预约演示
更新于:2026-02-25
Cysteine Hydrochloride
盐酸半胱氨酸
更新于:2026-02-25
概要
基本信息
原研机构 |
最高研发阶段批准上市 |
首次获批日期 中国 (1982-01-01), |
最高研发阶段(中国)批准上市 |
特殊审评快速通道 (美国) |
登录后查看时间轴
结构/序列
分子式C3H10ClNO3S |
InChIKeyQIJRTFXNRTXDIP-JIZZDEOASA-N |
CAS号7048-04-6 |
关联
19
项与 盐酸半胱氨酸 相关的临床试验NCT07293884
Efficacy of Nebulized Versus Intravenous N- Acetylcysteine in Airway Clearance in Broncheictasis
It has been deduced that reducing the production of mucus or improving the clearance of sputum in the airway is the key to enhance the therapeutic efficacy for bronchiectasis . Mucoactive drugs are commonly used to clear the airway in mucus hypersecretion diseases. Currently, investigators have found that N-acetylcysteine (NAC), an effective mucolytic agent, not only reduces the viscosity and elasticity of sputum, but it also has anti-inflammatory and antioxidant activity.
Additionally, the Spanish guidelines on the treatment of bronchiectasis indicate that the use of N-acetylcysteine should be considered for patients with bronchiectasis and COPD. Therefore, as a classic mucolytic agent with antioxidant and anti-inflammatory properties, N-acetylcysteine can be effective in the treatment of bronchiectasis
Additionally, the Spanish guidelines on the treatment of bronchiectasis indicate that the use of N-acetylcysteine should be considered for patients with bronchiectasis and COPD. Therefore, as a classic mucolytic agent with antioxidant and anti-inflammatory properties, N-acetylcysteine can be effective in the treatment of bronchiectasis
开始日期2025-12-01 |
申办/合作机构 |
NCT06622577
The Effect of Dietary Management and Cysteine Supplementation on Growth Parameters and Biochemical Control for Pediatric Qatari Patients Affected with Classical B6 Non-responsive Homocystinuria.
Classical homocystinuria (HCU) is an autosomal recessive disorder caused by the deficiency of an enzyme cystathionine β-synthase (CβS) that affects the catabolic pathway of the amino acid methionine (Met) which leads to an accumulation of high levels of methionine and Homocysteine causing complications in the multi system. Therefore, a strict dietary management is crucial to maintain good biochemical control, growth parameters and avoid complications. The main objective would be to analyze the impact of the Met restricted diet on growth parameters, biochemical markers and long-term complications in patients up to 18 years. In addition, the efficacy of dietary management with additional cysteine (Cys) supplementation for the patients up to 18 years would also be examined.
The participants of the study would be recruited from the metabolic and genetics clinic in Hamad General Hospital (HGH), Qatar. All Qatari participants with confirmed diagnosis of HCU <18 years of age will be included in the study. A mixed method study design would be used which include a cross sectional study design to assess the impact of methionine restricted diet on outcome variables and a single arm interventional study to analyze the effect of additional cysteine supplementation in patients from birth to 18 years. For the retrospective study, all the required data would be retrieved from electronic record from the Cerner of HMC and would be stored in a local drive with password protection. Further, all the eligible participants would be prospectively followed to supplement additional cysteine for the period of 6 months.
The collected data will be statistically analyzed using the "SPSS windows version 22.0 software. The study would improve better understanding of dietary management through the identified outcomes. The outcome of Cys supplementation will improve the protein tolerance, biochemical parameters, growth parameters and may standardize the Cys supplementation.
The participants of the study would be recruited from the metabolic and genetics clinic in Hamad General Hospital (HGH), Qatar. All Qatari participants with confirmed diagnosis of HCU <18 years of age will be included in the study. A mixed method study design would be used which include a cross sectional study design to assess the impact of methionine restricted diet on outcome variables and a single arm interventional study to analyze the effect of additional cysteine supplementation in patients from birth to 18 years. For the retrospective study, all the required data would be retrieved from electronic record from the Cerner of HMC and would be stored in a local drive with password protection. Further, all the eligible participants would be prospectively followed to supplement additional cysteine for the period of 6 months.
The collected data will be statistically analyzed using the "SPSS windows version 22.0 software. The study would improve better understanding of dietary management through the identified outcomes. The outcome of Cys supplementation will improve the protein tolerance, biochemical parameters, growth parameters and may standardize the Cys supplementation.
开始日期2024-12-01 |
申办/合作机构 |
NCT07184398
Use of a Cysteine-rich Whey Protein Isolate (Immunocal®) in Post COVID-19 Cognitive Impairment
Abstract Background: Post-COVID-19 cognitive impairment (PCCI), characterized by deficits in attention, memory, and executive functioning, remains a major challenge among patients with long COVID. Oxidative stress is a key contributor to this condition. Cysteine-rich whey protein isolate (CRWPI), such as Immunocal®, enhances intracellular glutathione production and may offer neuroprotective benefits.
Objective: To evaluate the efficacy of Immunocal® supplementation on cognitive function, particularly attention and memory, and functional performance in individuals with ICCP.
Methods: A randomized, controlled, parallel-group trial was conducted in Cali, Colombia, with 120 adults recovering from COVID-19 with mild to moderate cognitive impairment. Participants were randomly assigned to three groups: (1) immune® supplementation (CRWPI) (20 g/day), (2) neuropsychological rehabilitation, or (3) no intervention (control), for 12 weeks. Cognitive outcomes were assessed using the Montreal Cognitive Assessment (MoCA) and the NEUROPSI attention and memory test. Physical endurance was measured using the 30-second foot test (STST). Clinical symptoms evaluated through a medical assessment form, before and after the intervention, were also taken into account. Results: Both the Immunocal® and neurorehabilitation groups showed statistically significant improvements in all attention subdomains and memory types compared to the control. Immunocal® produced greater gains in divided attention and working memory, suggesting a specific advantage in cognitive domains sensitive to oxidative stress. STST performance was also significantly improved in the Immunocal® group. No significant improvements were seen in the control group.
Conclusion: Immunocal® supplementation significantly improves cognitive performance, comparable to structured neurorehabilitation, in individuals with ICCP. It also shows potential to improve physical endurance, clinical symptoms, and reduce fatigue. These findings support the integration of Immunocal® as a non-pharmacological intervention for cognitive dysfunction related to long COVID.
Objective: To evaluate the efficacy of Immunocal® supplementation on cognitive function, particularly attention and memory, and functional performance in individuals with ICCP.
Methods: A randomized, controlled, parallel-group trial was conducted in Cali, Colombia, with 120 adults recovering from COVID-19 with mild to moderate cognitive impairment. Participants were randomly assigned to three groups: (1) immune® supplementation (CRWPI) (20 g/day), (2) neuropsychological rehabilitation, or (3) no intervention (control), for 12 weeks. Cognitive outcomes were assessed using the Montreal Cognitive Assessment (MoCA) and the NEUROPSI attention and memory test. Physical endurance was measured using the 30-second foot test (STST). Clinical symptoms evaluated through a medical assessment form, before and after the intervention, were also taken into account. Results: Both the Immunocal® and neurorehabilitation groups showed statistically significant improvements in all attention subdomains and memory types compared to the control. Immunocal® produced greater gains in divided attention and working memory, suggesting a specific advantage in cognitive domains sensitive to oxidative stress. STST performance was also significantly improved in the Immunocal® group. No significant improvements were seen in the control group.
Conclusion: Immunocal® supplementation significantly improves cognitive performance, comparable to structured neurorehabilitation, in individuals with ICCP. It also shows potential to improve physical endurance, clinical symptoms, and reduce fatigue. These findings support the integration of Immunocal® as a non-pharmacological intervention for cognitive dysfunction related to long COVID.
开始日期2024-07-01 |
申办/合作机构 |
100 项与 盐酸半胱氨酸 相关的临床结果
登录后查看更多信息
100 项与 盐酸半胱氨酸 相关的转化医学
登录后查看更多信息
100 项与 盐酸半胱氨酸 相关的专利(医药)
登录后查看更多信息
70,744
项与 盐酸半胱氨酸 相关的文献(医药)2026-06-01·Bioactive Materials
Age-related GSS promoter methylation in BMSCs drives osteoporosis and the reversal by targeted GSH delivery
Article
作者: Li, Pan ; Guo, Shuo ; Qu, Mi ; Su, Kangkang ; Liu, Wenwen ; Zeng, Xianyan ; Yu, Dechen ; Luo, Zhuojing ; Liang, Zhuowen ; Zhang, Guangwei ; Zhang, Zhao ; Lei, Runbo ; Qin, Anhui ; Li, Jianxiong
Age-related osteoporosis arises from bone tissue with inadequate metabolic support for osteogenesis. We identified that DNA methylation-mediated suppression of glutathione synthetase (GSS) represents an upstream lesion limiting endogenous glutathione (GSH) synthesis and supply in aged bone, thereby constraining osteoblast differentiation. In turn, impaired GSH synthesis exacerbates oxidative stress levels and diminishes osteogenic capacity, and this metabolic bottleneck is independent of substrate availability: cysteine supplementation neither restored GSH synthesis flux in aged bone nor rescued its osteogenic deficits. To overcome this limitation, we developed an exosome-based GSH delivery platform using electroporation to efficiently load GSH. These exosomes are derived from CXCR4-enriched bone marrow mesenchymal stem cells (BMSCs), leveraging CXCR4-mediated homing to the bone marrow niche to enhance bone retention, stabilize GSH during loading and circulation, and elevate local GSH pools at osteogenic sites. In aged bone, this targeted system sustainably delivers GSH, alleviates oxidative stress, improves mitochondrial function, delays cellular senescence, and promotes osteogenesis. In summary, while DNA methylation acts upstream to constrain GSH synthesis in aging bone, therapeutically correcting the resultant metabolic deficit via bone-homing exosome-mediated GSH delivery restores osteogenic function and improves bone metabolism in aged individuals.
2026-05-01·SPECTROCHIMICA ACTA PART A-MOLECULAR AND BIOMOLECULAR SPECTROSCOPY
Scalable copper nanodendrites for next-generation SERS-based biosensors
Article
作者: Soma, Venugopal Rao ; Vendamani, V S ; Das, Sathi ; Yella, Pardhu
Scalable, complex geometry of copper nanodendrites (Cu NDs) is highly desirable for next-generation SERS-based biosensors because of the ease of preparation, strong plasmonic response, and lowcost. Highly branched copper nanodendrites (Cu NDs) are fabricated by a non-equilibrium galvanic replacement reaction (GRR) in this work. Remarkably, these Cu NDs create intense field-efficient spots at sharp tips and branch junctions, amplifying bimolecular vibrational signatures leading to superior detection at the trace levels. X-ray diffraction (XRD) and selective area electron diffraction (SAED) analyses indicate preferential growth of Cu NDs in the (111) direction. The versatility of the prepared Cu NDs is demonstrated by their ability to detect multiple biomolecules such as cytosine, adenine, Bovine serum albumin (BSA), and L-cysteine, which are promising biomarkers for early diagnosis of tumors. The sensitivity was evaluated using the analytical enhancement factor (AEF), yielding values of 108 for crystal violet and 104-105 for biomolecules. The estimated limit of detection (LOD) was 138 pM for CV, 27 nM for cytosine, 54 nM for adenine, 64 nM for BSA, and 380 nM for L-cysteine. The signal reproducibility at different locations of the sample surface was demonstrated by a low relative standard deviation (RSD) (∼3%). These results suggest that Cu NDs are robust, versatile, cost-effective, and scalable for future biosensor applications.
2026-05-01·SPECTROCHIMICA ACTA PART A-MOLECULAR AND BIOMOLECULAR SPECTROSCOPY
Luminescent sensor based on colloidal Ag
2
S quantum dots for oxytetracycline detection in milk
Article
作者: Ovchinnikov, O V ; Grevtseva, I G ; Smirnov, M S ; Kondratenko, T S
The paper presents the study results of luminescent sensor, based on colloidal Ag2S quantum dots, passivated with L-cysteine molecules (Ag2S/L-Cys QDs) for oxytetracycline (OTC) detection in raw milk. Its work is based on QDs luminescence quenching in the spectral region, which is free from the intrinsic luminescence of the raw milk and OTC (725 nm). A working sensor mechanism was proposed. It is based on the formation of a non-luminescent complex with charge transfer between OTC and Ag2S/L-Cys QDs due to binding the antibiotic with passivator. The sensor parameters were established. The limit of detection (LOD) is 0.53 μM in milk, limit of quantitative detection (LOQ) is 1.16 μM, linear range is 0-20 μM. These parameters are comparable with known luminescent sensor analogs for raw milk.
