预约演示
更新于:2025-05-07
calcium channel x ACE
更新于:2025-05-07
基本信息
关联
22
项与 calcium channel x ACE 相关的药物作用机制 ACE抑制剂 [+2] |
在研适应症 |
非在研适应症- |
最高研发阶段批准上市 |
首次获批国家/地区 新加坡 |
首次获批日期2016-05-23 |
作用机制 ACE抑制剂 [+1] |
在研适应症 |
非在研适应症- |
最高研发阶段批准上市 |
首次获批国家/地区 波兰 |
首次获批日期2016-03-09 |
作用机制 ACE抑制剂 [+1] |
在研适应症 |
非在研适应症- |
最高研发阶段批准上市 |
首次获批国家/地区 巴西 |
首次获批日期2012-04-21 |
115
项与 calcium channel x ACE 相关的临床试验CTR20250418
氨氯地平贝那普利胶囊(5 mg/10 mg)在健康受试者中餐后条件下的生物等效性试验
主要目的:在健康男性与女性受试者中于餐后条件下,评价受试制剂氨氯地平贝那普利胶囊【规格:每粒含苯磺酸氨氯地平5 mg(以氨氯地平计)和盐酸贝那普利10 mg,生产商:浙江华海药业股份有限公司】和Novartis Pharmaceuticals Corp为持证商的参比制剂氨氯地平贝那普利胶囊【Lotrel,规格:每粒含苯磺酸氨氯地平5 mg(以氨氯地平计)和盐酸贝那普利10 mg】的生物等效性。
次要目的:观察受试制剂氨氯地平贝那普利胶囊与参比制剂氨氯地平贝那普利胶囊(Lotrel)在健康受试者中的安全性。
开始日期2025-03-01 |
申办/合作机构 |
CTR20250421
氨氯地平贝那普利胶囊(5 mg/10 mg)在健康受试者中空腹条件下的生物等效性试验
主要目的:在健康男性与女性受试者中于空腹条件下,评价受试制剂氨氯地平贝那普利胶囊【规格:每粒含苯磺酸氨氯地平5 mg(以氨氯地平计)和盐酸贝那普利10 mg,生产商:浙江华海药业股份有限公司】和Novartis Pharmaceuticals Corp为持证商的参比制剂氨氯地平贝那普利胶囊【Lotrel,规格:每粒含苯磺酸氨氯地平5 mg(以氨氯地平计)和盐酸贝那普利10 mg】的生物等效性。
次要目的:观察受试制剂氨氯地平贝那普利胶囊与参比制剂氨氯地平贝那普利胶囊(Lotrel)在健康受试者中的安全性。
开始日期2025-02-19 |
申办/合作机构 |
CTR20250233
评估受试制剂培哚普利氨氯地平片(Ⅲ)(规格:10 mg:5 mg)与参比制剂开素达®(规格:10 mg:5 mg)在健康成年参与者空腹状态下的单中心、开放、随机、单剂量、交叉生物等效性研究
主要研究目的:研究空腹状态下单次口服受试制剂培哚普利氨氯地平片(Ⅲ)(规格:10 mg:5 mg,山东朗诺制药有限公司生产)与参比制剂培哚普利氨氯地平片(Ⅲ)(开素达®,规格:10 mg:5 mg,Servier (Ireland) Industries Ltd.生产)在健康成年参与者体内的药代动力学,评价空腹状态下口服两种制剂的生物等效性。
次要研究目的:评估受试制剂培哚普利氨氯地平片(Ⅲ)(规格:10 mg:5 mg)和参比制剂培哚普利氨氯地平片(Ⅲ)(开素达®,规格:10 mg:5 mg)在健康成年参与者中的安全性。
开始日期2025-02-08 |
申办/合作机构 |
100 项与 calcium channel x ACE 相关的临床结果
登录后查看更多信息
100 项与 calcium channel x ACE 相关的转化医学
登录后查看更多信息
0 项与 calcium channel x ACE 相关的专利(医药)
登录后查看更多信息
69
项与 calcium channel x ACE 相关的文献(医药)2025-05-01·Biochemical and Biophysical Research Communications
Discovering the natural source-derived antihypertensive compounds aspiring current therapeutic targets by computer-based drug design
