预约演示
更新于:2025-05-07
GPI
更新于:2025-05-07
基本信息
别名 AMF、Autocrine motility factor、glucose-6-phosphate isomerase + [9] |
简介 In the cytoplasm, catalyzes the conversion of glucose-6-phosphate to fructose-6-phosphate, the second step in glycolysis, and the reverse reaction during gluconeogenesis (PubMed:28803808). Besides it's role as a glycolytic enzyme, also acts as a secreted cytokine: acts as an angiogenic factor (AMF) that stimulates endothelial cell motility (PubMed:11437381). Acts as a neurotrophic factor, neuroleukin, for spinal and sensory neurons (PubMed:11004567, PubMed:3352745). It is secreted by lectin-stimulated T-cells and induces immunoglobulin secretion (PubMed:11004567, PubMed:3352745). |
关联
2
项与 GPI 相关的药物CN115772228
专利挖掘靶点 |
作用机制- |
在研适应症 |
非在研适应症- |
最高研发阶段药物发现 |
首次获批国家/地区- |
首次获批日期1800-01-20 |
WO2023088246
专利挖掘靶点 |
作用机制- |
在研机构 |
原研机构 |
在研适应症 |
非在研适应症- |
最高研发阶段药物发现 |
首次获批国家/地区- |
首次获批日期1800-01-20 |
100 项与 GPI 相关的临床结果
登录后查看更多信息
100 项与 GPI 相关的转化医学
登录后查看更多信息
0 项与 GPI 相关的专利(医药)
登录后查看更多信息
6,618
项与 GPI 相关的文献(医药)2025-06-01·Research in Veterinary Science
The effect of high-level dietary Laminaria digitata on the muscle proteome and metabolome of weaned piglets
Article
作者: Sergeant, Kjell ; Carvalho, Daniela F P ; Charton, Sophie A B ; Cocco, Emmanuelle ; de Almeida, André M ; Costa, Mónica M ; Freire, João P B ; Renaut, Jenny ; Sacarrão-Birrento, Laura ; Ribeiro, David M ; Leclercq, Céline C ; Prates, José A M
2025-05-01·Annals of Laboratory Medicine
Establishment of a Multilocus Sequence Typing Scheme for Pasteurella canis Using Isolates from Infected Humans and Diseased Companion Animals
Article
作者: Kim, Jae-Seok ; Goto, Mieko ; Shizuno, Kenichi ; Tsuyuki, Yuzo ; Takahashi, Takashi ; Yoshida, Haruno ; Maeda, Takahiro
2025-05-01·Clinical Anatomy
Number of serotonergic neurons in the subthalamic nucleus and globus pallidus internus could influence the effects of deep brain stimulation in Parkinson 's disease
Article
作者: Sharma, Mayank ; Rao, Asha ; Munawara, Rafika ; Gupta, Tulika
7
项与 GPI 相关的新闻(医药)2024-10-30
2024年10月29日,天津医科大学王晓虹团队在期刊《Nature Communications》上发表了题为“Protein O-GlcNAcylation coupled to Hippo signaling drives vascular dysfunction in diabetic retinopathy”的研究论文。从机制上讲,团队发现Yes相关蛋白(YAP)和带有PDZ结合基序(TAZ)的转录共激活因子是Hippo通路的关键效应子,在糖尿病视网膜病变中被O-GlcNAcyl化。这项研究强调了O-GlcNAc-Hippo轴在糖尿病视网膜病变发病机制中的关键作用,并表明其作为治疗靶点的潜力。