188
项与 盐酸半胱氨酸 相关的新闻(医药)2026-02-24
作者:椰椰
编辑:李宝珠
转载请联系本公众号获得授权,并标明来源
一文速览 2025 年 AI for Science 最值得关注的前沿论文。
过去一年,AI 与科学研究的关系正在发生一场深刻而安静的转变。从实验设计、数据建模到理论推演,人工智能正以前所未有的速度渗透进科研流程的核心环节,催生出一批难以用传统范式解释的突破性成果。2025 年,AI for Science 不再只是零散的技术应用,而是逐渐演化为一条清晰、系统、可复用的科研创新路径。
与早期「AI 辅助科研」的尝试不同,2025 年的一个显著变化在于:AI 不再只是工具,而是正在成为科研范式的一部分。越来越多的研究工作,从一开始就围绕「如何让模型参与科学发现」展开设计,这也催生了大量兼具方法创新与科学价值的高质量成果。
HyperAI超神经长期关注 AI4S 领域的进展与突破,通过系统解读前沿论文的方式,记录这一浪潮中的关键节点。一方面,我们希望将最新的研究成果与方法论进行结构化、普适化的整理,降低不同领域读者的理解门槛;另一方面,也希望通过持续输出,推动更多研究者、工程师与机构,真正认识到 AI 对科研生产力的深远影响。
值此岁末年初,正是回顾与展望的关键时刻。本文对「HyperAI超神经」在 2025 年解读过的 AI for Science 相关前沿论文进行了系统梳理与分类汇总,覆盖生物医药、医疗健康、材料化学、气象研究、天文学等多个方向,方便不同背景的读者快速检索与回顾。
查看更多前沿论文:
https://hyper.ai/cn/papers
查看研究相关数据集:
https://hyper.ai/cn/datasets
AI+ 生物医药
新的生成式模型及新评测基准,重塑无序蛋白集合预测能力
Advancing Protein Ensemble Predictions Across the Order-Disorder Continuum
*来源:bioRxiv
*作者:Peptone 公司、英伟达公司、麻省理工学院等组成的联合团队
*解读:重塑无序蛋白集合预测能力,英伟达/MIT/牛津大学/哥本哈根大学/Peptone等发布生成式模型及新评测基准
*论文:
https://www.biorxiv.org/content/10.1101/2025.10.18.680935v1
首个经人类皮层数据验证的神经元建模框架 NOBLE,超越传统 4200 倍速
NOBLE - Neural Operator with Biologically-informed Latent Embeddings to Capture Experimental Variability in Biological Neuron Models
*来源:NeurIPS 2025
*作者:苏黎世联邦理工学院、加州理工学院与阿尔伯塔大学等机构的联合团队
*解读:超越传统4200倍速!苏黎世联邦理工提出NOBLE,首个经人类皮层数据验证的神经元建模框架
*论文:
https://go.hyper.ai/Ramfp
图神经网络 PLACER,解决蛋白质构象异质性
Modeling protein-small molecule conformational ensembles with PLACER
*来源:《美国国家科学院院刊》(PNAS)
*作者:华盛顿大学 David Baker 教授的研究团队
*解读:解决蛋白质构象异质性的原子级建模挑战!David Baker团队PLACER框架解析
*论文:
https://www.biorxiv.org/content/10.1101/2024.09.25.614868v2
Squidiff 实现多场景转录组模拟,助力精准医学与空间医学发展
Squidiff: predicting cellular development and responses to perturbations using a diffusion model
*来源:Nature Methods
*作者:哥伦比亚大学、斯坦福大学的联合研究团队
*解读:哥大/斯坦福联手!Squidiff实现多场景转录组模拟,助力精准医学与空间医学发展
*论文:https://www.nature.com/articles/s41592-025-02877-y
血液细胞图像分类器,扩散模型助力白血病发现,能力超越临床专家
Deep generative classification of blood cell morphology
*来源:Nature
*作者:英国剑桥大学的研究团队
*解读:剑桥大学研发血液细胞图像分类器,扩散模型助力白血病发现,能力超越临床专家
*论文:
https://www.nature.com/articles/s42256-025-01122-7
Ctrl-DNA 框架,实现特定细胞基因表达的「靶向控制」
Ctrl-DNA: Constrained Reinforcement Learning for Cell-Specific Cis-Regulatory Element Design
*来源:NeurIPS 2025
*作者:多伦多大学团队联合昌平实验室
*解读:入选NeurIPS 2025,多伦多大学等提出Ctrl-DNA框架,实现特定细胞基因表达的「靶向控制」
*论文:
https://arxiv.org/abs/2505.20578
BoltzGen,可跨分子类型设计蛋白结合物,66%靶标获纳摩尔级亲和力
BoltzGen: Toward Universal Binder Design
*来源:
*作者:麻省理工学院与 Boltz 等多家机构
*解读:MIT团队开源BoltzGen,可跨分子类型设计蛋白结合物,66%靶标获纳摩尔级亲和力
*论文:
https://go.hyper.ai/3sx2K
全新蛋白质动态融合表征框架 FusionProt,实现迭代式信息交换,多项任务性能达到 SOTA
FusionProt: Fusing Sequence and Structural Information for Unified Protein Representation Learning
*来源:bioRxiv
*作者:以色列理工学院联合 Meta AI 的研究团队
*解读:Meta AI等提出全新蛋白质动态融合表征框架FusionProt,实现迭代式信息交换,多项任务性能达到SOTA
*论文:
https://go.hyper.ai/OXLYl
转录组引导的扩散模型 MorphDiff,为表型药物研发提速
Prediction of cellular morphology changes under perturbations with a transcriptome-guided diffusion model
*来源:Nature Communications
*作者:中国香港中文大学、穆罕默德·本·扎耶德人工智能大学等机构的研究人员
*解读:关联基因表达数据与细胞形态图像,港中文等开发转录组引导的扩散模型,为表型药物研发提速
*论文:
https://www.nature.com/articles/s41467-025-63478-z
AlphaPPIMI,PPIs 界面调节剂预测性能超越现有方法
Alphappimi: a comprehensive deep learning framework for predicting PPI-modulator interactions
*来源:Journal of Cheminformatics
*作者:中国石油大学和延世大学的联合研究团队
*解读:从「盲筛」到 「精准定位」,中国石油大学团队推出AlphaPPIMI,PPIs 界面调节剂预测性能超越现有方法
*论文:
https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-01077-2
scSiameseClu 在无监督单细胞聚类任务中达到 SOTA 性能
scSiameseClu: A Siamese Clustering Framework for Interpreting single-cell RNA Sequencing Data
*来源:IJCAI 2025
*作者:中国科学院、东北农业大学、澳门大学与吉林大学的研究团队
*解读:IJCAI 2025丨7个数据集验证:scSiameseClu 在无监督单细胞聚类任务中达到 SOTA 性能
*论文:
https://go.hyper.ai/00BhP
融合神经网络框架,高效预测蛋白质序列的多金属结合位点
A Modular Fusion Neural Network Approach to Efficiently Predict Multi-Metal Binding Sites in Protein Sequences
*来源:bioRxiv
*作者:香港科技大学的研究团队
*解读:香港科技大学提出融合神经网络框架,高效预测蛋白质序列的多金属结合位点
*论文:
https://go.hyper.ai/Y7DNU
ReaSyn 借鉴思维链类比分子合成,实现超高重建率与路径多样性
Rethinking Molecule Synthesizability with Chain-of-Reaction
*来源:arXiv
*作者:英伟达研究团队
*解读:英伟达提出ReaSyn,借鉴思维链类比分子合成,实现超高重建率与路径多样性
*论文:
https://arxiv.org/abs/2509.16084
无序区域结合蛋白设计新方法,专攻不可成药靶点
Design of intrinsically disordered region binding proteins
*来源:Science
*作者:David Baker 及其团队
*解读:登Science,David Baker团队提出无序区域结合蛋白设计新方法,专攻不可成药靶点
*论文:
https://www.science.org/doi/10.1126/science.adr8063
AMix-1 实现可扩展通用的蛋白质设计,设计蛋白变体活性提升50倍
AMix-1: A Pathway to Test-Time Scalable Protein Foundation Model
*来源:arXiv
*作者:清华大学智能产业研究院(AIR)周浩课题组联合上海人工智能实验室
*解读:设计蛋白变体活性提升50倍!清华AIR周浩团队基于贝叶斯流网络提出AMix-1,实现可扩展通用的蛋白质设计
*论文:
https://go.hyper.ai/6Lz0c
双向布朗桥扩散模型输出方差显著降低,提升虚拟染色结果可重复性
Virtual staining of label-free tissue in imaging mass spectrometry
*来源:Science Advances
*作者:UCLA 研究团队
*解读:输出方差显著降低!UCLA发布双向布朗桥扩散模型,提升虚拟染色结果可重复性
*论文:
https://go.hyper.ai/X9GEn
全原子扩散 Transformer ADiT,打破了周期性与非周期性系统的建模壁垒
All-atom Diffusion Transformers: Unified generative modelling of molecules and materials
*来源:ICML 2025
*作者:Meta FAIR、剑桥大学与麻省理工学院的联合科研团队
*解读:入选ICML 2025,Meta/剑桥/MIT提出全原子扩散Transformer框架,首次实现周期性与非周期性原子系统统一生成
*论文:
https://go.hyper.ai/27d7U
Full-Atom MPNN 显式建模每个氨基酸残基的序列身份和侧链结构
Sidechain conditioning and modeling for full-atom protein sequence design with FAMPNN
*来源:ICML 2025
*作者:斯坦福大学的团队联合加州帕洛阿尔托市 Arc 研究院
*解读:同时处理蛋白质主链和侧链信息,斯坦福等基于消息传递神经网络实现全原子结构建模
*论文:
https://go.hyper.ai/JUJDq
La-Proteina 实现原子级蛋白质设计突破,高精度生成多达 800 个残基的蛋白质
La-Proteina: Atomistic Protein Generation via Partially Latent Flow Matching
*来源:arXiv
*作者:NVIDIA 的研究团队联合加拿大魁北克人工智能研究所 Mila
*解读:英伟达实现原子级蛋白质设计突破,高精度生成多达800个残基的蛋白质
https://go.hyper.ai/3csT5
深度学习模型 SUICA 实现空间转录组切片中任一位置基因表达的预测
SUICA: Learning Super-high Dimensional Sparse Implicit Neural Representations for Spatial Transcriptomics
*来源:ICML 2025
*作者:东京大学郑银强老师组,麦吉尔大学丁俊老师组
*解读:数据降噪/生物信号强化/缓解dropout,深度学习模型SUICA实现空间转录组切片中任一位置基因表达的预测
*论文:
https://go.hyper.ai/C6Zcl
全新全原子蛋白质生成模型 APM 实现全原子设计与功能优化
An All-Atom Generative Model for Designing Protein Complexes
*来源:ICML 2025
*作者:湖南大学联合中国科学院大学、字节跳动 Seed 团队
*解读:支持蛋白质生成/折叠/逆折叠,湖大/中科大/字节提出APM模型,实现全原子设计与功能优化
*论文:
https://go.hyper.ai/TVp4i
深度学习模型 APEX,筛选潜在抗生素候选物
Computational exploration of global venoms for antimicrobial discovery with Venomics artificial intelligence
*来源:Nature Communications
*作者:美国宾夕法尼亚大学的研究团队
*解读:从动物毒液中挖掘386种全新抗菌肽,宾夕法尼亚大学开发深度学习模型APEX,筛选潜在抗生素候选物
*论文:
https://www.nature.com/articles/s41467-025-60051-6[*]
蛋白质语言模型 Prot42 实现长序列建模与高亲和力结合剂生成
Prot42: a Novel Family of Protein Language Models for Target-aware Protein Binder Generation
*来源:arXiv
*作者:阿布扎比 Inception AI 研究所与硅谷 Cerebras Systems 公司的联合研究团队
*解读:8k长序列建模,蛋白质语言模型Prot42仅利用目标蛋白序列即可生成高亲和力结合剂
*论文:
https://go.hyper.ai/cFupD
生物分子时间粗化动力学模拟器 UniSim,首次实现了跨分子类型、跨化学环境的统一时间粗化动力学模拟
UniSim: A Unified Simulator for Time-Coarsened Dynamics of Biomolecules
*来源:ICML 2025
*作者:清华大学、人民大学高瓴人工智能学院
*解读:入选 ICML 2025,清华/人大提出统一生物分子动力学模拟器 UniSim
*论文:
https://go.hyper.ai/5NWuO
SimplifiedBondfinder 基于 8.6 万蛋白质结构数据,融合量子力学计算的机器学习方法挖掘 69 个全新氮-氧-硫键
Revealing arginine-cysteine and glycine-cysteine NOS linkages by a systematic re-evaluation of protein structures
*来源:Communications Chemistry
*作者:乔治奥古斯特大学的团队
*解读:基于8.6万蛋白质结构数据,融合量子力学计算的机器学习方法挖掘69个全新氮-氧-硫键
*论文:
https://www.nature.com/articles/s42004-025-01535-w
利用蛋白质序列生成模型实现重叠基因设计,成功率极高
Design of overlapping genes using deep generative models of protein sequences
*来源:bioRxiv
*作者:美国华盛顿大学 David Baker 团队
*解读:David Baker 团队最新研究,利用蛋白质序列生成模型实现重叠基因设计,成功率极高
*论文:
https://doi.org/10.1101/2025.05.06.652464
预测框架 PUPS 创新地结合蛋白质语言模型和图像修复模型,实现单细胞级蛋白质定位
Prediction of protein subcellular localization in single cells
*来源:Nature Methods
*作者:麻省理工学院和哈佛大学的团队
*解读:融合蛋白质语言模型和图像修复模型,麻省理工与哈佛联手提出PUPS,实现单细胞级蛋白质定位
*论文:
https://go.hyper.ai/LeaQF
首个跨分子种类统一生成框架 UniMoMo,实现多类型药物分子设计
UniMoMo: Unified Generative Modeling of 3D Molecules for De Novo Binder Design
*来源:ICML 2025
*作者:清华大学刘洋老师组联合人大黄文炳老师组和字节 AI 制药团队
*解读:入选ICML 2025,清华/人大/字节提出首个跨分子种类统一生成框架UniMoMo,实现多类型药物分子设计
*论文:
https://go.hyper.ai/wZXZZ
深度学习框架 STAIG,揭示肿瘤微环境中的详细基因信息
STAIG: Spatial transcriptomics analysis via image-aided graph contrastive learning for domain exploration and alignment-free integration
*来源:Nature Communications
*作者:日本东京大学医科学研究所的研究团队
*解读:无需预对齐即可消除批次效应,东京大学团队开发深度学习框架STAIG,揭示肿瘤微环境中的详细基因信息
*论文:
https://www.nature.com/articles/s41467-025-56276-0
DRAKES 算法引入强化学习框架,首次实现了在离散扩散模型中对完整生成轨迹的可微奖励反向传播
Fine-Tuning Discrete Diffusion Models via Reward Optimization with Applications to DNA and Protein Design
*来源:ICLR 2025
*作者:美国麻省理工学院、哈佛大学、斯坦福大学、加州大学伯克利分校以及美国基因工程技术公司 Genentech 的研究人员
*解读:入选ICLR 2025,MIT/UC伯克利/哈佛/斯坦福等提出DRAKES算法,突破生物序列设计瓶颈
*论文:
https://doi.org/10.48550/arXiv.2410.13643
机器学习辅助的紫外吸收光谱法,基于 SVM 构建微生物污染检测模型
Machine learning aided UV absorbance spectroscopy for microbial contamination in cell therapy products
*来源:Scientific Reports
*作者:新加坡-麻省理工学院研究联盟、新加坡 A*SRL 实验室、新加坡国立大学、美国麻省理工学院的联合研究团队
*解读:30分钟内输出结果,新加坡国立大学/MIT等基于SVM构建微生物污染检测模型
*论文:
https://doi.org/10.1038/s41598-024-83114-y
蛋白质预训练新范式,解密蛋白质家族进化
Steering Protein Family Design through Profile Bayesian Flow
*来源:ICLR 2025
*作者:清华大学 AIR GenSI 研究组联合清华大学药学院
*解读:入选ICLR 2025 Oral,清华AIR周浩团队提出蛋白质预训练新范式,解密蛋白质家族进化