Article
作者: Beker, Merve ; Sarikamis-Johnson, Bahar ; Ercin, Nilufer ; Besli, Nail ; Kalkan-Cakmak, Rabia ; Celik, Ulkan
2024-08-12·Journal of Economic Entomology
Frequencies of insecticide resistance mutations detected by the amplicon sequencing in Plutella xylostella (Lepidoptera: Plutellidae) and Spodoptera exigua (Lepidoptera: Noctuidae) from China
Article
作者: Liu, Zheming ; Zhu, Hang ; Liu, Jia ; Zhou, Xiaomao ; Li, Kaiqin ; Zhou, Yong ; Ma, Haihao ; Man, Yilong ; Liu, Zhangyang
2024-08-01·Pesticide Biochemistry and Physiology
Insecticide resistance and characteristics of mutations related to target site insensitivity of diamondback moths in Taiwan
Article
作者: Chen, Chien-Yu ; Dai, Shu-Mei ; Hsu, Ju-Chun ; Chen, Yu-Hsien ; Huang, Li-Hsin ; Chang, Chia-Che
3
项与 calcium channel x ACE 相关的新闻(医药)2025-02-22
据世界卫生组织(WHO)的统计数据显示,目前高血压和高血脂分别是血管性疾病死亡的第一、第二位危险因素,即高血压与胆固醇水平的升高,已然成为心血管疾病产生严重健康后果的关键推手。高血压与高血脂的综合管理是临床常见的管理策略。那么,具体如何实施呢?
医学界心血管频道邀请到了复旦大学附属华山医院心内科山缨教授就“高血压患者的血压血脂综合管理”这一专题作了精彩的演讲。
高血压与血脂异常往往并存
《中国心血管病报告2017》显示,无论是在农村还是城市,心血管死亡率皆为肿瘤死亡率的两倍,是导致死亡的首要原因。
在心血管疾病的死亡原因中,动脉粥样硬化相关的心血管疾病死亡一直占据重要地位,其死亡率和发病率在1990年到2015年期间呈现出快速上升的趋势,与此同时,出血性中风的发病率趋于平缓。然而目前中国的高血压患病率仍高达27.5%,即每四个成年人中就有一人患有高血压,且控制率不足17%。山缨教授指出,如何开展更为有效的血压控制,是我们需要重视的议题。
血脂异常在中国人群中的患病率也在显著上升。研究数据显示,从2002年到2012年,中国的血脂异常患病率从原有的18.6%已攀升至40.4%,翻了一倍之多。
大多数情况下,高血压和血脂异常是并存的。有数据显示,高血压患者中合并血脂异常的比例高达61.5%。此外还有1/4的高血压患者合并糖耐量异常,一半的患者存在肥胖问题,单纯以血压增高为危险因素的高血压患者仅占10%。山缨教授强调,我们对高血压患者的管理不能仅局限于血压的控制,还必须关注血脂的管理,以最大程度降低心血管疾病的发病和死亡风险。
减少发病率和死亡率,从把握心血管危险分层和积极降压做起
山缨教授在谈到高血压合并血脂异常的治疗策略时,强调了四个关键方面:积极降压、评估患者心血管危险分层、降压降脂的药物以及高血压患者降脂治疗要点。
1.积极降压,获得血压降低本身带来的心血管获益
一项涵盖61万人的双盲随机对照研究的结果显示,收缩压降低10 mmHg可使心血管风险降低约10%,若能降低20 mmHg,则心脑血管风险可降低20%。此外,收缩压每下降10 mmHg,全因死亡率能显著下降13%,且这种获益对于不同基线收缩压水平的患者以及已有心血管疾病和不伴心血管疾病的高血压患者都是一致的。因此,降压治疗对于高血压人群来说至关重要。