https://www.nature.com/articles/s41467-024-53601-x
关于糖尿病视网膜病变
01
糖尿病的血管并发症,包括动脉粥样硬化、糖尿病肾病和视网膜病变,是该疾病最严重的表现之一。在这些并发症中,糖尿病视网膜病变(DR)是导致整体视力丧失和失明的主要原因。虽然激光光凝术和血管内皮生长因子(VEGF)中和疗法是治疗DR的常用方法,但它们有局限性。激光光凝疗法可破坏视网膜组织,导致暗点,并且由于这些药物的短暂作用,抗VEGF疗法对增殖性视网膜病变的有效性受到限制,因为需要频繁干预。此外,一些患者可能对VEGF靶向药物反应不佳或没有反应,在某些情况下,VEGF中和会加速光感受器的萎缩。
内皮细胞(EC)形成血管的最内层,这使它们与血液直接接触,并使它们能够感知血液中的营养水平。EC表现出高糖酵解活性,以与许多癌细胞相似的速度消耗葡萄糖,并且主要依靠糖酵解产生约85%的ATP6。糖酵解的几种关键调节因子,包括PFKFB3、ADORA2A和HK2,已被证明在调节ECs能量稳态和血管生成中起关键作用。
O-GlcNAc稳态失调与多种慢性人类疾病有关,包括心血管疾病、阿尔茨海默病和癌症。尽管如此,O-GlcNAc信号转导在调节ECs中的作用,特别是在糖尿病视网膜病变的情况下,仍然难以捉摸。团队研究了O-GlcNAcylation在糖尿病视网膜病变期间血管疾病中的作用,并评估了抗血管生成的治疗潜力。
抑制O-GlcNAcylation和YAP/TAZ,降低了DR病理
02
团队研究了两种抑制剂OSMI-1和Super-TDU在减少DR病理方面的治疗潜力。OSMI-1是一种OGT小分子抑制剂,之前已从高通量筛选结果优化。研究结果表明,OSMI-1处理显著增强了STZ模型中的壁细胞覆盖率。此外,它有效地减少了OIR模型中的新生血管形成、血岛面积和FITC-葡聚糖外渗,表明其作为DR治疗剂的潜力。
Super-TDU是一种靶向YAP-TEAD相互作用的抑制剂肽,抑制YAP介导的TEAD反式激活和YAP功能。在STZ模型中,Super-TDU和EXO超级TDU通过增加壁细胞覆盖率和减少脱细胞毛细血管的数量,表现出保护作用。EXO超级TDU与Super-TDU相比,表现出卓越的疗效。在OIR模型中,Super-TDU和EXO超级TDU有效抑制新生血管形成、视网膜出血和FITC-葡聚糖外渗。此外,EXO超级TDU与Super-TDU相比,表现出更好的疗效。这些结果表明,OGT或YAP/TAZ的药理学靶向,可能是减少DR病理的一种有前途的治疗方法。
基于外泌体的YAP/TAZ抑制剂递送,可缓解DR中的血管功能障碍。
YAP/TAZ调节ECs中的葡萄糖代谢和O-GlcNAcylation
03
与正常受试者相比,PDR患者的视网膜微血管ECs,表现出HK1、HK2、GPI、PGM3和OGT基因的上调,其中两个显示出统计学意义。YAP/TAZ可能是HBP和O-GlcNAcylation的关键调节因子。在YAP/TAZ敲低HRCECs中,总体O-GlcNAc修饰的蛋白水平显著降低,恢复YAP/TAZ,可以挽救O-GlcNAc水平。这些结果表明,O-GlcNAcylation激活YAP/TAZ,通过诱导促血管生成和葡萄糖代谢转录程序,在促进血管功能障碍中起关键作用。这种调节机制可能有助于形成正反馈回路,从而驱动DR中血管功能障碍的进展。
YAP/TAZ调节ECs中的葡萄糖代谢和O-GlcNAcylation。
总结
04
1. O-GlcNAcylation的作用:研究发现,响应高葡萄糖或缺氧暴露的蛋白质O-GlcNAcylation,在介导视网膜脉管系统中的内皮活化和病理性血管生成中起关键作用。
2. Hippo信号通路的激活:hyper-O-GlcNAcylation水平通过改变YAP磷酸化和蛋白质稳定性,导致Hippo通路的激活,这是关键的血管生成调节途径。
3. 治疗潜力:通过遗传学和药理学方法靶向O-GlcNAcylation-Hippo调节轴,可减轻STZ模型中的壁细胞丢失和血管渗漏,并减轻OIR模型中的病理性视网膜新生血管形成。
4. O-GlcNAcylation与YAP/TAZ的相互作用:研究发现,在糖尿病患者中,葡萄糖和缺氧通过向YAP/TAZ添加O-GlcNAc部分有效激活YAP/TAZ,促进血管渗漏和病理性视网膜新生血管形成。
5. 外泌体介导的药物递送系统:研究利用外泌体介导的药物递送系统加载治疗量的YAP/TAZ抑制剂Super-TDU,展示了对视网膜血管的高效和特异性靶向。
6. 结论:研究确定了O-GlcNAc修饰在DR中的关键作用,并揭示了Hippo信号通路和蛋白质O-GlcNAcylation之间的相互作用,强调了在糖尿病视网膜病变中靶向O-GlcNAcylation-Hippo信号轴的潜在治疗价值。
参考资料:
1. Ogurtsova, K. et al. IDF diabetes atlas: global estimates for the prevalence of diabetes for 2015 and 2040. Diabetes Res. Clin. Pract. 128, 40–50 (2017).