*论文:
https://go.hyper.ai/Dg5ha
玻尔兹曼对齐技术蛋白质结合自由能预测达 SOTA
Boltzmann-Aligned Inverse Folding Model as a Predictor of Mutational Effects on Protein-Protein Interactions
*来源:ICLR 2025
*作者:浙江大学计算机科学与技术学院沈春华教授团队联合澳大利亚阿德莱德大学、美国东北大学等团队
*解读:入选ICLR 2025!浙大沈春华等人提出玻尔兹曼对齐技术,蛋白质结合自由能预测达SOTA
*论文:
https://arxiv.org/abs/2410.09543
Proteina 模型参数超 RFdiffusion 5倍,从头设计蛋白质主链性能达 SOTA
Proteina: Scaling Flow-based Protein Structure Generative Models
*来源:ICLR 2025 Oral
*作者:英伟达联合魁北克人工智能研究所 Mila、蒙特利尔大学、麻省理工学院的研究团队
*解读:模型参数超 RFdiffusion 5倍!英伟达等发布 Proteina,从头设计蛋白质主链性能达 SOTA
*论文:
https://openreview.net/forum?id=TVQLu34bdw&nesting=2&sort=date-desc
UniGEM 模型首次基于扩散模型实现两任务协同增强
UniGEM: A Unified Approach to Generation and Property Prediction for Moleculess
*来源:ICLR 2025
*作者:清华大学联合中科院团队
*解读:首次实现分子生成与性质预测的统一,清华团队提出两阶段扩散生成机制,入选ICLR 2025
*论文:
https://openreview.net/pdf?id=Lb91pXwZMR
RFdiffusion 再进化,实现原子级精度的抗体从头设计
Atomically accurate de novo design of antibodies with RFdiffusion
*来源:bioRxiv
*作者:David Baker 团队及其合作者
*解读:David Baker团队新成果!RFdiffusion再进化,实现原子级精度的抗体从头设计
*论文:
https://doi.org/10.1101/2024.03.14.585103
首个蛋白质 -RNA 语言模型融合方案,结合亲和力预测刷新 SOTA
CoPRA: Bridging Cross-domain Pretrained Sequence Models with Complex Structures for Protein-RNA Binding Affinity Prediction
*来源:arXiv
*作者:清华大学、伦敦大学学院、莫纳什大学、北京邮电大学的联合团队
*解读:入选AAAI 2025!清华/伦敦大学学院等首创蛋白质-RNA语言模型融合方案,结合亲和力预测刷新SOTA
*论文:
https://arxiv.org/abs/2409.03773
Celcomen 模型首次在空间转录组学分析中实现因果推断可识别性
Estimation of single-cell and tissue perturbation effect in spatial transcriptomics via Spatial Causal Disentanglement
*来源:ICLR 2025
*作者:剑桥大学的研究团队
*解读:入选ICLR 2025!剑桥大学提出Celcomen模型,首次在空间转录组学分析中实现因果推断可识别性
*论文:
https://openreview.net/forum?id=Tqdsruwyac
AlphaFold-Metainference 方法精准预测无序蛋白质结构集合
AlphaFold prediction of structural ensembles of disordered proteins
*来源:Nature Communications
*作者:剑桥大学的研究团队
*解读:AlphaFold应用新里程碑!剑桥大学团队提出AlphaFold-Metainference,精准预测无序蛋白质结构集合
*论文:
https://www.nature.com/articles/s41467-025-56572-9
第二代 RNA 结构预测算法,多项基准测试超越 SOTA
Ab initio RNA structure prediction with composite language model and denoised end-to-end learning
*来源:bioRxiv
*作者:新加坡国立大学的张阳教授团队
*解读:新加坡国立大学张阳团队开发第二代RNA结构预测算法,多项基准测试超越SOTA
*论文:
https://www.biorxiv.org/content/10.1101/2025.03.05.641632v1
创新性 4D 扩散模型结合分子动力学模拟数据,能够同时预测多个时间步长的蛋白质运动轨迹
4D Diffusion for Dynamic Protein Structure Prediction with Reference and Motion Guidance
*来源:arXiv
*作者:复旦大学、上海科学智能研究院和南京大学的研究团队
*解读:AlphaFolding填补蛋白质动态结构预测空白!复旦大学等提出4D扩散模型,成果入选AAAI 2025
*论文:
https://arxiv.org/abs/2408.12419
PepPrCLIP 破解「不可成药」难题,创建几乎总是与目标蛋白质更匹配的肽
De novo design of peptide binders to conformationally diverse targets with contrastive language modeling
*来源:Science Advances
*作者:杜克大学的研究团队
*解读:有望开发癌症新疗法!杜克大学用PepPrCLIP破解「不可成药」难题
*论文:
https://www.science.org/doi/10.1126/sciadv.adr8638
MOLRL 基于强化学习优化靶向分子,成功率可达100%
Targeted Molecular Generation With Latent Reinforcement Learning
*来源:ChemRxiv
*作者:Cellarity 公司和英伟达的研究团队
*解读:成功率可达100%,药物开发公司Cellarity联手英伟达,基于强化学习优化靶向分子
*论文:
https://go.hyper.ai/H4JhR
进化驱动的病毒变异驱动力预测框架 E2VD,预测精度提升 67%
A unified evolution-driven deep learning framework for virus variation driver prediction
*来源:Nature Machine Intelligence
*作者:北京大学信息工程学院田永鸿教授、陈杰副教授,联合广州国家实验室周鹏研究员指导博士生聂志伟、硕士生刘旭东等
*解读:登Nature子刊!北大团队用AI预测新冠/艾滋病/流感病毒进化方向,精度提升67%
*论文:
https://www.nature.com/articles/s42256-024-00966-9
AI+ 医疗健康
Healthcare Agent 问诊主动性及相关性超越GPT-4等闭源模型
Healthcare agent: eliciting the power of large language models for medical consultation
*来源:Nature Artificial Intelligence
*作者:武汉大学和南洋理工大学研究团队
*解读:从伦理保障到病史管理,武汉大学等提出Healthcare Agent,问诊主动性及相关性超越GPT-4等闭源模型
*论文:
https://go.hyper.ai/09lYX
ICA-Var 的多变量分析方法,基于基因测序和机器学习的废水流行病学评估,病毒检出时间最高提前 4 周
Early detection of emerging SARS-CoV-2 Variants from wastewater through genome sequencing and machine learning
*来源:Nature Communications
*作者:内华达大学拉斯维加斯分校的研究团队
*解读:登Nature子刊,基于基因测序和机器学习的废水流行病学评估,病毒检出时间最高提前4周
*论文:
https://www.nature.com/articles/s41467-025-61280-5
医学 GraphRAG 刷新问答准确性记录,在 11 个数据集评测上达 SOTA
Medical Graph RAG: Towards Safe Medical Large Language Model via Graph Retrieval-Augmented Generation
*来源:ACL 2025
*作者:牛津大学、卡内基梅隆大学与爱丁堡大学的联合团队
*解读:ACL 2025丨牛津大学等提出医学GraphRAG,刷新问答准确性记录,在11个数据集评测上达SOTA
*论文:
https://go.hyper.ai/OaMIE
首个 VR 运动干预系统 REVERIE(「灵境」),重塑青少年脑-身-心健康
Adaptive AI-based virtual reality sports system for adolescents with excess body weight: a randomized controlled trial
*来源:Nature Medicine
*作者:上海交通大学医学院附属第六人民医院/主动健康战略与发展研究院李华婷教授团队、上海交通大学计算机学院/人工智能教育部重点实验室盛斌教授团队通过医工交叉合作研究,携手上海体育大学王继红研究员团队、上海科技大学/上海临床研究中心曾嵘教授团队及新加坡国立大学林水德教授团队
*解读:钱学森「灵境」 预言成真!上交/上体/清华等构建全球首个VR运动干预系统REVERIE,重塑青少年脑-身-心健康
*论文:
https://www.nature.com/articles/s41591-025-03724-5
NeuralCohort 方法基于多维度 EHR 数据实现细粒度患者队列建模,住院时间预测准确率提升 16.3%
NeuralCohort: Cohort-aware Neural Representation Learning for Healthcare Analytics
*来源:ICML 2025
*作者:新加坡国立大学联合浙江大学
*解读:新加坡国立大学基于多维度EHR数据实现细粒度患者队列建模,住院时间预测准确率提升16.3%
*论文:
https://openreview.net/forum?id=bqQVa6VRvm
全球首个 HIE 领域临床思维图谱模型,神经认知结果预测任务上性能提升 15%
Visual and Domain Knowledge for Professional-level Graph-of-Thought Medical Reasoning
*来源:ICML 2025
*作者:哈佛医学院、波士顿儿童医院、纽约大学及 MIT-IBM 沃森实验室的跨学科团队
*解读:入选ICML 2025!哈佛医学院等推出全球首个HIE领域临床思维图谱模型,神经认知结果预测任务上性能提升15%
*论文:
https://openreview.net/forum?id=tnyxtaSve5
分层蒸馏多示例学习框架 HDMIL,快速处理千兆像素病理全切片图像
Fast and Accurate Gigapixel Pathological Image Classification with Hierarchical Distillation Multi-Instance Learning
*来源:CVPR 2025
*作者:哈尔滨工业大学江俊君教授、江奎副教授和张永兵教授团队
*解读:入选CVPR 2025,哈工大团队提出分层蒸馏多示例学习框架HDMIL,快速处理千兆像素病理全切片图像
*论文:
https://go.hyper.ai/B3RMf
专为 3D 血管分割而设计的基础模型 vesselFM,能够在零样本、单样本和少样本场景中实现优于现有先进模型的分割能力和泛化能力。
vesselFM: A Foundation Model for Universal 3D Blood Vessel Segmentation
*来源:CVPR 2025
*作者:苏黎世大学、苏黎世联邦理工学院和慕尼黑工业大学的研究人员
*解读:性能远超SAM系模型,苏黎世大学等开发通用3D血管分割基础模型,入选CVPR 2025
*论文:
https://go.hyper.ai/lVad9
图编码混合生存模型基于 800 万真实数据,识别具有一致特征和生存结局的亚表型
Identification of predictive subphenotypes for clinical outcomes using real world data and machine learningn
*来源:Nature Communication
*作者:美国康奈尔大学与再生元制药公司
*解读:基于800万真实数据,康奈尔大学团队利用图神经网络精准预测肺癌患者生存期,发现3类致命亚型
*论文:
https://doi.org/10.1038/s41467-025-59092-8
融合策略AI模型,实现多中心、跨专科感染性休克死亡风险的精准预测
Artificial intelligence based multispecialty mortality prediction models for septic shock in a multicenter retrospective study
*来源:npj digital medicine
*作者:华中科技大学同济医学院附属同济医院、医药卫生管理学院研究团队
*解读:登Nature子刊!华中科技大学提出融合策略AI模型,实现多中心、跨专科感染性休克死亡风险的精准预测
*论文:
https://go.hyper.ai/faMLL
2 种新型癌症预测算法,基于血液指标实现 15 种癌症早期预测
Development and external validation of prediction algorithms to improve early diagnosis of cancer
*来源:Nature Communications
*作者:伦敦玛丽女王大学与牛津大学研究团队
*解读:牛津大学等深挖746万成年人健康数据开发早筛算法,基于血液指标实现15种癌症早期预测
*论文:
https://go.hyper.ai/L7gNm
多智能体对话 (MAC) 框架,显著提升 LLMs 的诊断能力
Enhancing diagnostic capability with multi-agents conversational large language models
*来源:npj digital medicine
*作者:四川大学华西医院、华西生物医学大数据中心、浙江大学医学院、北京邮电大学等团队
*解读:模拟医生会诊,四川大学华西医院团队开发多智能体对话框架助力疾病诊断
*论文:
https://www.nature.com/articles/s41746-025-01550-0#Tab6
首个全模态医疗图像重识别框架,在 11 个数据集上的评测达 SOTA
Towards All-in-One Medical Image Re-Identification
*来源:
*作者:上海人工智能实验室联合多家知名高校
*解读:入选CVPR 2025,上海AI Lab等提出首个全模态医疗图像重识别框架,在11个数据集上的评测达SOTA
*论文:
https://arxiv.org/pdf/2503.08173
多对一回归模型 M2OST,利用数字病理图像精准预测基因表达
M2OST: Many-to-one Regression for Predicting Spatial Transcriptomics from Digital Pathology Images
*来源:AAAI 2025
*作者:中国浙江大学的林兰芬教授研究团队联合浙江杭州之江实验室以及日本立命馆大学
*解读:入选AAAI 2025,浙江大学提出多对一回归模型M2OST,利用数字病理图像精准预测基因表达
*论文:
https://arxiv.org/abs/2409.15092
MindGlide 模型,实现多发性硬化症病变量化
Enabling new insights from old scans by repurposing clinical MRI archives for multiple sclerosis research
*来源:Nature Communications
*作者:英国伦敦大学学院团队
*解读:最大化挖掘临床MRI数据价值,UCL团队提出MindGlide模型,实现多发性硬化症病变量化
*论文:
https://www.nature.com/articles/s41467-025-58274-8
全球首个针对大模型在辅助基层医生培训中实际效能的前瞻性真实世界证据
Large language models for diabetes training: a prospective study
*来源:Science Bulletin
*作者:上海交通大学盛斌教授团队联合上海体育大学毛丽娟教授团队、携手清华大学黄天荫教授团队和上海市糖尿病研究所贾伟平教授团队等多学科力量,联合美国杜克大学、约翰霍普金斯大学以及澳洲墨尔本大学等国际顶尖学府和研究机构
*解读:医生培训迎来 DeepSeek 外挂!上体/上交/清华合作研究证实大模型可成为基层医生培训 「黄金搭档」
*论文:
https://www.sciencedirect.com/science/article/pii/S2095927325000891
AcneDGNet 的深度学习算法准确率远超初级皮肤科医生,实现痤疮病变检测与分级
Evaluation of an acne lesion detection and severity grading model for Chinese population in online and offline healthcare scenarios
*来源:Scientific Reports
*作者:北京大学国际医院皮肤科韩钢文及其团队
*解读:准确率远超初级皮肤科医生,北大国际医院等开发深度学习算法,实现痤疮病变检测与分级
*论文:
https://www.nature.com/articles/s41598-024-84670-z
多模态医学影像分割模型,实现三维影像自动分割与交互
VISTA3D: A Unified Seamentation Foundation Model For 3D Medical lmaging
*来源:arXiv
*作者:英伟达联合阿肯色大学医学院、美国国立卫生研究院及牛津大学
*解读:精度提升5.2%,英伟达等发布多模态医学影像分割模型,实现三维影像自动分割与交互
*论文:
https://doi.org/10.48550/arxiv.2406.05285
对多切面超声心动图的心脏结构进行精准分割,有效减少冗杂度
EchoONE: Segmenting Multiple echocardiography Planes in One Model
*来源:CVPR 2025
*作者:深圳大学医学部生物医学工程学院医学超声图像计算实验室
*解读:入选CVPR 2025!深圳大学团队等提出EchoONE,可精准分割多切面超声心动图
*论文:https://arxiv.org/abs/2412.02993
当前具有最大规模参数量的生物医学大语言模型 MedFound
A generalist medical language model for disease diagnosis assistance
*来源:Nature Medicine
*作者:北京邮电大学、北京大学第三医院、三峡大学
*解读:开源1760亿参数通用医学语言模型!北邮/北大/三峡大学提出MedFound,推理能力接近专家医师
*论文:https://www.nature.com/articles/s41591-024-03416-6