降压目标一般为130/80 mmHg,对于高龄老年患者可放宽至140/90 mmHg,虚弱的高血压患者则根据耐受性和预计生存情况设定目标。降压达标的速度要求是在3个月以内。降压方法主张联合用药,如血管紧张素转换酶抑制剂(ACEI)/血管紧张素Ⅱ受体阻滞剂(ARB)+钙通道阻滞剂(CCB),若仍不能控制,可加上利尿剂。国际高血压协会推荐的联合降压方案是A+C+D,在必要时可加用螺内酯。
2.评估心血管危险分层,确定高血压患者低密度脂蛋白胆固醇(LDL-C)靶目标,LDL-C控制达标是关健
除了降压治疗,对高血压患者还需评估其心血管危险因素并进行分层。不同高血压程度和胆固醇程度的患者,心血管死亡率存在显著差异。收缩压在160 mmHg以上且总胆固醇在240 mg/dl以上的人,与收缩压130 mmHg以下且总胆固醇200 mg/dl以下的人相比,心血管死亡风险增加17.6倍。
遗传流行病学研究也表明,基因导致LDL-C长期低水平的人群,心血管风险显著低于通过药物降脂的人群。LDL-C水平和收缩压的绝对值小幅变化,都能带来心血管风险的差异,且暴露的累积时间也是心血管风险的危险因素,早期干预对心血管事件的获益极大。
CTT研究显示,不同收缩压基线水平下,LDL-C每降低1 mmol/L,心血管风险可降低约20%。ASCOT-LLA研究将高血压合并3个心血管危险因素的患者随机分入降脂组和安慰剂组,结果显示降压联合降脂组在3.3年时心血管风险降低30%以上。
3.他汀类药物是LDL-C达标的药物治疗基石
山缨教授提到在ASCOT-LLA研究提前终止后,研究者将安慰剂组的患者也纳入降压联合降脂治疗组,继续随访10年。结果发现那些在前3年就开始接受阿托伐他汀联合降压治疗的患者,与从第4年开始接受该治疗方案的患者相比,全因死亡率低14%。这提示我们早期启动降压联合降脂治疗(尤其是他汀类药物)能够带来心血管保护作用,这种获益不仅在短期内显现,而且能够持续多年。
4.单片复方制剂在降压、降脂、降压联合降脂治疗中的价值
单片复方制剂(polypill)是一种包含多种药物成分的复合制剂,用于同时实现降压和降脂的目标。TIPS-3研究是一项针对无心血管疾病但有中高心血管风险的患者,给予含有降压和降脂成分的polypill的随机对照试验。研究结果显示使用polypill的患者心血管事件风险下降了21%。这提示即使在尚未发生心血管疾病的高危人群中,这种综合管理策略也能显著降低心血管事件的风险。
仅在转氨酶达到这个值,才考虑停他汀类药物
根据《中国心血管疾病防治指南》,高血压患者的心血管风险分为低危、中危、高危、极高危和超高危。
他汀类药物是中国降脂治疗的基石,常规剂量可使LDL-C降低30%~50%。中国他汀药物可及性高。2023年指南提出谨慎判断他汀不耐受,仅在转氨酶增高3倍以上或激酶增高4倍以上时才考虑减量或停药。
2019年美国心脏协会(AHA)数据显示,每1万例患者接受他汀治疗5年,LDL-C降低2 mmol/L,可减少1000例二级预防患者事件,预防500例一级预防患者事件。严重肝病发生率极低,新增糖尿病100例,但血脂管理对血糖异常患者更重要,获益远高于风险。除他汀外,还有胆固醇吸收抑制剂、PCSK9抑制剂和血脂康胶囊等降脂药物。
小结
最后,山缨教授总结道,高血压患者需每年至少检查一次血脂,追求降压和降脂双重达标;无论是否采用药物治疗,均应重视生活方式干预,制定个体化指导建议;刚启动降脂药物治疗的患者,或调整治疗剂量的患者,建议4~6周复查血脂;所有动脉粥样硬化性心血管疾病(ASCVD)高危、极高危和超高危的高血压患者,均须立即同时启动降压联合降LDL-C药物治疗,及早实现血压和血脂双达标;降压和降脂治疗需长期坚持,并维持达标,最终降低心血管事件。
专家简介
相关阅读
《柳叶刀》:仅需20分钟微创消融,有望同时治愈高血压和原酮症!
不吃降压药,微创手术也能强效降压!符合这些条件的高血压患者,更能从中受益
安贞医院周玉杰领衔:强效降压25.2 mmHg!微创手术有望为中国高血压患者带来新选择
JAHA:强化降压达到<140 mmHg,高血压患者就无需太担心中风风险
ESC近5万人研究:高血压患者什么时候吃降压药更好?早上和晚上吃有区别吗?