2. Antonetti, D. A., Klein, R. & Gardner, T. W. Diabetic retinopathy. N. Engl. J. Med. 366, 1227–1239 (2012).
2023-11-02
EDINA, Minn., Nov. 2, 2023 /PRNewswire/ -- Surgical Information Sciences, a medical device company focused on improved visualization of brain structures for deep brain stimulation (DBS) surgery, announced the commencement of its post-market study, the Visualization of the STN and GPi for DBS Surgery in Patients with Parkinson's Disease (VISION Study). This study aims to evaluate the potential of SIS technology in enhancing the accuracy of DBS implant placement for patients battling Parkinson's Disease. The VISION Study will involve the participation of 90 patients across multiple sites, and it represents a significant step in the pursuit of improved treatment for those living with this challenging condition.
Millions of people worldwide are impacted by Parkinson's Disease and it affects their daily lives. DBS has shown promise in managing parkinsonian symptoms, but its success is closely tied to the precision of implant placement within the subthalamic nucleus (STN) and globus pallidus internus (GPi). SIS believes that its cutting-edge visualization technology has the potential to significantly improve the accuracy of this procedure, ultimately leading to improved patient outcomes.
Dr. Patrick Senatus, a neurosurgeon at Hartford Hospital in Hartford, CT, commented, "The VISION Study presents an opportunity to assist surgeons and programming physicians in treating patients with Parkinson's Disease. By enhancing visualization, the accuracy of DBS implant placement may be improved through further customizing targeting of therapeutic brain regions. I am enthusiastic about the possibilities that this research holds."