对比度驱动医学图像分割的通用框架,实现医学图像精准分割
ConDSeg: A General Medical Image Segmentation Framework via Contrast-Driven Feature Enhancement
*来源:AAAI 2025
*作者:中国地质大学、百度
*解读:入选AAAI 2025!解决医学图像分割软边界与共现难题,中国地质大学等提出图像分割模型ConDSeg
*论文:
https://arxiv.org/abs/2412.08345
在 2 种传染性和 14 种非传染性疾病的 9 个基准数据集上均为最优
A multimodal multidomain multilingual medical foundation model for zero shot clinical diagnosis
*来源:Nature Portfolio
*作者:牛津大学、亚马逊、罗切斯特大学、GlaxoSmithKline、西湖大学医学人工智能实验室
*解读:牛津/亚马逊/西湖大学/腾讯等提出多模态多领域多语言医学模型M³FM,可用于零样本临床诊断
*论文:
https://www.nature.com/articles/s41746-024-01339-7
分类准确率可达 97%,显著高于人类观察者的 82%
Deep learning versus human assessors: forensic sex estimation from three-dimensional computed tomography scans
*来源:Scientific Reports
*作者:澳大利亚西澳大学、新南威尔士大学、印度尼西亚哈萨努丁大学
*解读:准确率达97%,澳大利亚团队新成果基于深度学习凭颅骨CT鉴定性别,赶超人类法医
*论文:
https://www.nature.com/articles/s41598-024-81718-y
双向逐步特征对齐的未对齐医学图像融合方法
BSAFusion: A Bidirectional Stepwise Feature Alignment Network for Unaligned Medical Image Fusion
*来源:AAAI 2025
*作者:昆明理工大学、中国海洋大学
*解读:入选AAAI 2025!可实现多模态医学图像对齐与融合,国内两大高校联合提出BSAFusion
*论文:https://arxiv.org/abs/2412.08050
新型分层多智能体框架,涵盖了 362 种常见疾病
KG4Diagnosis: A Hierarchical Multi-Agent LLM Framework with Knowledge Graph Enhancement for Medical Diagnosis
*来源:AAAI-25 Bridge Program
*作者:华威大学、克兰菲尔德大学、剑桥大学、牛津大学
*解读:助力诊断362种常见疾病!剑桥/牛津/华威大学等提出多Agent大语言模型框架,自动化构建医疗知识图谱
*论文:https://arxiv.org/abs/2406.05285
AI+ 材料化学
不足 10 万结构数据训练,PET-MAD 原子模拟精度媲美专业模型
PET-MAD as a lightweight universal interatomic potential for advanced materials modeling
*来源:Nature Communications
*作者:瑞士洛桑理工学院
*解读:以不足10万结构数据训练,瑞士洛桑联邦理工提出PET-MAD,原子模拟精度媲美专业模型
*论文:
https://www.nature.com/articles/s41467-025-65662-7
计算成本减半,将人类的化学推理形式化为机器可理解的框架
ChemOntology: A Reusable Explicit Chemical Ontology-Based Method to Expedite Reaction Path Searches
*来源:ACS Catalysis
*作者:日本北海道大学
*解读:计算成本减半,化学反应发现工具ChemOntology将人类直觉「编码」到系统中,加速反应路径搜索
*论文:
https://pubs.acs.org/doi/10.1021/acscatal.5c06298
MOF-ChemUnity:一个结构化、可扩展、可拓展的知识图谱
MOF-ChemUnity: Literature-Informed Large Language Models for Metal–Organic Framework Research
*来源:ACS Publications
*作者:加拿大多伦多大学、加拿大国家研究委员会清洁能源创新研究中心
*解读:从9,874篇文献到1.5万晶体结构,MOF-ChemUnity重构MOF全景知识,推动材料发现进入「可解释AI」时代
*论文:
https://pubs.acs.org/doi/10.1021/jacs.5c11789
将 Graphormer 的全局注意力机制与 CGCNN 相融合
CGformer: Transformer-enhanced crystal graph network with global attention for material property prediction
*来源:Matter
*作者:上海交通大学人工智能与微结构实验室
*解读:新材料研发提速!上交大团队开发新AI材料设计模型CGformer,融合全局注意力机制
*论文:
https://www.cell.com/matter/abstract/S2590-2385(25)00423-0
FASTSOLV 模型:实现任意温度下的小分子溶解度预测
Data-driven organic solubility prediction at the limit of aleatoric uncertainty
*来源:Nature Communication
*作者:麻省理工学院
*解读:推理速度快50倍,MIT团队提出FASTSOLV模型,实现任意温度下的小分子溶解度预测
*论文:
https://www.nature.com/articles/s41467-025-62717-7
利用 MOFs 合成后即可获得的信息来预测其潜在性能和用途
Connecting metal-organic framework synthesis to applications using multimodal machine learning
*来源:Nature Communications
*作者:加拿大多伦多大学
*解读:多模态模型加速新材料与工业应用匹配,无需完整晶体结构即可预测材料性质
*论文:
https://www.nature.com/articles/s41467-025-60796-0
首次构建能够同时处理超材料设计三大模态的统一框架
UNIMATE: A Unified Model for Mechanical Metamaterial Generation, Property Prediction, and Condition Confirmation
*来源:ICML 2025
*作者:美国弗吉尼亚理工学院、 Meta AI
*解读:超材料设计破局!Meta AI等提出UNIMATE,首次实现拓扑生成/性能预测等任务的统一建模
*论文:
https://go.hyper.ai/FoAWw
覆盖 2 亿分子质谱图,构建全球最大规模质谱数据集 GeMS
Self-supervised learning of molecular representations from millions of tandem mass spectra using DreaMS
*来源:Nature Biotechnology
*作者:捷克科学院有机化学与生物化学研究所
*解读:覆盖2亿分子质谱图,捷克科学院发布DreaMS模型,构建全球最大规模质谱数据集GeMS
*论文:
https://go.hyper.ai/uNbqL
在单一机器学习模型中同时学习广义势能及其对外部刺激的响应函数
Unified differentiable learning of electric response
*来源:Nature Communications
*作者:哈佛大学、Robert Bosch LLC
*解读:从石英到铁电材料,哈佛大学提出等变机器学习框架,加速材料大规模电场模拟
*论文:
https://go.hyper.ai/18TWg
基于 LLM 从 8.8 万篇论文中提取 1.4 万种材料的化学组成
Data-driven material screening of secondary and natural cementitious precursors
*来源:Communication Materials
*作者:美国麻省理工学院
*解读:MIT团队利用大模型筛选25类水泥熟料替代材料,相当于减排12亿吨温室气体
*论文:
https://go.hyper.ai/ZOAaW
将全面的 SSE 数据库与 LLM 和 ab initio 元动力学模拟相结合
Unraveling the Complexity of Divalent Hydride Electrolytes in Solid-State Batteries via a Data-Driven Framework with Large Language Model
*来源:Angewandte Chemie-International Edition
*作者:日本东北大学、中国四川大学、日本芝浦工业大学
*解读:中日团队联合攻关,利用大模型解析氢化物固态电解质传导机制,建立可靠活化能预测模型
*论文:
https://go.hyper.ai/isQRi
在高达 TB 级别的多组分高分辨质谱数据库中,检索离子同位素分布
Discovering organic reactions with a machine-learning-powered deciphering of tera-scale mass spectrometry data
*来源:Nature Communications
*作者:俄罗斯科学院
*解读:登Nature子刊,俄罗斯研究团队基于机器学习实现万亿级质谱数据搜索,发现未知化学反应
*论文:
https://go.hyper.ai/ak7bN
一种基于扩散模型的生成式人工智能结构解析方法 PXRDnet
Ab initio structure solutions from nanocrystalline powder diffraction data via diffusion models
*来源:Nature Materials
*作者:哥伦比亚大学、斯坦福大学
*解读:首次实现纳米晶体端到端解析,哥大团队提出PXRDnet,成功解析200种复杂模拟纳米晶体
*论文:
https://go.hyper.ai/r1K6b
基于机器学习成功制备 10 种光驱动有机晶体
Machine learning-driven optimization of the output force in photo-actuated organic crystals
*来源:Digital Discovery
*作者:日本早稻田大学
*解读:效率提升73倍!日本研究团队基于机器学习成功制备10种光驱动有机晶体
*论文:
https://go.hyper.ai/RU0ro
成功预测金属的电声相互作用,效率提高 5 倍
Accelerating superconductor discovery through tempered deep learning of the electron-phonon spectral function
*来源:npj Computational Materials
*作者:美国佛罗里达大学、田纳西大学
*解读:超导材料搜索效率提升5倍!佛罗里达大学等用深度学习变革材料发现,成果登Nature子刊
*论文:
https://www.nature.com/articles/s41524-024-01475-4
通过梯度提升决策树技术,实现对 RHEAs 和 RCCAs 抗氧化性能的高精度预测
Advancing refractory high entropy alloy development with AI-predictive models for high temperature oxidation resistance
*来源:Scripta Materialia
*作者:法国波尔多大学、日本国立材料科学研究所、中国台湾国立清华大学、比利时鲁汶大学、比利时 WEL 研究所
*解读:高熵合金新发现!多团队联手实现抗氧化性高精度预测,增加铝/铬/硅含量可有效改善
*论文:
https://doi.org/10.1016/j.scriptamat.2024.116394
结合局部消息传递与全局注意力机制,能精准预测分子光电性能
RingFormer: A Ring-Enhanced Graph Transformer for Organic Solar Cell Property Prediction
*来源:AAAI 2025
*作者:香港理工大学
*解读:入选AAAI 2025!香港理工大学团队基于图Transformer,精准预测有机材料分子光电性能
*论文:
https://doi.org/10.48550/arXiv.2412.09030
一种无机逆合成规划方法,成功地促进了无机材料合成的效率和准确性
Retrieval-Retro: Retrieval-based Inorganic Retrosynthesis with Expert Knowledge
*来源:NeurIPS 2024
*作者:韩国化学技术研究所 、韩国科学技术院
*解读:无机材料逆合成效率飙升,韩国团队推出Retrieval-Retro,成果入选NeurIPS 2024
*论文:
https://doi.org/10.48550/arXiv.2410.21341
基于扩散模型,根据目标空间群生成结构
A generative model for inorganic materials design
*来源:Nature
*作者:微软
*解读:直接设计目标属性材料!微软MatterGen模型重磅开源,用生成式AI重新定义材料逆向设计新范式
*论文:
https://www.nature.com/articles/s41586-025-08628-5
AI+ 农林畜牧业
覆盖近 1.5 万个物种,刷新生物声学分类检测 SOTA
Perch 2.0: The Bittern Lesson for Bioacoustics
*来源:arXiv
*作者:Google DeepMind、Google Research
*解读:覆盖近1.5万个物种,谷歌DeepMind发布Perch 2.0,刷新生物声学分类检测SOTA
*论文:
https://arxiv.org/abs/2508.04665
首次将 219 个基于傅里叶变换、香农熵等数学理论的新型序列描述符纳入特征空间
PlantLncBoost: key features for plant lncRNA identification and significant improvement in accuracy and generalization
*来源:New Phytologist
*作者:山东理工大学、北京林业大学、广东省农业科学院、巴西圣保罗大学、英国罗莎琳德富兰克林医科大学、瑞典于默奥大学
*解读:整合多源植物转录组数据,山东理工大学等构建PlantLncBoost模型,跨物种lncRNA预测准确率最高达96%
*论文:
https://go.hyper.ai/F7pkc
AI+ 气象研究
攻克了「渐进式噪声调度」与「时间损失加权」的协同设计问题
Elucidated Rolling Diffusion Models for Probabilistic Weather Forecasting
*来源:NeurIPS 2025
*作者:英伟达、加州大学圣迭戈分校
*解读:入选NeurIPS 2025,英伟达提出ERDM模型,解长期预报难题,中远期预报持续领先EDM基准
*论文:
https://doi.org/10.48550/arXiv.2506.20024
一种新型潜在扩散模型,可用于高精度概率性次季节至季节尺度天气预报
OmniCast: A Masked Latent Diffusion Model for Weather Forecasting Across Time Scales
*来源:NeurIPS 2025
*作者:加州大学洛杉矶分校、美国阿贡国家实验室
*解读:效率至高提升20倍!加州大学开发OmniCast,解决自回归天气预报模型误差累计问题
*论文:
https://go.hyper.ai/YANIu
超本地预测模型,实现了提前数天预测大部分强降雨事件
Hyperlocal Extreme Rainfall Forecasts in Mumbai: Convolutional Neural Network Transfer Learning-Based Downscaling Approach
*来源:SSRN
*作者:印度理工学院孟买分校、马里兰大学
*解读:准确度提升400%!印度季风预测模型基于36个气象站点,实现城区尺度精细预报
*论文:
https://go.hyper.ai/j05Vt
仅需 2 分钟,ACE2 可完成一次 4 个月季节预报
Skilful global seasonal predictions from a machine learning weather model trained on reanalysis data
*来源:npj Climate and Atmospheric Science
*作者:英国埃克塞特哈德利中心气象局、埃克塞特大学、美国艾伦人工智能研究所(Ai2)
*解读:机器学习vs.动力学模型,Ai2最新研究:仅需2分钟,ACE2可完成一次4个月季节预报
*论文:
https://go.hyper.ai/YyRfT
一个将球面信号处理与隐马尔可夫集合框架相结合的概率机器学习天气预报系统
FourCastNet 3: A geometric approach to probabilistic machine-learning weather forecasting at scale
*来源:arXiv
*作者:英伟达、美国劳伦斯伯克利国家实验室、加州大学伯克利分校、美国加州理工学院
*解读:1分钟内完成15天预报,英伟达/UC伯克利等提出机器学习天气预报系统FCN3,支持单卡极速推理
*论文:
https://arxiv.org/pdf/2507.12144
无需依赖传统数值天气预报模型,即可实现高技巧性的天气预报
End-to-end data-driven weather prediction
*来源:Nature
*作者:剑桥大学、图灵研究所、多伦多大学、微软科学智能中心、欧洲中期天气预报中心、英国南极调查局、谷歌 DeepMind
*解读:登Nature,剑桥大学等发布首个端到端的数据驱动天气预报系统,预测速度提升数十倍
*论文:
https://www.nature.com/articles/s41586-025-08897-0
AI+ 天文学
利用卷积神经网络,成功识别出 7 个高质量类星体透镜候选体
Quasars acting as Strong Lenses Found in DESI DR1
*来源:arXiv
*作者:斯坦福大学、SLAC 国家加速器实验室、北京大学、意大利国家天体物理研究院布雷拉天文台、伦敦大学学院、加州大学伯克利分校
*解读:斯坦福/北大/UCL/UC伯克利联手,利用CNN从81万类星体中精准识别7个罕见透镜样本
*论文:
https://arxiv.org/abs/2511.02009
首个面向天文学的大规模多模态基础模型家族 AION-1
AION-1: Omnimodal Foundation Model for Astronomical Sciences
*来源:NeurIPS 2025
*作者:加州大学伯克利分校、剑桥大学、牛津大学等
*解读:首个天文多模态基础模型AION-1诞生!UC伯克利等基于2亿天文目标预训练,成功构建泛化性多模态天文AI框架
*论文:
https://openreview.net/forum?id=6gJ2ZykQ5W
一键获取 2023—2024 年 AI4S 领域高质量论文及深度解读文章 ⬇️
往期推荐
戳“阅读原文”,免费获取海量数据集资源!