欢迎投稿:学术成果、前沿进展、临床干货等主题均可,点此了解投稿详情。
免责声明:药明康德内容团队专注介绍全球生物医药健康研究进展。本文仅作信息交流之目的,文中观点不代表药明康德立场,亦不代表药明康德支持或反对文中观点。本文也不是治疗方案推荐。如需获得治疗方案指导,请前往正规医院就诊。
分享,点赞,在看,传递医学新知
AHA会议临床结果
2025-02-17
2025年世界肾脏病大会(WCN 2025)于2025年2月6日至9日在印度新德里举行。大会期间,热点辩论环节尤为引人关注,其中Swapnil Hiremath教授就与Indranil Dasgupta教授就“改善全球肾脏病预后组织(KDIGO)慢性肾脏病患者血压控制目标的现实可达性”这一主题展开了辩论。Swapnil Hiremath教授认为KDIGO慢性肾脏病患者血压控制目标是可及的,小编整理了Swapnil Hiremath教授的论述内容,一起来看看吧~
关于降压目标的设定
全球疾病负担数据显示,高血压与众多不良健康结局密切相关,尤其是心脏病和中风。普遍观点认为极高血压值(例如180 mmHg或160 mmHg)才构成威胁,但事实上,风险自血压达到120 mmHg时便已开始悄然上升。正因如此,多数心血管事件的发生往往集中在血压介于120~160 mmHg区间的人群中。因此,单纯因设定140 mmHg或160 mmHg等较为容易达成的血压目标值而妥协,并非科学的应对策略。
这并非仅凭一个研究得出的结论,一项涵盖了多项大型试验的综合荟萃分析研究显示,血压每降低约10 mmHg,主要不良心血管事件(MACE)的风险就能显著降低约20%。血压与中风及心力衰竭的发生紧密相关,其适度下降可使中风和心力衰竭风险降低约30%,并使死亡风险降低约10%。
治疗高血压的首要目的并非延缓慢性肾脏病(CKD)的进展,因为其对肾衰竭的直接影响相对有限。实际上,控制高血压主要是为了降低心脏病和中风的风险。或许有人认为,这应该是心脏病学家的职责范畴,而非肾病学家的任务。然而,这种观点有失偏颇,因为大多数CKD患者的致死原因并非肾衰竭或透析,而是心脏病。对于肾小球滤过率(GFR)低于60 ml/min/1.73m2的患者而言,心血管疾病更是首要死因。因此,若期望CKD患者拥有更长的生存期,就必须妥善管理其心血管风险因素,而控制血压无疑是其中的关键一环。
那么,为何将120 mmHg作为血压控制的目标值呢?如前所述,血压每降低10 mmHg,相关风险便会降低。这一结论主要基于SPRINT试验的结果。在SPRINT试验中,总体数据显示,不仅MACE的风险降低了25%,全因死亡率也下降了约27%。
深入探究SPRINT试验中的CKD亚组数据,我们发现,尽管该亚组的入组人数未达到预期,但仍能清晰展现出全因死亡率的持续下降趋势。在CKD患者中,全因死亡率降低了28%,心血管风险降低了43%。尽管样本量有限,但数据显示在CKD群体中,风险降低的幅度依然十分明显。
既然如此,为何有人持反对观点?这或许是对老年患者的年龄顾虑。然而,当我们观察针对老年群体的数据,会发现其效果甚至更为显著。考虑到老年患者中风风险本就偏高,通过合理降压,可以帮助他们延长寿命。因此,年龄不应成为治疗的阻碍。