"Today marks a momentous occasion for SIS. We are proud to take the first step toward demonstrating improved outcomes of DBS surgery through patient specific visualization of target structures and lead placement," said Brad Swatfager, President and Chief Executive Officer. "Our team has worked hard to develop this cutting-edge technology, and we look forward to collaborating with healthcare professionals across the country to explore its full potential and improve the experience for physicians and patients."
About Surgical Information Sciences: Surgical Information Sciences has developed software designed to enhance clinical images for the visualization of anatomical structures of the brain. The software applies deep learning models to predict and visualize the shape and location of the structures and is currently cleared by the FDA and has CE Mark.
For more information about Surgical Information Sciences, please visit our website at .
Media Contact: Susan Vnoucek, [email protected]
SOURCE Surgical Information Sciences
2023-06-06
关注并星标CPHI制药在线 ---智药研习社6月线下会议报名---6月5日,诺华的「盐酸伊普可泮胶囊」被CDE拟纳入优先审评,用于成人阵发性睡眠性血红蛋白尿症(PNH)患者的治疗。 盐酸伊普可泮(iptacopan,LNP023)是诺华研发的首个靶向补体旁路途径B因子的口服抑制剂,作用于补体系统C5末端通路的上游,同时控制血管内溶血和血管外溶血,弥补了抗C5抗体的不足,被开发用于治疗多种补体介导疾病,如阵发性睡眠性血红蛋白尿症(PNH)、C3肾小球病(C3G)、IgA肾病(IgAN)、非典型溶血性尿毒症综合征(aHUS)、膜性肾病(MN)、狼疮性肾炎(LN)以及免疫性血小板减少性紫癜(ITP)和冷凝集素病(CAD)等。 2023年5月,诺华宣布评估iptacopan单药治疗未接受过补体抑制剂(包括抗C5抗体)治疗的PNH成人患者疗效和安全性的3期临床试验达到主要终点。该研究是一项单臂、开放标签、多中心研究,共入组40例平均血红蛋白(Hb)<10g/dL、乳酸脱氢酶(LDH)>1.5x正常值上限且未接受过补体抑制剂治疗的成人PNH患者。 结果显示:试验达到主要研究终点,92.2%患者在不输注红细胞(RBCT)情况下达到血红蛋白水平较基线升高≥2g/dl。次要终点也显示出临床意义上的改善,如62.8%的患者在RBCT情况下血红蛋白水平较基线升高≥12g/dl,97%的患者避免输血。 在国内,iptacopan先后于2022年3月和2023年2月被CDE纳入突破性治疗品种,分别用于治疗 C3肾小球病(C3G)和阵发性睡眠性血红蛋白尿症(PNH)。此次iptacopan被CDE纳入优先审评,将进一步加速其PNH适应症在国内的获批上市。 