2026-02-18
·知乎专栏
摘要本文系统分析了宇宙空间辐射对人体健康的危害机制,以及防辐射药物的发展历程与作用原理。研究发现,宇宙辐射由银河宇宙射线、太阳高能粒子及地球辐射带三种主要来源构成,具有高能、穿透性强、难以屏蔽的特点,通过直接电离和间接自由基氧化两种机制导致DNA损伤、细胞凋亡及器官功能障碍。与化学类防辐射药物相比,天然药物特别是中药成分因其多靶点、多机制协同作用、低毒高效等特性,展现出在航天辐射防护领域的独特优势。研究总结了中药中主要抗辐射成分的化学分类及其作用机制,并探讨了其在航天辐射防护中的应用前景与挑战。**研究结果表明,中药抗辐射成分通过激活Nrf2/ARE、TLR4、Bcl-2/Bax等多条信号通路,形成全面的辐射防护体系,有望为未来深空探测任务中宇航员健康保障提供重要支持**。关键词宇宙空间辐射;辐射防护;天然药物;中药成分;航天健康一、宇宙空间辐射的危害机制1.1 宇宙辐射的来源与特性宇宙空间辐射主要来源于三个方面:银河宇宙射线(GCRs)、太阳高能粒子(SPEs)及地球辐射带。其中,GCRs是来自超新星爆发的高速带电粒子(质子占比90%),具有接近光速的能量,穿透力最强,是深空旅行最大的辐射威胁。太阳高能粒子则包括太阳风携带的质子/电子流、太阳耀斑产生的X射线与伽马射线,以及日冕物质抛射形成的高能粒子流。地球辐射带(范艾伦带)主要由地磁场捕获的超高能电子/质子构成,在低地球轨道上是航天员主要的辐射来源之一。**宇宙辐射的能量远超地面常规辐射源**,例如,GCRs中的高能重离子穿透力极强,即使在航天器的屏蔽下仍能对航天员造成显著伤害。据统计,国际空间站(ISS)宇航员每日辐射剂量超过0.5毫希沃特(mSv),是地表日常照射量的200倍,长期任务(如6个月)的总辐射剂量可达50-100 mSv。更为严峻的是,NASA预测火星任务可能面临超过1000倍于地球的辐射剂量,这将对宇航员健康构成严峻挑战。1.2 宇宙辐射对人体的损伤机制宇宙辐射对人体的损伤主要通过两种机制实现:直接电离损伤和间接自由基氧化损伤。**直接电离损伤**是指高能辐射粒子直接与生物大分子(如DNA、蛋白质等)发生相互作用,导致分子键断裂、结构破坏和功能丧失。这种损伤可造成DNA单链或双链断裂,其中DNA双链断裂(DSB)尤为危险,可能导致细胞死亡或不可逆的遗传变异。在航天员中,这种损伤主要表现为造血系统功能障碍(如白细胞减少、血小板减少)、皮肤损伤、眼底病变等。**间接自由基氧化损伤**则是宇宙辐射与人体水分子相互作用,产生大量活性氧(ROS)如羟基自由基(?OH)、超氧阴离子(O2-)等。这些自由基可攻击细胞膜、线粒体、细胞核等结构,导致脂质过氧化、蛋白质氧化修饰和DNA损伤,引发慢性氧化应激。研究表明,辐射产生的ROS会激活Nrf2/ARE信号通路,诱导细胞衰老(SASP),并抑制DNA修复相关基因(如AtRAD54、AtRAD51)的表达,从而加剧辐射损伤。此外,辐射还会导致免疫系统功能衰退,如T细胞活性异常,增加感染风险。1.3 宇航员特有的辐射健康风险长期太空辐射暴露对宇航员造成的健康风险具有特殊性,主要体现在以下方面:首先,**辐射与微重力的协同效应**会加剧损伤。研究发现,微重力环境下,辐射诱导的DNA损伤修复基因表达受到抑制,自由基(ROS)产生增加,同时会激活细胞衰老相关信号通路。这种协同效应导致航天员对辐射的耐受性显著降低,修复能力减弱。其次,**性别差异**在航天辐射防护中不容忽视。研究表明,雌性哺乳动物的辐射耐受阈值比雄性低10%-20%,这与雌激素水平降低导致的抗氧化能力下降有关。这一发现对宇航员选拔和辐射防护策略制定具有重要意义。第三,**器官特异性损伤**在航天环境中更为明显。NASA的"天地"双胞胎实验发现,长期太空辐射暴露会增加心血管疾病和神经系统紊乱的风险。辐射诱导的血管损伤和加速动脉粥样硬化过程是航天员心血管系统的主要威胁,而神经系统损伤则可能导致认知功能下降、情绪障碍和神经炎症。此外,免疫系统功能低下也是航天员面临的重要挑战,表现为T细胞活性异常,感染风险增加。二、防辐射药物的发展历程2.1 化学类防辐射药物的发展防辐射药物的研究始于20世纪40年代,经历了从早期探索到现代发展的几个关键阶段。**早期探索阶段(1940-1960年代)**:1949年,半胱氨酸首次被证实具有辐射防护作用,成为抗辐射药物研究的起点。半胱胺等含硫化合物可提供氢原子阻断氧化反应,但存在低血压等副作用,且在空气中极不稳定,口服效果差。**临床转化阶段(1970-1990年代)**:20世纪70年代,美国辉瑞公司开始研发氨磷汀(WR-2721),该药物是一种含硫的有机磷化合物,1986年在美国首次获批用于治疗乳腺癌。随后,氨磷汀逐渐扩展至多种癌症治疗,1990年代获FDA批准作为首个临床抗辐射药物,主要用于放化疗患者防护。然而,氨磷汀存在低血压、呕吐等严重副作用,且需静脉注射,限制了其广泛使用。**改良与创新阶段(2000-2020年代)**:为克服氨磷汀的局限性,研究人员开发了结构改良药物如PrC-210,实现了口服吸收并降低了消化道反应。同时,氮氧化物类化合物(如Tempol)通过清除羟自由基发挥作用,口服生物利用度达27.5%,显著提升了实用价值。此外,激素衍生物(如雌激素类似物)通过调节细胞周期增强辐射耐受性,但需在照射前72小时内给药。**应急防护阶段(2020年代至今)**:2023年,世界卫生组织(WHO)时隔16年更新了辐射应急药物清单,新增了细胞因子类药物(如G-CSF、GM-CSF)应对骨髓损伤。这些药物通过促进造血干细胞增殖分化,缓解骨髓抑制,提高受照个体的存活率。同时,稳定碘(如碘化钾)被推荐用于减少放射性碘在甲状腺的积累,普鲁士蓝则用于清除体内放射性铯和铊。2.2 天然类防辐射药物的兴起与化学类药物相比,天然类防辐射药物的研究起步较晚,但近年来发展迅速,特别是在航天医学领域。**早期探索(1980-1990年代)**:1982年,强亦忠等研究人员首次通过动物实验验证了某些市售胃肠中药对辐射的防护作用,为天然药物抗辐射研究奠定了基础。1985年,中国航天医学工程研究所张瑞钧教授团队研发出"微达康"(原名"抗微一号"),由刺五加、黄芪、女贞子等中药组成,1986年列入国家星火计划,1987年获国防科工委科技进步一等奖。该药物在临床试验中显示对放化疗患者白细胞保护总有效率达94.2%,血小板保护有效率达96.4%。**机制研究阶段(1990-2010年代)**:1990年代后,研究人员开始系统研究中药抗辐射的活性成分和作用机制。研究表明,中药中的多糖、黄酮、皂苷、酚类等成分具有显著的抗辐射活性。例如,黄芪甲苷通过调控Bcl-2/Bax蛋白表达,保护辐射损伤细胞与骨髓造血细胞;川芎嗪通过调控造血微环境提升放射损伤修复能力;红景天苷通过抗缺氧和抗辐射作用,保护辐射引起的造血功能损伤。**航天应用探索阶段(2010年代至今)**:随着中国载人航天工程的发展,天然药物特别是中药在航天辐射防护中的应用受到广泛关注。2022-2023年间,中国空间站开展了冬虫夏草、六味地黄丸等中药的微重力环境稳定性实验,验证其在太空环境中的防护潜力。2025年,中国空间站《科学研究与应用进展报告》中提到,中药复方制剂如"太空养心丸"、"强骨抗萎方"等已应用于航天员健康保障。此外,中药提取物如刺五加、天麻等活性成分在微重力环境下表现出对神经系统的保护作用,通过上调脑源性神经营养因子(BDNF)改善突触可塑性。**国际合作阶段(2020年代至今)**:2022年6月5日,中国与巴基斯坦联合开展的药用植物研发项目搭载神舟十四号载人飞船进行实验,验证了冬虫夏草等药用植物在太空环境中的活性。2023年,北京大学药学院曾克武研究员团队通过神舟十五号载人飞船搭载30余种传统中药活性分子、提取物和方剂,进行为期半年的药物稳定性实验,为后期航天中药研发提供了早期筛选和基础数据支持。三、化学类与天然类防辐射药物的比较3.1 化学类防辐射药物的特点化学类防辐射药物具有以下特点:**作用靶点明确**:化学类药物通常具有明确的作用靶点,如氨磷汀通过脱磷酸代谢产物WR-1065清除自由基,选择性保护唾液腺、骨髓等组织;Tempol通过捕获超氧阴离子降低氧化应激损伤。**研发历史长**:化学类药物研发起步早,已有多个药物通过临床试验并获FDA等监管机构批准。例如,氨磷汀于1990年代获FDA批准用于放化疗患者防护;普鲁士蓝于2003年获FDA批准用于治疗铯-137和铊辐射污染损伤。**临床应用广泛**:化学类药物在临床应用较为成熟,如氨磷汀在美国被广泛用于头颈部癌放疗前的防护,可减少X线照射引起的骨髓抑制、肾损伤等并发症。此外,稳定碘已被纳入WHO辐射应急药物清单,用于减少放射性碘在甲状腺的积累。**局限性明显**:化学类药物也存在明显局限性。首先,**毒性显著**:氨磷汀临床使用中常见的副作用包括低血压(需停药)、过敏反应、血钙降低等;半胱胺存在低血压、呕吐等副作用,且在空气中极不稳定。其次,**稳定性差**:许多化学类药物在微重力和辐射环境下容易降解。例如,模拟实验显示阿托伐他汀等药物在太空辐射环境下活性显著降低。第三,**给药途径受限**:氨磷汀需静脉注射,限制了其在航天环境中的应用便利性。第四,**作用机制单一**:多数化学类药物主要针对单一靶点,如自由基清除或DNA保护,缺乏多靶点协同作用。3.2 天然类防辐射药物的特点天然类防辐射药物,特别是中药成分,具有以下特点:**多靶点协同作用**:中药成分通常具有多靶点、多机制协同作用的特点。例如,黄芪甲苷通过调控Bcl-2/Bax蛋白表达保护骨髓造血细胞;同时,其含有的多糖成分可增强免疫功能,形成多重防护机制。这种多靶点特性使得天然药物能够全面应对辐射引起的复杂生理病理变化。**安全性高**:与化学类药物相比,天然药物通常毒性较低,副作用较少。例如,微达康对放化疗患者白细胞保护的总有效率分别为94.2%,而主要副作用为轻度胃肠不适。刺五加、天麻等中药活性成分在航天应用中显示出良好的安全性,未报告严重不良反应。**成分多样性**:中药中含有多糖、黄酮、皂苷、酚类、生物碱等多种生物活性成分,这些成分协同作用可提供更全面的防护。例如,当归多糖能显著提升辐射大鼠骨髓有核细胞、白细胞数量;而红景天苷则通过抗缺氧和抗辐射作用保护造血功能。**航天环境适应性好**:研究表明,中药成分在微重力和辐射环境下表现出较好的稳定性。例如,中国空间站实验显示,枸杞多糖在太空环境中仍能有效清除自由基,保护视网膜;当归多糖对电离辐射诱发的大鼠肠道屏障损伤具有预防作用。此外,中药复方制剂如"太空养心丸"已成功应用于航天员心血管系统保护。**临床转化案例少**:尽管天然药物在动物实验中表现出良好的抗辐射活性,但其临床转化案例相对较少。目前仅有微达康等少数中药制剂获得临床应用许可。大多数中药成分仍处于实验室研究或临床前试验阶段。3.3 天然药物在防辐射领域的优势**多靶点协同防护**:天然药物特别是中药成分能够同时作用于多个信号通路,形成协同防护效应。例如,黄芪甲苷一方面通过调控Bcl-2/Bax蛋白表达抑制细胞凋亡,另一方面通过提高SOD活性清除自由基。这种多靶点特性使得天然药物能够更全面地应对辐射引起的复杂损伤。**低毒高效特性**:中药成分通常具有较低的毒性,适合长期使用。例如,微达康对放化疗患者的白细胞保护有效率达94.2%,且主要副作用为轻度胃肠不适。相比之下,氨磷汀等化学药物需要密切监测血压变化,且部分患者需停药处理。**环境适应性强**:中药成分在微重力和辐射环境下表现出较好的稳定性。例如,中国空间站实验显示,中药提取物在太空环境中仍能保持其活性,且未观察到显著降解。这为长期航天任务中的辐射防护提供了便利条件。**整体调节能力**:中药成分具有整体调节机体功能的能力,能够同时改善航天员面临的多种航天特因环境应激。例如,刺五加、天麻等中药活性成分不仅能够保护神经系统免受辐射损伤,还能改善微重力引起的认知功能障碍和情绪失调。**航天医学需求契合**:天然药物特别是中药成分的多靶点、整体调节特性与航天医学对复合应激防护的需求高度契合。航天员面临微重力、辐射、昼夜节律紊乱等多重应激,单一靶点药物难以全面应对。中药成分的多机制协同作用能够同时保护多个系统,提高航天员的整体适应能力。四、中药抗辐射成分及作用机制4.1 中药抗辐射成分的化学分类根据化学结构特点,中药抗辐射成分可分为以下几类:**多糖类**:多糖是一类具有生理活性的天然产物,能够增强机体免疫、保护造血系统、清除自由基。常见的中药多糖成分包括:- **黄芪多糖**:通过调控PI3K/AKT通路激活Nrf2/HO-1,减轻辐射导致的肠黏膜损伤- **灵芝多糖**:通过激活TLR4信号通路促进造血功能恢复,实验显示可提升30%受照小鼠存活率- **枸杞多糖**:清除自由基,保护视网膜免受辐射损伤- **当归多糖**:显著提升辐射大鼠骨髓有核细胞、白细胞数量,改善肠道菌群失调- **红景天多糖**:增强免疫功能,减轻辐射引起的免疫抑制**黄酮类**:黄酮类化合物具有显著的抗氧化和自由基清除能力,能够保护DNA、减轻氧化损伤。常见的中药黄酮成分包括:- **黄芪总黄酮**:提高超氧化物歧化酶活性,增强机体免疫力- **大豆异黄酮**:通过激活Nrf2通路增强抗氧化酶表达,对电离辐射诱导的骨髓损伤具有防护作用- **红景天苷**:通过抗缺氧和抗辐射作用,保护造血功能- **茶多酚**:清除自由基,减轻辐射引起的氧化损伤- **川芎嗪**:调控造血微环境,提升放射损伤修复能力**皂苷类**:皂苷类化合物结构复杂,具有多靶点防护特性。