既往指南将血压控制目标设定为140/90 mmHg,但SPRINT试验的发布突然提出了一个更为严格的目标——降至120 mmHg。这一挑战性的新标准自然引起了广泛争议,我认为这些声音应当被理性对待。
SPRINT试验是一项设计严密、执行严谨的研究,其结果支持了120 mmHg作为更优的血压控制目标。此外,中国的ESPRIT试验(纳入患者数超过SPRINT,达11000名)也采用了相似策略,研究结果显示,强化降压组和标准降压组的主要终点事件发生分别为547例(9.7%)和623例(11.1%),证实了将血压控制在120 mmHg以下可降低MACE风险。
另外,近期发表的BPROAD试验(针对12000例糖尿病患者)同样显示,与140 mmHg相比,将血压控制在120 mmHg能降低MACE风险达21%。因此,设定更低的血压目标值(120 mmHg)对保护CKD患者的心脏,并降低死亡风险是有益的。
关于血压测量的问题
在血压测量方面,我们的表现确实不尽如人意。一项针对159名医学生的调研揭示,仅有一人能够准确执行血压测量操作。
测量血压前有诸多注意事项,如避免饮用咖啡或茶、不吸烟以及保持膀胱排空等。膀胱充盈状态可能使血压升高约30 mmHg,因此确保患者在测量前排空膀胱至关重要。此外,正确的体位、快速放气等技巧虽已被倡导多年,但实施效果并不理想。
在此背景下,自动血压测量设备显得尤为重要。尽管这些设备也曾备受争议,但它们在实际应用中确实展现出了有效性。例如,已停产的BP TRUE设备通过取五次测量的平均值,测量血压从147 mmHg降低至136 mmHg。
另一项的研究也证实了这一点,使用与SPRINT试验类似的设备,测量血压降低了12.7 mmHg。这些研究强调了正确测量血压的重要性,以及采用自动化设备在提升测量准确性和效率方面的潜力。
您或许会提出疑问,是否可以直接将手动测量值扣除13 mmHg作为标准血压?答案是否定的,因为误差的范围可能介于-46~+20之间。
一项发表在《内科学年鉴》杂志的研究探讨了医护人员在场与否对血压测量结果的影响,结果表明,即便有他人在场,血压测量结果亦无显著差异。
研究团队在巴尔的摩一个历史悠久的市场中进行了测量,尽管环境喧嚣,但通过实施5分钟的静息时间及采用正确技术,发现嘈杂环境与安静环境下的测量结果并无差异。这意味着,在候诊室内测量血压同样可行,无需因环境而担忧测量准确性。若未能准确测量血压,则不应轻易启动高血压治疗方案。上述试验中采用的测量手段(无论是自动化设备还是手动操作)均符合规范。
血压控制的关键在于精确测量,正确的测量能让看似不可能达成的降压目标变得可及。
关于药物治疗
一项关于不依从性和难治性高血压的系统性综述研究显示,高血压患者的不依从性发生率高达约35%,而若采用直接观察疗法(DOT)或尿液检测,不依从的比例可能攀升至近半数。这一议题与Dasgupta医生的关注点紧密相关,相信他会认同患者的不依从性是一个不可忽视的关键因素。因此,当患者在增加药物剂量后血压仍居高不下时,请务必考虑不依从性的可能性。
合理的药物使用可以用ACDc来记忆:A[血管紧张素转换酶抑制剂(ACEI)/血管紧张素II受体拮抗剂(ARB)]、C(钙通道阻滞剂)、D(利尿剂)。
有一种误解认为噻嗪类利尿剂在CKD患者中无效。CLICK研究显示,即使在CKD患者中,使用低剂量的氯噻酮(6.25 mg或12.5 mg,隔日一次)也可以将血压降低约11 mmHg。当然,应用的同时我们也要注意避免不良反应。
