预约演示
更新于:2025-05-07
PUUV glycoprotein
更新于:2025-05-07
基本信息
别名- |
简介- |
关联
1
项与 PUUV glycoprotein 相关的药物作用机制 PUUV glycoprotein 抑制剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期1800-01-20 |
100 项与 PUUV glycoprotein 相关的临床结果
登录后查看更多信息
100 项与 PUUV glycoprotein 相关的转化医学
登录后查看更多信息
0 项与 PUUV glycoprotein 相关的专利(医药)
登录后查看更多信息
2
项与 PUUV glycoprotein 相关的文献(医药)2023-10-01·The Journal of general virology
Validation of an antigenic site targeted by monoclonal antibodies against Puumala virus.
Article
作者: Iheozor-Ejiofor, Rommel Paneth ; Levanov, Lev ; Lundkvist, Åke ; Rissanen, Ilona ; Kedari, Ashwini ; Plyusnin, Alexander ; Vapalahti, Olli
PLOS Pathogens1区 · 医学
Crystal Structure of Glycoprotein C from a Hantavirus in the Post-fusion Conformation
1区 · 医学
ArticleOA
作者: Tischler, Nicole D ; Willensky, Shmuel ; Dessau, Moshe ; Bignon, Eduardo A ; Modis, Yorgo ; Bar-Rogovsky, Hagit
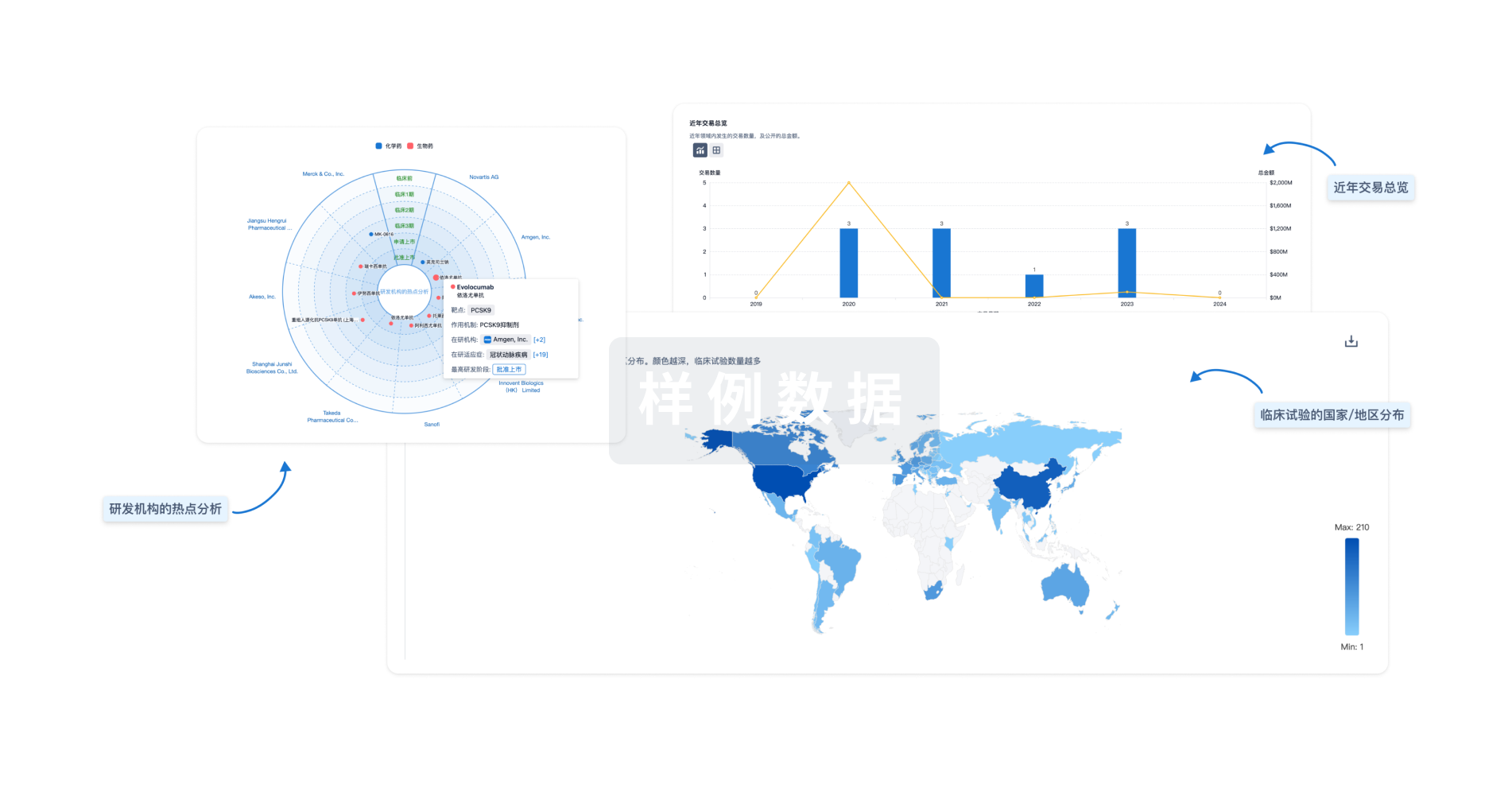
分析
对领域进行一次全面的分析。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