常见的中药皂苷成分包括:- **人参皂苷Rb1**:通过抑制线粒体凋亡通路保护肠道上皮细胞- **三七皂苷**:改善辐射后造血功能,增加血细胞数量- **刺五加皂苷**:减轻对免疫功能的损伤,刺激造血系统功能- **黄芪甲苷**:保护骨髓造血细胞,加速粒细胞增殖与动员**酚类物质**:酚类物质具有酚羟基结构,能够有效清除自由基,减轻氧化损伤。常见的中药酚类成分包括:- **茶多酚**:对DNA的辐射损伤具有保护作用,具有较强的抗氧化能力- **阿魏酸**:显著升高辐射细胞中谷胱甘肽和烟酰胺腺嘌呤二核苷酸磷酸含量,对受辐射内皮细胞产生保护作用- **白藜芦醇**:单剂量腹腔注射可显著抑制辐射小鼠血液淋巴细胞DNA损伤- **葡萄核多酚**:对急性放射损伤具有良好防护作用,对辐射损伤引发的骨髓细胞增殖活性改变和细胞凋亡具有保护作用**生物碱类**:生物碱类成分能够有效保护造血系统,减少辐射损伤。常见的中药生物碱成分包括:- **川芎嗪**:显著升高辐射小鼠骨髓有核细胞、白细胞和红细胞数量,呈剂量依赖性- **苦参碱**:抑制小鼠骨髓细胞端粒酶活性高表达,诱导骨髓细胞凋亡,作为有效的辐射防护药物- **胡椒碱**:保护造血系统,减少辐射损伤- **骆驼蓬碱**:具有抗辐射活性,保护机体免受辐射伤害4.2 中药抗辐射的主要作用机制中药抗辐射作用机制复杂多样,主要包括以下几个方面:**自由基清除机制**:这是中药抗辐射最基础的作用机制。辐射导致的间接损伤主要源于水分子电离产生的ROS(如羟基自由基、超氧阴离子等),这些自由基可攻击细胞膜、线粒体和DNA,导致细胞损伤。中药中的多酚类(如茶多酚、阿魏酸)、黄酮类(如红景天苷、川芎嗪)等成分具有丰富的酚羟基,能够直接清除ROS,减轻氧化损伤。例如,阿魏酸可通过提供氢原子中和羟自由基,恢复生命体正常机能;茶多酚则通过清除自由基,减轻辐射引起的氧化损伤。**DNA保护与修复机制**:中药成分能够保护DNA免受辐射直接损伤,并促进受损DNA的修复。例如,黄芪甲苷对辐射损伤细胞与骨髓造血细胞具有保护作用,能使损伤的脾细胞向正常逆转;红景天苷通过抗缺氧和抗辐射作用,保护造血功能。此外,中药中的某些成分(如水熊虫提取物中的Dsup蛋白)能够直接结合DNA,减少高能粒子对DNA的损伤。**免疫调节机制**:辐射对免疫系统造成显著损伤,中药成分可通过多种途径调节免疫功能。例如,灵芝多糖通过激活TLR4信号通路增强免疫应答;黄芪甲苷提高血清及回肠组织中超氧化物歧化酶(SOD)水平,降低丙二醛(MDA)含量,减轻辐射对免疫系统的损伤。微达康的研究表明,中药复方能够提高辐射后机体的免疫功能,减少感染风险。**抗氧化应激通路激活**:中药成分能够激活内源性抗氧化应激通路,增强细胞自身抗氧化能力。**Nrf2/ARE通路是中药抗辐射最重要的机制之一**。研究表明,唐古特大黄多糖组分1(RTP1)能够减轻辐射所致肠黏膜损伤和氧化应激,且是Nrf2激活剂,能够诱发Nrf2活化转位入核,从而增强细胞自身的抗氧化能力。此外,中药成分还能够激活其他抗氧化通路,如SOD、CAT等抗氧化酶的表达。**信号通路调控**:中药成分能够调控多种细胞信号通路,减轻辐射损伤。例如,HKST激活TLR4受体后上调Bcl-2蛋白表达,使辐射诱导的细胞凋亡减少45%;β葡聚糖通过TLR2受体激活NF-κB信号通路,减轻辐射损伤。刺五加、天麻等中药活性成分通过上调脑源性神经营养因子(BDNF)改善突触可塑性,减轻辐射和失重诱导的神经损伤。**造血系统保护机制**:辐射对造血系统造成严重损伤,中药成分能够通过多种途径保护造血功能。例如,黄芪甲苷能使损伤的脾细胞向正常逆转,加速粒细胞增殖与动员作用,促进成熟迟粒细胞成熟进入储存池,从而进入外周血循环;红景天苷通过抗缺氧和抗辐射作用,对造血功能损伤具有保护效果。微达康的研究表明,中药复方能够促进60Co照射后全血5-羟色胺恢复,并对组织5-羟色胺及外周血白细胞具有保护作用。4.3 航天特殊环境下中药抗辐射效果的评估在航天特殊环境下,中药抗辐射效果受到多种因素影响,需要特别评估:**微重力与辐射的协同效应**:微重力环境会影响中药成分的吸收、分布和代谢,进而影响其抗辐射效果。研究表明,微重力可能抑制辐射旁效应诱导的DNA损伤修复基因(如AtRAD54和AtRAD51)的表达,增加ROS产生。然而,中药成分的多靶点特性可能在一定程度上抵消这种协同效应带来的负面影响。例如,刺五加、天麻等中药活性成分在微重力环境下仍能有效保护神经系统功能,通过上调BDNF改善突触可塑性。**太空环境稳定性**:中药成分在太空环境中的稳定性是航天应用的关键问题。中国空间站实验显示,中药提取物在太空环境中仍能保持其活性,且未观察到显著降解。例如,枸杞多糖在太空环境中仍能有效清除自由基,保护视网膜;当归多糖对电离辐射诱发的大鼠肠道屏障损伤具有预防作用。北京大学药学院曾克武研究员团队通过神舟十五号载人飞船搭载30余种传统中药活性分子、提取物和方剂,系统评估了其在太空环境中的稳定性,为航天中药研发提供了重要数据支持。**航天员个体化用药需求**:航天员面临复合应激(微重力、辐射、昼夜节律紊乱等),需要个体化用药方案。中药复方制剂因其整体调节特性,能够同时应对多种应激因素。例如,"太空养心丸"已成功应用于航天员心血管系统保护;"强骨抗萎方"用于预防骨质丢失;"太空燮理汤"则用于调节下丘脑-垂体-肾上腺轴和神经递质系统。这些复方制剂体现了中药在航天医学中的独特优势。五、中药在航天辐射防护中的应用前景5.1 现有航天中药应用案例目前,中药已在航天辐射防护中展现出应用潜力,部分案例已取得积极成果:**微达康的应用**:微达康是由中国航天医学工程研究所研制的抗辐射中药制剂,由刺五加、黄芪、女贞子等中药组成。该药物已在长春医科大学第一附属医院等国内八大医疗机构完成临床试验,对放化疗患者的白细胞保护总有效率超过94%,且副作用轻微,主要表现为轻度胃肠不适。这一特性使其成为航天辐射防护的潜在候选药物。**刺五加、天麻的神经保护作用**:研究表明,刺五加、天麻等中药活性成分可通过上调脑源性神经营养因子(BDNF)改善突触可塑性,减轻辐射诱导的神经损伤。脑机海河实验室团队已将这一发现应用于"太空针灸"技术,通过便携式穴位刺激装置调节航天员脑电节律及改善情绪认知,已在空间站任务中实现应用。**中药复方的综合防护作用**:中药复方如"太空养心丸"、"强骨抗萎方"等已在航天员健康保障中发挥重要作用。"太空养心丸"主要用于航天员心血管系统保护;"强骨抗萎方"则针对航天特因环境引起的骨质丢失。这些复方制剂体现了中药在航天医学中的整体调节优势。**航天育种与中药抗辐射研究**:航天育种技术为中药抗辐射研究提供了新途径。2004年,长征二号运载火箭首次搭载灵芝菌种进行太空实验;2005年,中国第22颗返回式科学与技术试验卫星搭载了肉苁蓉等13种药用植物种子,开启了中药太空育种研究的序幕。2022-2023年间,中国空间站开展了冬虫夏草、六味地黄丸等中药的微重力环境稳定性实验,验证了其在太空环境中的防护潜力。5.2 航天中药抗辐射研究的挑战与对策尽管中药在航天辐射防护中展现出巨大潜力,但仍面临一系列挑战:**航天环境适应性验证不足**:目前大多数中药抗辐射研究是在地面实验室条件下进行的,缺乏对航天特殊环境(微重力、辐射)适应性的系统验证。**对策**:加强地面模拟实验与在轨实验的结合,利用中国空间站的实验平台开展中药在微重力和辐射环境下的稳定性及药效学研究。**作用机制复杂难以阐明**:中药成分多样、作用机制复杂,难以像化学药物那样明确靶点和作用途径。**对策**:采用现代分子生物学和组学技术(如基因组学、蛋白质组学、代谢组学)系统研究中药抗辐射的作用机制,揭示其多靶点协同防护的科学内涵。**标准化与质量控制难题**:中药成分复杂、质量控制难度大,难以确保批次间的一致性。**对策**:建立中药提取物的标准指纹图谱,采用现代分析技术(如高效液相色谱、质谱联用)进行质量控制,确保航天中药的稳定性和有效性。**临床转化周期长**:中药从实验室研究到临床应用的转化周期长,成本高。**对策**:优化研究流程,建立"地面筛选-活体上行-在轨饲养-活体下行"的空间小型哺乳动物实验全流程生命支持保障和实验技术体系,加速中药抗辐射成分的筛选和验证。**航天医学标准缺乏**:目前缺乏针对航天环境的中药应用标准和指南,限制了其在航天医学中的规范化应用。**对策**:参考中国载人航天工程办公室发布的航天员健康防护指南,结合中药特点制定航天中药应用标准,明确剂量、给药途径和适用条件。5.3 未来航天中药抗辐射研究方向基于当前研究进展和航天医学需求,未来中药抗辐射研究应聚焦以下方向:**多靶点协同防护研究**:深入研究中药成分多靶点协同防护的分子机制,特别是Nrf2/ARE、TLR4、Bcl-2/Bax等信号通路的相互作用。通过系统分析中药成分在辐射环境下的基因表达谱和蛋白质互作网络,揭示其多机制协同防护的科学内涵。**航天特因环境适应性研究**:针对微重力与辐射的协同效应,开展中药成分在航天特殊环境中的药效学和药代动力学研究。利用中国空间站的实验平台,评估中药成分在微重力和辐射环境下的稳定性及作用效果,为航天中药研发提供实验依据。**中药复方优化与创新**:基于航天员面临的复合应激特点,优化现有中药复方,开发针对航天特因环境的新型中药复方。例如,结合抗辐射与抗微重力成分,开发能够同时应对多种航天环境挑战的复方制剂,提高航天员的整体适应能力。**中药制剂形式创新**:针对航天环境对药物稳定性的影响,开发新型中药制剂形式,如缓释制剂、纳米制剂等,提高中药在航天环境中的稳定性和生物利用度。同时,探索中药与航天特有技术(如"太空针灸")的结合应用,拓展中药在航天医学中的应用范围。**航天中药数据库建设**:建立航天中药成分数据库和药效学评价体系,系统整理中药抗辐射成分的化学结构、作用机制和防护效果,为航天中药研发提供数据支持。同时,开发基于人工智能的中药抗辐射成分筛选和配伍优化系统,提高研究效率和准确性。六、结论宇宙空间辐射对人体健康的危害复杂多样,主要包括直接电离损伤和间接自由基氧化损伤两种机制。**辐射通过破坏DNA、蛋白质等生物大分子,导致细胞死亡或遗传变异,同时与微重力环境产生协同效应,进一步加剧航天员的健康风险**。在航天特因环境下,辐射不仅影响造血系统,还对心血管系统、神经系统和免疫系统造成显著损伤,且雌性航天员的辐射耐受性较雄性低10%-20%,这些特点对辐射防护策略提出了更高要求。防辐射药物经历了从早期化学类药物(如半胱氨酸、氨磷汀)到现代天然类药物(特别是中药成分)的演进。化学类药物虽然作用靶点明确、研发历史长、临床应用广泛,但存在毒性显著、稳定性差、给药途径受限等局限性,难以满足长期航天任务的需求。相比之下,**天然药物特别是中药成分以其多靶点协同作用、低毒高效特性、环境适应性强等优势,逐渐成为航天辐射防护的研究热点**。中药抗辐射成分主要包括多糖类(如黄芪多糖、灵芝多糖)、黄酮类(如黄芪总黄酮、红景天苷)、皂苷类(如人参皂苷Rb1、三七皂苷)、酚类(如茶多酚、阿魏酸)和生物碱类(如川芎嗪、苦参碱)等。这些成分通过自由基清除、DNA保护与修复、免疫调节、抗氧化应激通路激活、信号通路调控和造血系统保护等多种机制协同作用,形成全面的辐射防护体系。其中,**Nrf2/ARE通路是中药抗辐射最重要的机制之一**,通过激活这一通路,中药成分能够增强细胞自身的抗氧化能力,减轻辐射引起的氧化应激。在航天特殊环境下,中药抗辐射研究已取得初步成果。微达康、刺五加、天麻等中药成分已在临床试验或航天实验中显示出良好的抗辐射效果。中药复方如"太空养心丸"、"强骨抗萎方"等也已在航天员健康保障中发挥重要作用。然而,中药在航天辐射防护中的应用仍面临航天环境适应性验证不足、作用机制复杂难以阐明、标准化与质量控制难题、临床转化周期长以及航天医学标准缺乏等挑战。未来,中药抗辐射研究应聚焦多靶点协同防护、航天特因环境适应性、中药复方优化与创新、中药制剂形式创新以及航天中药数据库建设等方向。**通过系统研究中药成分在航天特殊环境中的稳定性和作用机制,结合现代生物技术和人工智能技术,有望开发出适用于长期航天任务的天然抗辐射药物,为航天员健康提供全面保障**。七、展望随着中国载人航天工程的快速发展和深空探测任务的推进,航天员面临的辐射风险将日益增加。中药抗辐射研究作为航天医学的重要组成部分,具有广阔的发展前景。一方面,中药成分的多靶点、整体调节特性与航天医学对复合应激防护的需求高度契合。通过深入研究中药成分在航天特殊环境中的作用机制,有望开发出能够同时应对多种航天环境挑战的抗辐射药物,提高航天员的整体适应能力。另一方面,航天特殊环境为中药研究提供了独特的平台。利用空间站的实验装置,可以在真实微重力和辐射环境下评估中药成分的稳定性和药效学特点,为中药现代化研究提供新的思路和方法。同时,航天育种技术也为中药抗辐射研究提供了新途径,通过太空环境诱导中药植物产生耐辐射突变株,可为开发新一代抗辐射中药提供遗传资源。此外,**中医药与航天医学的深度融合将推动航天医学"中国方案"的形成**。脑机海河实验室团队提出的"在轨辨证评估体系"结合人工智能技术,有望实现中药在航天环境中的精准应用。天津中医药大学与航天联合实验室的建立,以及《中国空间站科学研究与应用进展报告》中对中药复方制剂的肯定,都表明中医药在航天医学领域的重要性日益凸显。总之,中药抗辐射研究不仅对保障航天员健康具有重要意义,也将为航天事业的发展提供重要支持。通过加强基础研究、技术创新和标准建设,中药有望成为未来深空探测任务中航天员辐射防护的重要选择,为人类探索宇宙的征程保驾护航。
2026-02-16
·同写意
100+全球先锋领袖现场开讲、50+顶尖新基建机构抢先入驻、1000+产业精英面对面链接,中国ADC和核药产业界规格最高、影响力最大、汇聚创新力量最全的年度品牌盛会!