肌酐仅仅是一个数值,其临床意义需结合具体情况分析。当为患者采取治疗措施后,尽管肌酐值有所上升,但患者的临床状况得到改善,这种情况是可以接受的。我理解这并不容易,因为患者也在密切关注这个数值。
例如,在治疗心力衰竭患者时,使用利尿剂后,肌酐值的上升往往是预期之内且积极的信号。然而,如果是由于其他原因(如败血症或肾毒性药物导致的急性肾损伤)引起的自发性肌酐上升,则需要另作考虑。我并非建议忽视急性肾损伤(AKI),但如果是为了治疗目的而诱导产生的AKI,许多研究表明,相关生物标志物并不会显著上升,此时肌酐数值的波动不必过分担忧。
Don't STOP-ACEi研究讨论了是否应在晚期CKD患者中停用ACEi和ARB。结果显示,如果停用这些药物,肌酐值会下降,但这并不会延缓肾病的进展。继续使用ACE抑制剂和ARB的患者,MACE风险降低了9%(尽管没有统计学意义)。
另一项发表在《内科学年鉴》的研究讨论了在晚期CKD患者中使用ACEi和ARB的效果。结果显示,继续使用这些药物可以降低肾衰竭的风险。我想强调的是,对于蛋白尿阳性CKD患者、糖尿病CKD患者或心力衰竭患者,不要随意停用ACEi和ARB。患者有明确的适应证,可以继续使用这些药物。
最后,关于α受体阻滞剂,从医学角度来看,α受体阻滞剂并未给患者带来实质性的益处,除了可能降低某些数值和提供短暂的心理安慰。众多研究已证实了这一点,尽管它们可能使肌酐值看似正常,且不会导致低钾血症,但实质上并未改善患者的健康状况。
因此,我们应选择更合适的药物组合。例如,对于糖尿病患者、心力衰竭患者或有蛋白尿适应证的患者,SGLT2抑制剂是不错的选择。在动态血压监测(ABPM)下,SGLT2抑制剂也能有效降低血压约4~9 mmHg。
此外,胰高血糖素样肽-1(GLP-1)受体激动剂也在许多地方逐渐得到认可。对于肥胖患者或CKD患者,GLP-1受体激动剂能够减轻体重并降低血压,因此值得考虑使用。
相关阅读
《自然-医学》:降糖、减重、降压,还能治肾病!司美格鲁肽“一药多效”再获力证
2024年指南:肾病9大食养建议,这样吃喝…更合理
显著改善肾功能,创新肾病疗法2期临床结果登上NEJM
欢迎投稿:学术成果、前沿进展、临床干货等主题均可,点此了解投稿详情。
免责声明:药明康德内容团队专注介绍全球生物医药健康研究进展。本文仅作信息交流之目的,文中观点不代表药明康德立场,亦不代表药明康德支持或反对文中观点。本文也不是治疗方案推荐。如需获得治疗方案指导,请前往正规医院就诊。
分享,点赞,在看,传递医学新知
临床终止临床研究
2024-03-22
关注并星标CPHI制药在线 3月19日,瑞士制药公司Idorsia开发的创新药物Aprocitentan(阿普昔腾坦片,商品名Tryvio)正式获FDA批准上市,用于治疗难治性高血压患者。这一里程碑式的进展,标志着医学界在高血压治疗领域取得了重大突破。 针对难治性高血压,30年来第一个新机制降压药 高血压是心脑血管疾病相关死亡的重要危险因素。目前常用的降压药有6大类,分别是:血管紧张素转换酶抑制剂(ACEI)、血管紧张素受体拮抗剂(ARB)、钙通道阻滞剂(CCB)、噻嗪类/噻嗪样利尿剂、β受体阻滞剂(BB)、血管紧张素受体脑啡肽酶抑制剂(ARNI),分别有特定的作用机制。 