关于PNH及其治疗进展 阵发性睡眠性血红蛋白尿症(PNH)是一种罕见的获得性克隆性造血干细胞疾病,全球发病率约为1~2/百万, 亚洲地区发病率相较西方高。PNH是由于患者造血干细胞的PIG-A基因突变,使部分或完全血细胞膜糖化磷脂酰肌醇(GPI)锚合成障碍,造成血细胞表面的GPI锚连蛋白(如补体调节蛋白CD55、CD59)缺失,细胞灭活补体能力减弱,从而引起细胞被破坏和发生溶血等。PNH患者的临床表现主要为不同程度的发作性血管内溶血、阵发性血红蛋白尿、骨髓造血功能衰竭和静脉血栓形成。 补体系统是机体固有免疫的组成部分。在补体介导的溶血反应中,补体激活早期阶段被称为近端通路。PNH患者的红细胞因CD55缺乏导致C3裂解增多,C3a和C3b的形成增加,C3b介导的调理作用使红细胞在脾和肝脏被吞噬,导致血管外溶血(EVH)。C3a及C3b形成后产生的补体级联反应被称为远端通路。PNH患者的红细胞因CD59的缺乏使C5b更多的转化为膜攻击复合物(MAC),导致红细胞直接裂解,即血管内溶血(IVH)。 抗C5是公认的PNH标准治疗。目前,全球监管机构已批准3款PNH药物,即Soliris(依库珠单抗,阿斯利康/Alexion)、Ultomiris(ravulizumab,阿斯利康/Alexion)和Empaveli (pegcetacoplan,Apellis公司),其中仅Soliris在国内获批上市。 • Soliris是全球首个获批的C5补体抑制剂,通过选择性抑制末端补体C5蛋白质的激活来发挥作用,2007年3月被FDA批准用于治疗PNH,随后又被批准用于治疗非典型溶血性尿毒症综合征(aHUS)、抗乙酰胆碱受体(AchR)抗体阳性的难治性全身型重症肌无力(gMG)和抗水通道蛋白-4(AQP4)抗体阳性的成年患者的视神经脊髓炎症(NMOSD)。 • Ultomiris是Soliris的升级版,是第一种每8周给药一次的长效C5补体抑制剂,2018年12月被FDA批准用于治疗PNH,随后又被批准用于治疗aHUS、gMG和NMOSD。 • Empaveli是FDA批准的首个治疗PNH的补体C3靶向疗法,它是一种合成的环肽,与聚乙二醇聚合物偶联,特异性结合于C3和C3b。2021年5月,Empaveli 被FDA批准用于治疗未经治疗的成人PNH患者,以及既往接受过补体C5抑制剂Soliris (依库珠单抗)和Ultomiris (ravulizumab)治疗的PNH患者。 此外,目前全球还有多款在研PNH新药,详见下表。作用靶点上,除了C5,在研PNH新药靶点还涉及CFB、CFD等。药物类型上,除了单抗和环肽,还包括RNAi药物,如Cemdisiran。 研发进度上,罗氏的Crovalimab和诺华的Iptacopan进展较快。2022年8月Crovalimab治疗PNH的上市申请获CDE受理,且被纳入优先审评。Crovalimab是罗氏通过连续单克隆抗体回收技术(Smart-Ig)工程化改造自主研发的的新一代C5抑制剂,可以阻断补体C5裂解为C5a和C5b。 与已批准的C5抑制剂相比,Crovalimab高度可溶,提供持续完整的末端补体途径抑制,允许小剂量皮下注射(1-4 mL),可每月一次给药,同时作为皮下注射剂,患者可以在家自行给药,大大提升了依从性;其特异性靶向表位,能覆盖到更多临床情形的PNH患者,并表现出良好的耐受性和可接受的安全性。 此外,目前全球还有多款在研PNH新药进入后期临床,如Pozelimab、Nomacopan等。预计不久的将来,PNH领域将迎来更多治疗选择。 【企业推荐】智药研习社近期课程报名来源:CPHI制药在线声明:本文仅代表作者观点,并不代表制药在线立场。本网站内容仅出于传递更多信息之目的。如需转载,请务必注明文章来源和作者。投稿邮箱:Kelly.Xiao@imsinoexpo.com▼更多制药资讯,请关注CPHI制药在线▼点击阅读原文,进入智药研习社~
优先审批临床3期突破性疗法上市批准临床结果
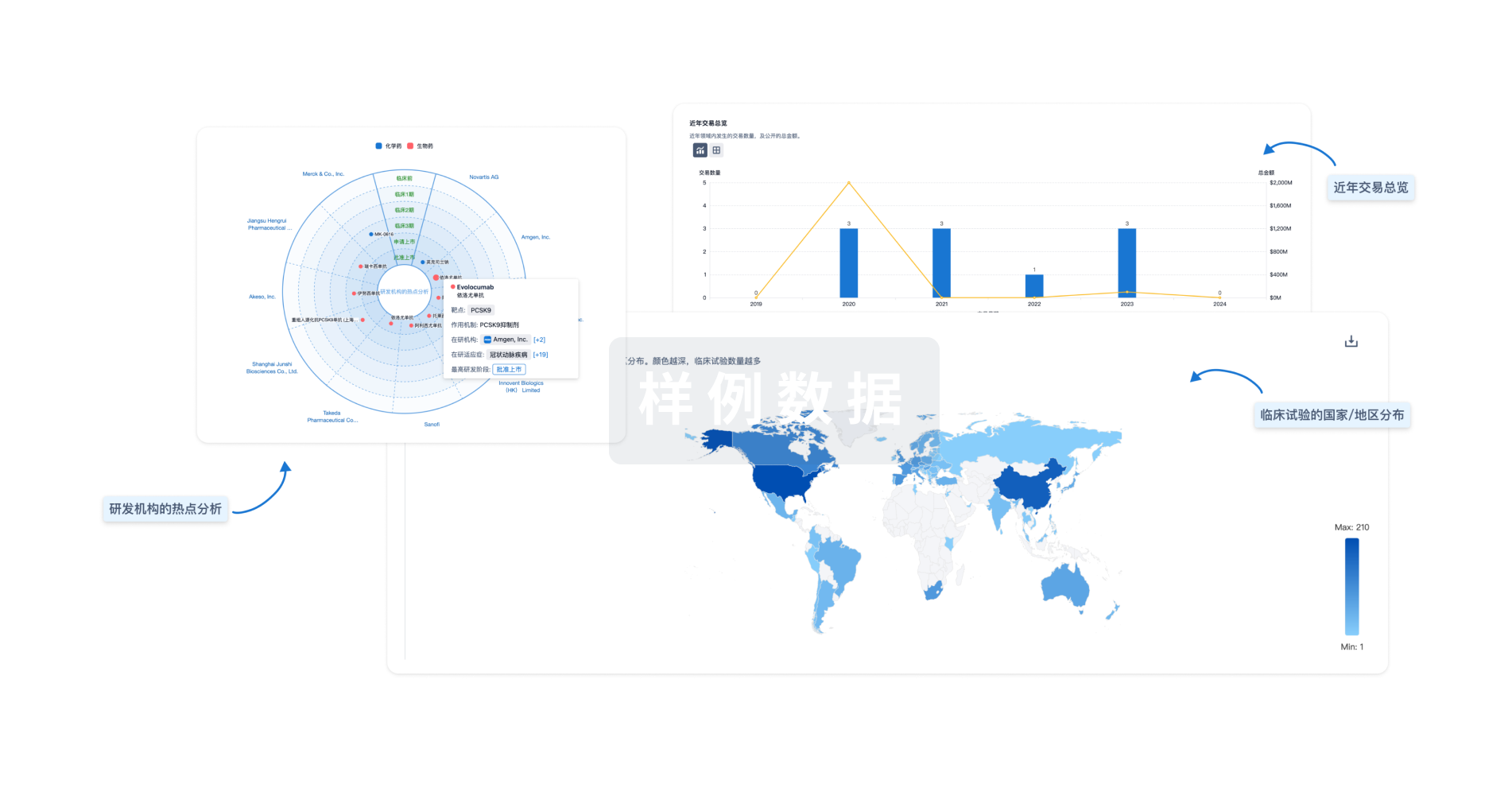
分析
对领域进行一次全面的分析。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