抗体偶联药物(ADC)通过将高特异性单克隆抗体与强效细胞毒性药物共价连接,实现了对肿瘤细胞的靶向杀伤。连接抗体与药物分子的“接头”(Linker)不仅发挥着简单的物理连接功能,更是决定ADC药代动力学特性、治疗效果和临床安全性的关键要素。
今天为大家解读一篇关于ADC药物linker相关的文章。包括linker设计的化学基础、结构分类、作用机制及其对ADC生物学行为的影响,以期为大家在ADC药物linker的理性设计与临床开发提供全面的专业参考。
TONACEA
01
ADC linker的基本概念与设计原则
01
ADC的结构组成与linker功能定位
抗体-药物偶联物(ADC)由三个核心组成部分构成:
√ 单克隆抗体:提供对肿瘤细胞表面抗原的特异性识别能力;
√ 细胞毒性药物(“弹头”或“载荷”):发挥高效的肿瘤细胞杀伤作用;
√ Linker:共价连接抗体与药物分子,形成稳定的偶联复合物。
Linker的功能超越了单纯的物理连接,需要满足以下关键要求:
1.保持化学稳定性:在药物制剂储存过程中保持结构完整性;
2.维持血浆稳定性:在系统循环期间抵抗酶解、水解等降解过程;
3.实现靶向释放:在肿瘤细胞内快速、有效地释放活性药物分子;
4.调控药代动力学:影响ADC的分布、代谢和清除行为。
02
Linker设计的基本要求:稳定性与释放性的平衡
理想的ADC linker必须在系统稳定性与细胞内释放效率之间达到精确平衡:
• 系统稳定性需求:
√ ADC在循环中保持完整;
√ 避免药物在血浆中过早释放导致的系统毒性;
√ 维持足够长的循环半衰期以确保肿瘤组织充分暴露。
• 细胞内释放需求:
√ ADC被肿瘤细胞内吞后,接头能够快速响应细胞内环境;
√ 高效释放活性药物分子;
√ 释放机制具有肿瘤特异性,减少对正常细胞的影响。
图1. ADC的结构示意图,展示了抗体、Linker和细胞毒性药物的空间排列关系。
TONACEA
02
Linker连接位点的化学基础与工程策略
01
传统连接位点的化学特性• 赖氨酸(Lysine)残基偶联
√ 化学基础:赖氨酸残基的ε-氨基具有高亲核性;
√ 常用连接化学:N-羟基琥珀酰亚胺酯(NHS ester)与氨基反应形成酰胺键;
√ 位点特征:
①每个IgG抗体分子含有约80-100个可及赖氨酸残基
②反应生成异质性偶联产物,载药量分布范围广(0-8个药物分子/抗体)
③每个批次的载药量分布遵循二项分布规律,但批次间具有良好的重现性
• 半胱氨酸(Cysteine)残基偶联
√ 化学基础:通过还原链间二硫键产生反应性硫醇基团;
√ 常用连接化学:硫醇与马来酰亚胺基团通过Michael加成反应形成硫醚键;
√ 位点特征:
①IgG抗体含有4个链间二硫键(重链-重链、重链-轻链),还原后可生成8个游离硫醇
②较赖氨酸偶联具有更高的位点特异性
③仍会产生一定程度的异质性产物
图2. ADC的质谱分析图,展示了美登素类ADC的载药量分布,平均DAR=3.5)
02
工程化连接位点的创新策略• Thiomab技术:工程化半胱氨酸定点偶联
√ 技术原理:通过基因工程在抗体特定位置引入半胱氨酸残基
√ 技术流程:
①抗体表达过程中,工程化半胱氨酸的巯基与培养基中的胱氨酸形成混合二硫键;
②偶联前通过还原反应生成游离硫醇;
③部分氧化以恢复对结构完整性重要的链内和链间二硫键;
④工程化半胱氨酸保持硫醇形式,用于与马来酰亚胺接头反应。
√ 技术优势:
①实现高度均一的定点偶联;
②控制每个抗体的药物分子数;
③改善药代动力学特性的可预测性。
• 糖基化位点工程
√ 传统方法:通过高碘酸钠氧化抗体Fc区的N-连接糖链,生成醛基后与含肼基接头反应;
局限性:可能引起非特异性蛋白质氧化
√ 现代策略:
①代谢标记:在含叠氮糖(如Ac4ManNAz或Ac4GalNAz)的培养基中表达抗体,将叠氮基引入糖链;
②化学重塑:利用β(1,4)-半乳糖基转移酶将含叠氮基的糖单元引入抗体糖链
③点击化学偶联:叠氮基与张力环炔烃(如环辛炔)进行无铜点击化学反应;
√ 优势:提供高特异性连接位点,不影响内源性氨基酸的反应性。
• 非天然氨基酸插入
√ 实现方法:
①正交tRNA/氨酰tRNA合成酶系统:使用能够被非天然氨基酸酰化的正交tRNA;
②无细胞表达系统:包含特定tRNA和相应氨酰tRNA合成酶的无细胞提取物。
√ 偶联化学:
①点击化学:含叠氮基的非天然氨基酸与含环炔烃的接头反应;
②肟连接:含羟氨基的非天然氨基酸与含醛基的接头反应;
√ 应用实例:Axup等(2012)通过该方法制备了均质化抗Her2偶联物。
• 转谷氨酰胺酶(Transglutaminase)介导的偶联
√ 酶学基础:微生物转谷氨酰胺酶催化谷氨酰胺侧链与伯胺形成共价键;
√ 天然位点利用:人IgG的Q295位谷氨酰胺(通常被N297糖基化阻碍可及性);
√ 工程化策略:引入谷氨酰胺标签序列,扫描抗体表面最适反应位点;
√ 应用:实现高度特异性和均一性的抗体偶联。
TONACEA
03
ADC药物linker的分类与作用机制
01
不可裂解Iinker• 硫醚类linker的化学特性
√ 马来酰亚胺-硫醇化学:在中性水相条件下高效反应形成稳定硫醚键;
√ 替代化学:硫醇与卤代乙酰胺基团反应(需较高pH和过量试剂);
√ 应用药物:主要与微管蛋白抑制剂连接,如美登素(maytansine)和奥瑞他汀(auristatin)衍生物。
图3.美登素类衍生物结构
图4. 抗体-SMCC-DM1的偶联方案
• 代表性不可裂解linker结构
SMCC接头(N-琥珀酰亚胺-4-(马来酰亚胺甲基)环己烷-1-羧酸酯)。
√ 应用:Ado-曲妥珠单抗-美坦新(T-DM1,Kadcyla®)的偶联化学。
√ 结构特征:含疏水环己基核心。
√ 偶联机制:
①NHS酯端与抗体赖氨酸残基反应;
②马来酰亚胺端与DM1的硫醇基反应;
√ 代谢产物:Lys-SMCC-DM1(唯一代谢物)。
图5. 抗体- SMCC -DM1生成单一代谢物Lys- SMCC -DM1
PEG4Mal linker
√ 结构改进:用亲水性四乙二醇(PEG4)替代SMCC的疏水环己基;
√ 药理学优势:在表达P-糖蛋白(MDR表型)的细胞中显示更高活性;
√ 代谢产物:Lys-PEG4Mal-DM1;
√ 类似应用:BMPEO(双马来酰亚胺三氧乙二醇)接头,含PEG3间隔基。
图6. 抗体-PEG₄-Mal-DM1的偶联方案
• 不可裂解linker的细胞内代谢途径
√ 释放机制:需要抗体在溶酶体中完全降解;
√ 代谢产物特征:保留连接氨基酸残基(如赖氨酸);
√ 细胞渗透性:通常较低,限制“旁观者效应”。
• 临床实例:抗CD70-mc-MMAF
√ 接头类型:马来酰亚胺-己酰(mc)不可裂解接头;
√ 载药量:平均每个抗体连接4个MMAF分子;
√ 结构改进:用溴乙酰胺-己酰(bac)替代mc接头。
①提高血浆稳定性;
②增加25%的肿瘤内药物暴露;
③但在786-O肾细胞癌异种移植模型中,疗效无统计学显著差异。
图7. 抗体-mc-MMAF的结构示意图
02
可裂解linker• 酸不稳定linker
1)腙键(Hydrazone)linker
√ 设计原理:利用肿瘤细胞内吞体和溶酶体的酸性环境(pH4-5);
√ pH敏感性:中性pH(血浆)稳定,酸性pH快速水解;
√ 应用药物:主要与DNA靶向药物连接,如加利车霉素(calicheamicin)和阿霉素(doxorubicin)。
图8. calicheamicin和doxorubicin的结构
2)临床实例分析
吉妥珠单抗奥佐米星(Gemtuzumab ozogamicin,Mylotarg®)
√ 靶点:CD33(急性髓系白血病);
√ 药物:N-乙酰-γ₁-加利车霉素;
√ 接头:4-(4'-乙酰苯氧基)-丁酸双功能接头;
√ 载药量:每个抗体4-6个加利车霉素分子,50%抗体未偶联。
√ 释放机制:
①酸性环境中腙键水解;
②硫醇介导还原;
③Masume-Bergman环化反应;
④生成苯炔双自由基,导致DNA链断裂。
√ 临床历程:最初获批后因确认性III期临床试验未显示临床获益而自愿撤市,现以分次给药方案重新评估。
图9. 通过腙键接头连接的抗体-加利车霉素的释放和作用机制
伊妥珠单抗奥佐米星(Inotuzumab ozogamicin,CMC-544)
√ 靶点:CD22(B淋巴细胞恶性肿瘤);
√ 采用相同酸不稳定接头:4-(4'-乙酰苯氧基)-丁酸;
√ 临床评估:因总生存期无获益而暂停治疗CD22阳性侵袭性非霍奇金淋巴瘤的评估。
3)酸不稳定linker的局限性
√ 缺乏特异性:可在任何酸性环境中裂解,包括正常组织;
√ 可能过早释放:导致系统毒性增加;
√ 依赖pH梯度:肿瘤微环境的pH异质性可能影响释放效率。
• 可还原linker
1)二硫键linker的设计基础
√ 还原电位差异:血浆与细胞内还原环境的显著差异;
√ 血浆还原剂:半胱氨酸(~5μM),白蛋白硫醇(~0.6mM但不可及);
√ 细胞内还原剂:还原型谷胱甘肽(1-10mM),蛋白质二硫键异构酶家族;
√ 设计目标:血浆中稳定,细胞内高效还原。
2)二硫键稳定性的调控策略
√ 空间位阻效应:通过甲基化增加二硫键周围的空间位阻;
√ 研究模型:HuC242抗体(靶向CanAg抗原);
√ 关键发现:
①二硫键两侧各一个甲基(总计两个甲基)比单侧两个甲基提供更高稳定性;
②甲基化位置对稳定性影响显著;
③稳定性与ADC体内药代动力学特性良好相关。
图10. 甲基位阻增加对二硫键稳定性的影响
3)二硫键还原的细胞定位
√ 理论预测:还原反应在胞质(pH 7.4)比在溶酶体(pH 5)更有效。
√ 实验证据:某些细胞系中,内吞体中即发生一定程度的二硫键还原。
①提供溶酶体外的替代代谢途径;
②可能对细胞毒性效应产生优势;
√ 示踪研究:与FRET荧光团通过二硫键连接的叶酸在内吞体中发生显著还原。
图11. 抗体- SPDB -DM4的代谢
4)二硫键linker的优势
√ 释放中性代谢物:可自由扩散进入邻近细胞;
√ 旁观者效应:增强实体瘤穿透和杀伤效率;
√ 肿瘤特异性:利用肿瘤细胞内高还原环境。• 肽类 linker
1)肽类 linker 的设计优势
√ 单键裂解机制:肽键水解即可释放药物(对比不可裂解接头需双键裂解);
√ 亲水性调节:通过氨基酸选择提高接头亲水性;
√ 酶特异性:利用肿瘤细胞高表达的蛋白酶(如组织蛋白酶B)。
2)代表性肽 linker 结构
Val-Cit-PABC-MMAE系统
结构组成:缬氨酸-瓜氨酸(Val-Cit)二肽 + 对氨基苄基氨基甲酸酯(PABC)"自毁"间隔基。
图12. mc-Val-Cit-PABC-MMAE结构
√ 裂解酶:溶酶体蛋白酶(如组织蛋白酶B)
√ 释放机制:
①Val-Cit肽键被蛋白酶水解;
②释放PABC-MMAE中间体;
③PABC自毁,生成游离MMAE。
√ 扩散特性:MMAE可扩散至邻近细胞,产生旁观者效应。
图13. mc-Val-Cit-PABC-MMAE的代谢和“自毁”过程
3)临床成功案例:Brentuximab Vedotin(Adcetris™)
√ 靶点:CD30(霍奇金淋巴瘤,间变性大细胞淋巴瘤);
√ 接头结构:mc-Val-Cit-PABC;
√ 临床优势:相比通过5-苯甲酰戊酸-AE酯腙连接的MMAE偶联物,显示更好的稳定性和特异性。
图14. Brentuximab vedotin的结构
4)肽类 linker 的变体研究
√ Phe-Lys-PABC linker 与Val-Cit接头比较:
①在小鼠和人类血浆中稳定性较低;
②预计半衰期:小鼠血浆12.5天 vs 30天(Val-Cit);
③预计半衰期:人类血浆80天 vs 230天(Val-Cit)。
5)肽类 linker 的亲水性改造
√ 应用需求:连接疏水性药物,如DNA小沟结合剂(MGB)
√ 设计策略:
缬氨酸-赖氨酸-四乙二醇接头;
缬氨酸-赖氨酸-对氨基苄醚自毁间隔基;
√ 优势:避免聚集形成,提高载药量。
• β-葡萄糖醛酸 linker
1)酶响应机制
√ 裂解酶:β-葡萄糖醛酸酶(β-glucuronidase);
√ 酶分布:溶酶体中丰富,某些肿瘤中过表达;
√ 临床相关性:人乳腺肿瘤和瘤周组织的主要药物代谢酶系统。
2)结构特点与应用
√ 应用药物:MMAE、MMAF、阿霉素丙基噁唑;
√ 靶点抗体:c1F6(抗CD70)、cAC10(抗CD30);
√ 亲水性优势:
允许高载药量(可达8个药物分子/抗体);
适用于连接疏水性药物(如DNA小沟结合剂);
生成单体ADC,避免聚集。
3)代谢途径
图15. 抗体-β-葡萄糖醛酸-细胞毒素的代谢
TONACEA
04
Linker对ADC生物学行为的调控机制
理想情况下,ADC药物具有前药特性:其设计为在血浆系统循环中保持非活性状态,但在细胞内化后被激活。
图16. 抗体-药物偶联物的细胞内加工
01
Linker与ADC的细胞内加工和活化• ADC的细胞内旅程
①表面结合:ADC特异性结合肿瘤细胞表面抗原;
②内吞作用:网格蛋白介导的囊泡形成;
③内吞体转运:pH逐渐下降至5-6;
④溶酶体融合:pH进一步降至4,富含蛋白水解酶和酯酶;
⑤ADC降解:生成由细胞毒性药物共价连接于氨基酸残基的代谢物。
• Linker 设计对活化速率的影响
√ 肽类接头的优势:单键裂解即可释放药物,活化速率快;
√ 酶分布考虑:组织蛋白酶B同时存在于内吞体和溶酶体。
导致每个抗体递送的药物量减少;
可能引起正常组织中毒素的早期释放;
潜在优势:允许更快速的活化;
潜在风险:可能在抗体通过FcRN回收过程中在内吞体裂解。• 酸不稳定 Linker 的特异性问题
√ 机制限制:依赖pH而非酶特异性;
√ 潜在风险:在任何酸性环境中都可能裂解,包括正常组织;