尽管这些药物在降压方面疗效显著,但仍有部分成人高血压患者控压效果不理想,这部分患者即为难治性高血压。患有难治性高血压的患者,靶器官损害风险增加、预后差,需要更加重视。 Aprocitentan是一款新型口服双重内皮素A/B受体(ETA/ETB)拮抗剂,由Idorsia公司开发。Aprocitentan可有效抑制内皮素-1(ET-1)与ETA和ETB的结合。Aprocitentan是马昔腾坦的活性代谢产物,但具有更长的半衰期(48h vs 14h)。 FDA此次批准Aprocitentan上市主要基于III期PRECISION研究。该研究是一项多中心、盲法、随机III期临床试验,主要和关键次要终点分别是收缩压从基线到第4周和第40周的变化, 24 h动态血压变化等。 研究分为4个部分连续进行:第1部分是为期4周的双盲期,其中704 例患者按 1:1:1 的比例随机接受 Aprocitentan 12.5 mg QD(1天1次)、Aprocitentan 25 mg QD 和安慰剂治疗; 第2部分是为期32周(第4~第36周)的单盲研究,所有患者接受 Aprocitentan 25 mg QD 治疗; 第3部分是为期12周(第36~第48周)的随机、双盲、安慰剂控制撤退期,患者按1:1重新随机分配至 Aprocitentan 25 mg QD 或安慰剂治疗; 第 4 部分是所有患者停用Aprocitentan观察4周。 结果显示,第4周时,接受Aprocitentan 12.5 mg、Aprocitentan 25 mg和安慰剂治疗的坐位收缩压(SiSBP)相比,降低幅度显著大于安慰剂组。具体而言,相较于安慰剂组,Aprocitentan 12.5 mg 和 Aprocitentan 25 mg 两组SiSBP分别下降 3.8 mmHg(P = 0.0042)和 3.7 mmHg(P = 0.0046)。 在第36~40周期间,接受Aprocitentan治疗的患者的SiSBP较安慰剂组持续降低,差异为-5.8mmHg(p<0.0001),维持时间长达48周。 研究中最常见的不良反应为轻中度水肿或液体潴留,接受 Aprocitentan 12.5 mg、25 mg 和安慰剂的患者中发生率分别为 9.1%、18.4% 和 2.1%。 结果表明,在难治性高血压患者中,Aprocitentan耐受性良好,在4 周内的降压效果优于安慰剂,而且降压效果在持续服用40周期间保持稳定。 在递交Aprocitentan上市申请时,Idorsia的CEO表示,高血压领域已经30多年来无创新机制产品上市,而Aprocitentan将是一种以全新机制治疗难治性高血压的药物。 另外,Aprocitentan除具潜力抑制内皮素通路外,其与其他药物产生药物间相互作用的机率低,这些特征都让Aprocitentan有潜力成为治疗顽固性高血压患者的药物。 ET受体拮抗剂研发现状 内皮素(ET)是一种强效肽类血管收缩剂,1988年由日本科学家Yanagisawa及其同事发现。ET由血管内皮细胞分泌,家族中共有3个成员:ET-1、ET-2和ET-3。这些ET各长21个氨基酸,链内有两个二硫键限制其整体结构。 