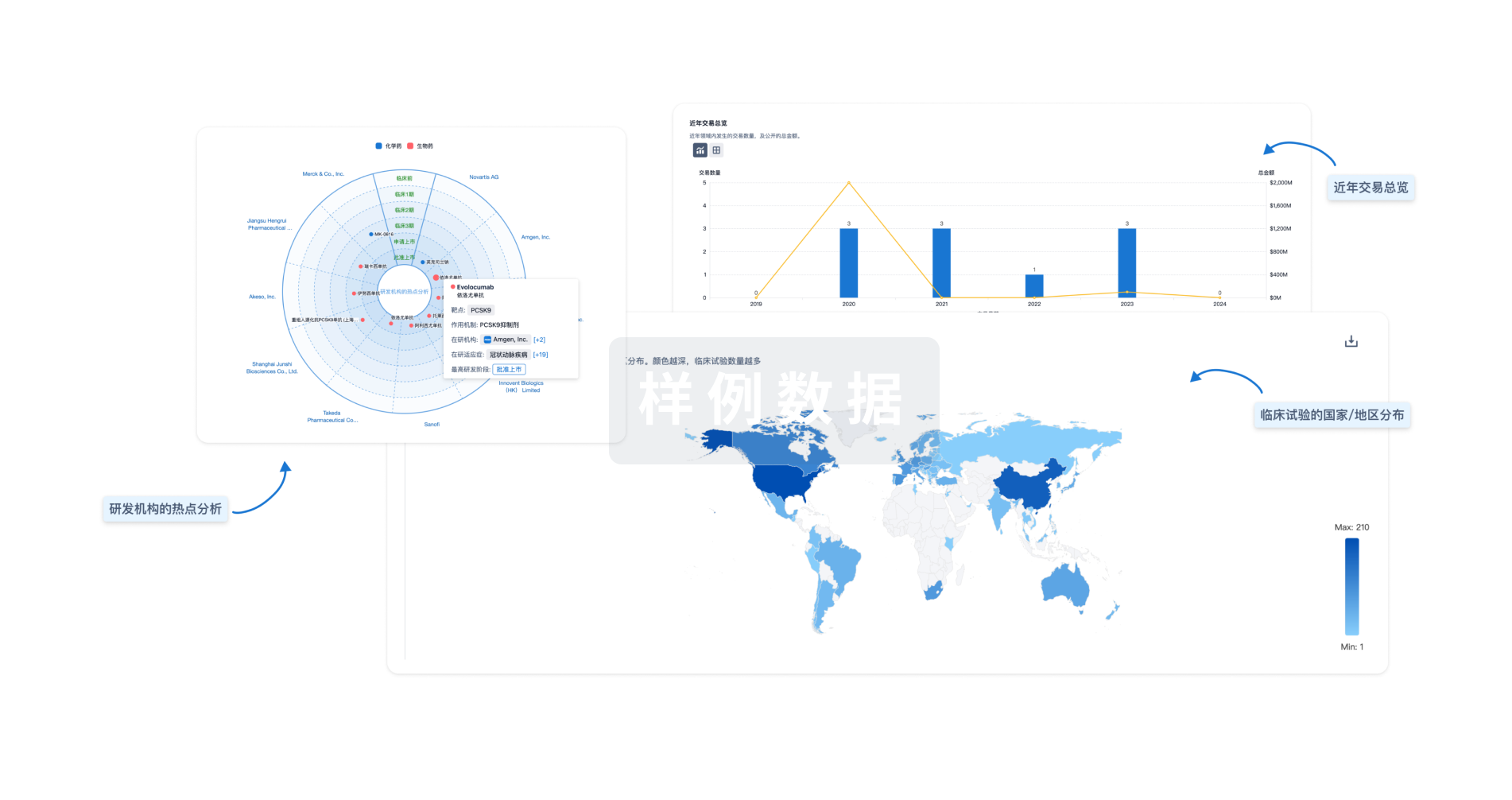
分析
对领域进行一次全面的分析。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用