人体共有两种类型的ET受体:ETA 型受体(ETA)与 ET-1 和 ET-2 的结合亲和力高于与 ET-3 的结合亲和力;ETB 型受体(ETB)与这三种异肽的结合亲和力相当。其中ET-1及其受体ETA作为病原因素和诱发因子在心血管疾病中被广泛研究,具有强效收缩血管作用。ETA主要分布于平滑肌细胞,介导血管平滑肌细胞增殖、纤维化、炎症反应、血管收缩等效应,与心血管病的发生密切相关。选择性阻断ETA能够阻断由内皮素引起的血管收缩,发挥降压作用。 目前,多家公司正在布局ETA或ETB受体选择性拮抗剂或双重拮抗剂。其中以ETA拮抗剂最为广泛,适应症涵盖肾病、心血管疾病等。 阿斯利康的zibotentan,这是一款小分子ETA选择性拮抗剂,拟与达格列净组成联合疗法,用于治疗慢性肾脏病和肝硬化。 诺华收购的ETA小分子拮抗剂atrasentan目前处于III期临床阶段,拟用于治疗IgA肾病和蛋白尿性肾小球疾病。诺华计划今年向FDA提交申请,以期atrasentan在美国加速批准。 在国内,智康弘义研发的ETA小分子拮抗剂SC0062,正在开展针对IgA肾病和糖尿病肾脏病的II期临床研究。在先前I期健康志愿者的临床研究结果表明,SC0062在展现出了良好的安全性、药代动力学和药效学特征。 信立泰研发的小分子ETA拮抗剂已获批临床,适应症为轻、中度原发性高血压。 目前,顽固性高血压是典型的未满足临床需求,在全球超过10亿高血压患者中,顽固性高血压患者的发病率为10-12%。他们尽管接受了至少3种以上不同种类的抗血压药物联用,血压仍然得不到控制。长期不受控的高血压可能对心脏与血管造成伤害,进而增加患者发生心脏病、肾衰竭、血管性痴呆与脑卒中的风险。 此次Aprocitentan的获批上市,为这类顽固性高血压患者提供了全新机制的降压药物。 参考来源: 1. https://www.msdmanuals.cn/home/heart-and-blood-vessel-disorders/high-blood-pressure/high-blood-pressure 2. https://ir.travere.com/news-releases/news-release-details/retrophin-announces-corporate-name-change-travere-therapeutics 3.Vignon-Zellweger N,Heiden S,Miyauchi T,et al.Endothelin and endothelin receptors in the renal and cardiovascular systems[J].Life sciences,2012,91(13-14):490-500.【智药研习社近期课程预告】来源:CPHI制药在线声明:本文仅代表作者观点,并不代表制药在线立场。本网站内容仅出于传递更多信息之目的。如需转载,请务必注明文章来源和作者。投稿邮箱:Kelly.Xiao@imsinoexpo.com▼更多制药资讯,请关注CPHI制药在线▼点击阅读原文,进入智药研习社~
临床结果上市批准临床3期临床成功
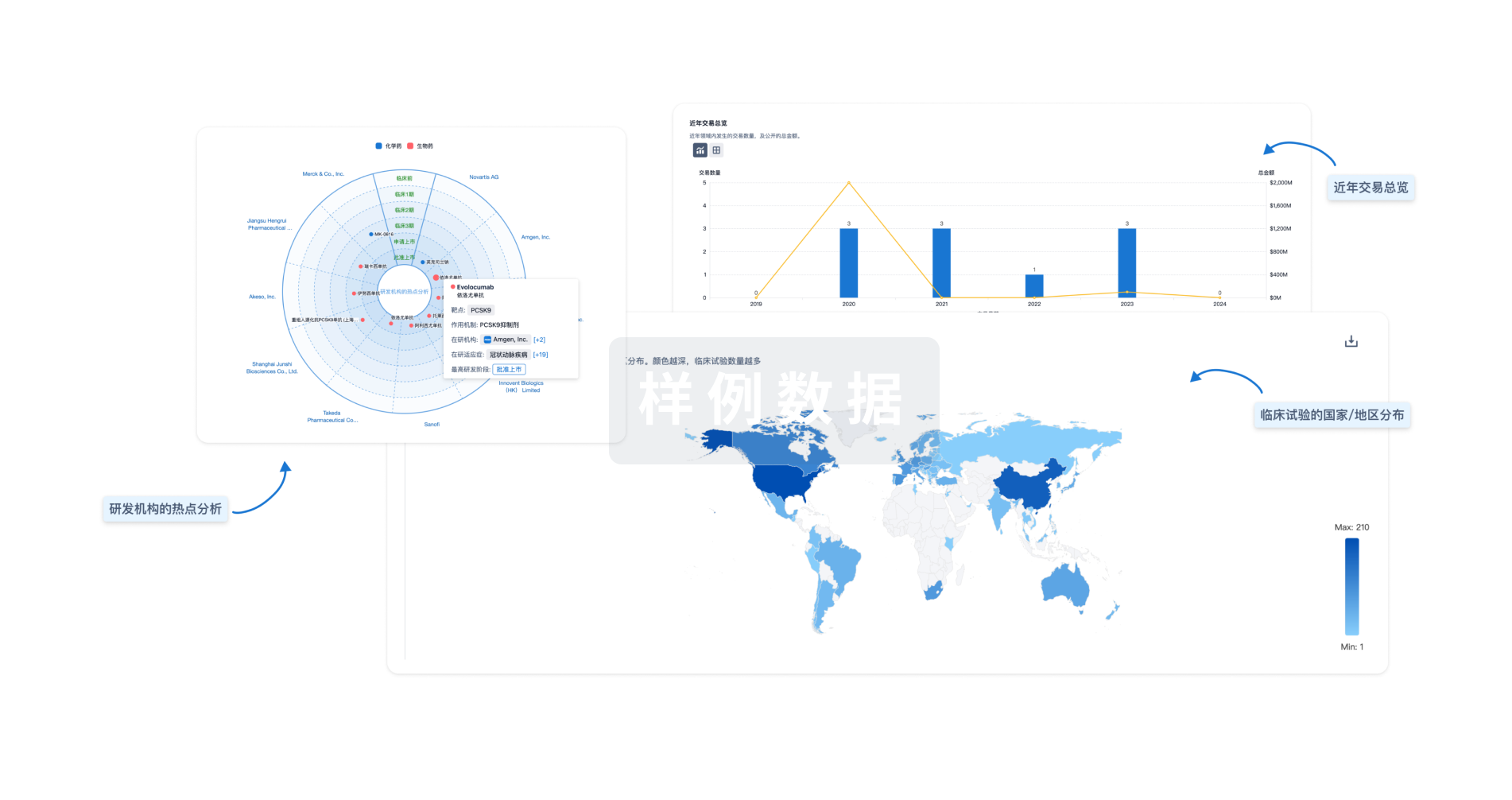
分析
对领域进行一次全面的分析。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用