√ 毒性关联:与游离药物的膜渗透性相关。
02
Linker设计克服多药耐药(MDR)• MDR的机制与挑战
主要介导者:多药转运蛋白MDR1(P-糖蛋白);
√ 功能机制:ATP依赖性药物外排泵;
√ ADC药物底物:包括加利车霉素、阿霉素、紫杉烷类、美登素类、海兔毒素类似物。• Linker 设计策略规避MDR
1. 亲水性linker的优势
√ PEG4Mal-DM1:相比SMCC-DM1在MDR阳性细胞中活性更高;
√ 作用机制:亲水性接头产生的代谢物(Lys-PEG4Mal-DM1)不易被MDR1识别和外排。
2. 磺酸化SPDB linker
√ 结构改进:在SPDB接头中引入远端磺酸基;
√ 应用实例:抗EpCAM-sulfo-SPDB-DM4;
√ 活性比较:在COLO 205MDR细胞系和异种移植模型中优于非磺酸化版本。
3.研究模型验证
√ 细胞模型:
HCT-15(结肠腺癌,天然高表达MDR1);
UO-31(肾腺癌,天然高表达MDR1);
COLO 205MDR(工程化高表达MDR1)。
√ 实验结果:
MDR1导致对微管蛋白抑制剂的敏感性降低6-18倍;
亲水性 linker 可部分克服MDR表型。
图17. 规避MDR耐药的Sulfo-SPDB接头结构
• 临床开发中的ADC
√ IMGN853:靶向叶酸受体,sulfo-SPDB-DM4连接;
√ 适应症:卵巢癌、非小细胞肺癌。
03
Linker增强实体瘤活性的机制• 实体瘤的治疗挑战
√ 结构复杂性:异质性组织构成;
√ 靶抗原表达不均:限制可靶向的癌细胞群体;
√ 肿瘤穿透障碍:
抗体类疗法分子量大;
肿瘤内血管分布不均;
即使均匀表达的抗原,抗体结合也呈斑片状分布。
• “旁观者效应” linker 的设计
√ 定义:释放可扩散的游离药物,杀伤邻近不表达靶抗原的细胞。
√ 实现机制:
可还原接头:释放中性、可扩散代谢物;
肽接头与自毁间隔基:生成细胞膜可渗透的毒素;
√ 体内优势:
补偿不完全的肿瘤穿透;
杀伤异质性肿瘤细胞群体;
在实体瘤模型中显示出更好的体内活性;
√ 体外-体内差异:旁观者效应的优势在体外模型中不一定明显。
04
Linker 的血浆稳定性机制• 硫醚键的稳定性问题
逆Michael反应(Retro-Michael Reaction)
√ 现象:硫醇-马来酰亚胺键在血浆中可能发生交换反应;
√ 交换对象:与血清白蛋白的游离硫醇基团交换;
√ 后果:导致ADC失活和潜在的系统毒性。
位点依赖的稳定性差异
√ 溶剂暴露位点(如Fc区的S396C):更易发生逆Michael反应。
√ 带正电环境位点(如轻链的V205C):稳定性更高。
√ 机制差异:
溶剂暴露环境中:易发生马来酰亚胺交换;
带正电环境中:马来酰亚胺环打开,形成稳定的琥珀酰亚胺衍生物,防止交换。
• 美登素类 linker的特殊性
√ 化学相似性:同样使用硫醇-马来酰亚胺化学;
√ 稳定性差异:对逆Michael反应抵抗性更强;
√ 可能机制:DM1硫醇供体的pKa较高;
√ 释放机制研究:游离DM1主要通过β-消除释放,随后硫醚键氧化为亚砜;
体外人工假象:硫醚氧化在体内不太可能发生,因血浆氧化还原电位受严格调控。• 二硫键linker的稳定性调控
√ 位阻效应:甲基化程度与体外对DTT还原的抗性正相关;
√ 药代动力学关联:体外稳定性与体内PK特性良好相关;
√ 实例比较:
huC242-SPDB-DM4(二甲基):较长血浆半衰期;
huC242-SPDB-DM3/huC242-SPP-DM1(单甲基):较短血浆半衰期。
05
Linker 稳定性与ADC活性的复杂关系• 稳定性-活性的平衡原理
核心原则:最高稳定性的接头不一定产生最佳疗效。
示例分析1:SPP-DM1 vs SPP-DM4
√ 稳定性比较:SPP-DM4:三个甲基位阻,体外稳定性更高,血浆半衰期218小时;SPP-DM1:一个甲基位阻,血浆半衰期47小时;
√ 暴露量比较:SPP-DM4的AUC(22,712 h·μg/mL)显著高于SPP-DM1(5,186 h·μg/mL)。
√ 疗效对比:huC242-SPP-DM1在COLO 205和HT-29异种移植模型中疗效更好。
√ 机制解释:SPP-DM1在细胞内更易释放活性代谢物,更高的暴露量无法补偿更慢的细胞内活化速率。
示例分析2:整合素靶向偶联物
√ CNTO365:SPDB-DM4(双甲基位阻);
√ CNTO366:SPP-DM4(三甲基位阻);
√ 疗效比较:在HT-29结肠癌和A-549肺癌模型中,CNTO365疗效更好。
示例分析3:T-DM1 vs T-SPP-DM1
√ 稳定性比较:T-DM1(T-SMCC-DM1)血浆稳定性更好,半衰期更长;
√ 肿瘤代谢物积累:尽管清除更快、肿瘤定位更少,T-SPP-DM1在肿瘤中生成的代谢物量与T-DM1相似;
√ 疗效结果:在BT474-EEI异种移植模型中两者疗效相似;
√ 作用机制差异:T-SPP-DM1具有旁观者活性,而T-DM1没有。
图18. 曲妥珠单抗-SPP-DM1(T-SPP-DM1)与T-DM1(T- SMCC -DM1)的药代动力学特性及代谢物在BT474 EEI 来源的肿瘤异种移植模型中的蓄积
• 疗效预测的多因素考量
√ 药代动力学特性:血浆稳定性、清除率、分布容积;
√ 肿瘤内激活速率:决定活性代谢物的生成速度;
√ 杀伤模式:直接毒性、旁观者效应;
√ 经验性测试的必要性:每个ADC必须独立评估上述因素的综合效应。
06
Linker对ADC安全性的影响机制• Linker与肝脏解毒作用
肝脏的核心作用
√ 主要代谢场所:抗体代谢的主要部位;
√ ADC代谢物浓度:在肝脏中显著积累。
研究模型:huN901-美登素类偶联物
√ 放射性标记:氚标记美登素的C-20甲氧基;
√ 研究目的:评估接头对ADC解毒的影响。
解毒机制分析
(1)不可裂解接头(SMCC-DM1)
√ 代谢产物:Lys-SMCC-[³H]DM1;
√ 毒性降低:因细胞渗透性差,细胞毒性降低50倍以上。
(2)可裂解二硫键接头(SPP-DM1,SPDB-DM4)
√ 初级代谢物:赖氨酸连接的美登素类类似物;
√ 次级代谢途径:肝脏中的还原、S-甲基化、NADPH依赖性氧化;
√ 终产物:S-甲基亚砜和S-甲基砜衍生物;
√ 毒性降低:5-50倍(多种人类癌细胞系测试)。
图19. 抗体-鹅膏蕈素偶联物的肝脏解毒作用
• Linker对ADC生物分布的影响
研究方法
√ ¹²⁵I标记抗体骨架:追踪ADC整体组织分布;
√ ³H标记美登素C-20甲氧基:追踪药物部分的分布。
关键发现
①抗体主导分布:偶联与未偶联抗体的生物分布特征相似;
②肝胆消除途径:30-50%注射剂量在给药后2小时至2天内在胆囊回收;
③种属一致性:T-[³H]DM1在大鼠中,7天内80%放射性经粪便排泄;
④接头影响有限:不同接头(SMCC、SPDB)的ADC分布特征相似。
安全性差异的来源
√ 主要因素:代谢物的细胞渗透性和效力;
√ 次要因素:接头依赖的组织分布差异有限。• 临床安全性观察:稳定性与耐受性的负相关
临床药代动力学特征
√ 影响因素:接头稳定性 + 抗原介导的清除;
√ 美登素类偶联物:人体PK与接头对DTT还原的敏感性和临床前动物PK一致。
Cantuzumab案例研究
√ 相同抗体:抗huC242(靶向CanAg)。
√ 不同接头比较:
Cantuzumab mertansine(huC242-SPP-DM1):轻度位阻二硫键,人体血浆半衰期2天;
Cantuzumab ravtansine(huC242-SPDB-DM4):高度位阻二硫键,半衰期4.6天。
√ PK差异解释:反映了SPP-DM1和SPDB-DM4在CanAg背景下的稳定性差异
其他ADC比较
√ SAR3419(huB4-SPDB-DM4):靶向CD19,半衰期7.9天;
√ Ado-曲妥珠单抗-美坦新(T-DM1):半衰期4.4天,主要反映抗原介导的清除。
最大耐受剂量(MTD)的linker相关性
关键观察:稳定性-耐受性负相关
√ 化学稳定性排序:SMCC > SPDB > SPP(SMCC最稳定,SPP最不稳定);
√ 临床耐受性排序:SPP > SPDB > SMCC(SPP MTD最高,SMCC最低)。
√ 潜在机制:
稳定接头:缓慢释放,持续低水平毒性积累;
不稳定接头:快速清除,急性但可逆毒性。
需要进一步研究的临床问题
①这种负相关是否适用于所有美登素类偶联物(可能产生不同的小分子量代谢物);
②类似趋势是否存在于其他细胞毒性载荷中。
图20. 接头稳定性与人体最大耐受剂量的负相关关系
TONACEA
05
Linker设计的临床转化考量与未来方向
01
从临床前到临床的转化挑战• 预测性的局限性
√ 体外-体内相关性:体外稳定性测试不能完全预测体内行为;
√ 种属差异:动物模型与人类在代谢、分布和清除方面的差异;
√ 肿瘤微环境模拟:临床前模型难以完全复制人类肿瘤的复杂性。• 个体化差异考虑
√ 肿瘤异质性:不同肿瘤类型、同一肿瘤不同区域的差异;
√ 患者间变异:代谢酶表达、肝脏功能、免疫状态差异;
√ 药物相互作用:ADC与合并用药的潜在相互作用。
02
生产工艺与质量控制挑战• 均质化ADC的生产
√ 工程化策略:定点偶联技术(Thiomab、非天然氨基酸等)的可扩展性;
√ 工艺复杂性:多步反应、纯化策略、分析验证;
√ 成本效益:复杂生产工艺对最终产品成本的影响。
• 质量控制要点
√ 载药量分布:即使使用定点偶联技术,仍需严格控制;
√ 聚集倾向:特别是高载药量或疏水性接头的情况;
√ 稳定性监控:制剂稳定性、运输稳定性、体内稳定性。
03
Linker设计的未来发展方向• 智能响应型接头
√ 多重触发机制:pH、酶、还原电位、温度等组合响应;
√ 肿瘤微环境特异性:响应缺氧、特定蛋白酶过表达、异常代谢物;
√ 时空控制释放:提高肿瘤特异性,最小化正常组织暴露。• 连接化学的创新
√ 更稳定的连接键:抵抗逆Michael反应等降解途径;
√ 正交反应化学:提高偶联选择性和效率;
√ 模块化设计:便于不同抗体-药物组合的快速开发。• 多学科融合策略
√ 化学-生物学整合:理性设计结合高通量筛选;
√ 计算模拟辅助:预测接头稳定性、代谢途径和毒性特征;
√ 临床反馈驱动:基于真实世界证据的迭代优化。
04
临床开发策略建议• 早期评估重点
√ 接头稳定性全面评估:体外稳定性、血浆稳定性、靶细胞释放效率;
√ 代谢产物表征:明确主要代谢物及其药理活性;
√ 毒性特征早期识别:特别是肝脏毒性和血液学毒性。
• 临床研究设计考量
√ 剂量探索策略:考虑接头稳定性对MTD的影响;
√ 药代动力学-药效学关系:建立接头特性与临床效应的关联;
√ 生物标志物开发:识别可能预测疗效或毒性的生物标志物。
• 监管考量
√ 质量控制标准:接头相关特性的质量控制策略;
√ 可比性研究:生产工艺变更对ADC特性的影响评估;
√ 风险评估:基于接头特性的安全性风险管理计划。
TONACEA
06
总结与展望
01
Linker在ADC开发中的核心地位
ADC linker已从简单的化学连接工具演变为多功能、智能化的关键模块,其设计需要综合考虑:
• 化学稳定性要求:在制剂储存和系统循环中保持完整性;
• 生物学响应性:在靶细胞中高效、特异地释放活性药物;
• 代谢可控性:生成具有适当药理学特性的代谢物;
• 安全性谱:最小化正常组织毒性,最大化治疗指数;
• 生产工艺可行性:支持规模化、可重复的生产。
02
Linker 设计的核心平衡原则
成功的ADC linker 设计必须在多个相互制约的因素间找到最佳平衡点:
• 系统稳定性 vs 细胞内释放效率;
• 肿瘤特异性 vs 释放机制可靠性;
• 药代动力学优化 vs 药效学最大化;
• 疗效增强 vs 毒性控制。
03
Linker 技术的未来影响
随着对接头-药物-抗体相互作用的深入理解和技术进步,新一代ADC接头有望:
• 提高治疗指数:通过更精确的肿瘤特异性释放;
• 克服耐药性:通过智能设计规避MDR等耐药机制;
• 扩展适应症:特别是实体瘤治疗的改进;
• 实现个性化治疗:根据肿瘤特征定制接头特性。
04
对ADC开发者的建议
• 早期投入接头研究:将接头设计作为ADC开发的核心组成部分;
• 综合评估策略:结合化学、生物学和药理学方法全面评估接头特性;
• 临床转化思维:从开发早期就考虑临床可行性和监管要求;
• 迭代优化理念:基于不断积累的知识和技术进步持续改进接头设计。
ADC linker设计的进步将继续推动抗体-药物偶联物领域的发展,为癌症患者提供更有效、更安全的治疗选择。随着多学科合作的深化和技术创新的加速,我们有理由期待更智能、更精准的ADC接头技术在未来癌症治疗中发挥越来越重要的作用。
抗体药物偶联物细胞疗法
100 项与 盐酸半胱氨酸 相关的药物交易
登录后查看更多信息
研发状态
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| 营养紊乱 | 美国 | 2019-04-16 | |
| 寻常痤疮 | 日本 | 1987-05-23 | |
| 药物性皮炎 | 日本 | 1987-05-23 | |
| 湿疹 | 日本 | 1987-05-23 | |
| 渗出性红斑 | 日本 | 1987-05-23 | |
| 白细胞减少 | 日本 | 1987-05-23 | |
| 荨麻疹 | 日本 | 1987-05-23 | |
| 膳食补充剂 | 美国 | 1986-10-22 | |
| 肿瘤 | 中国 | 1982-01-01 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
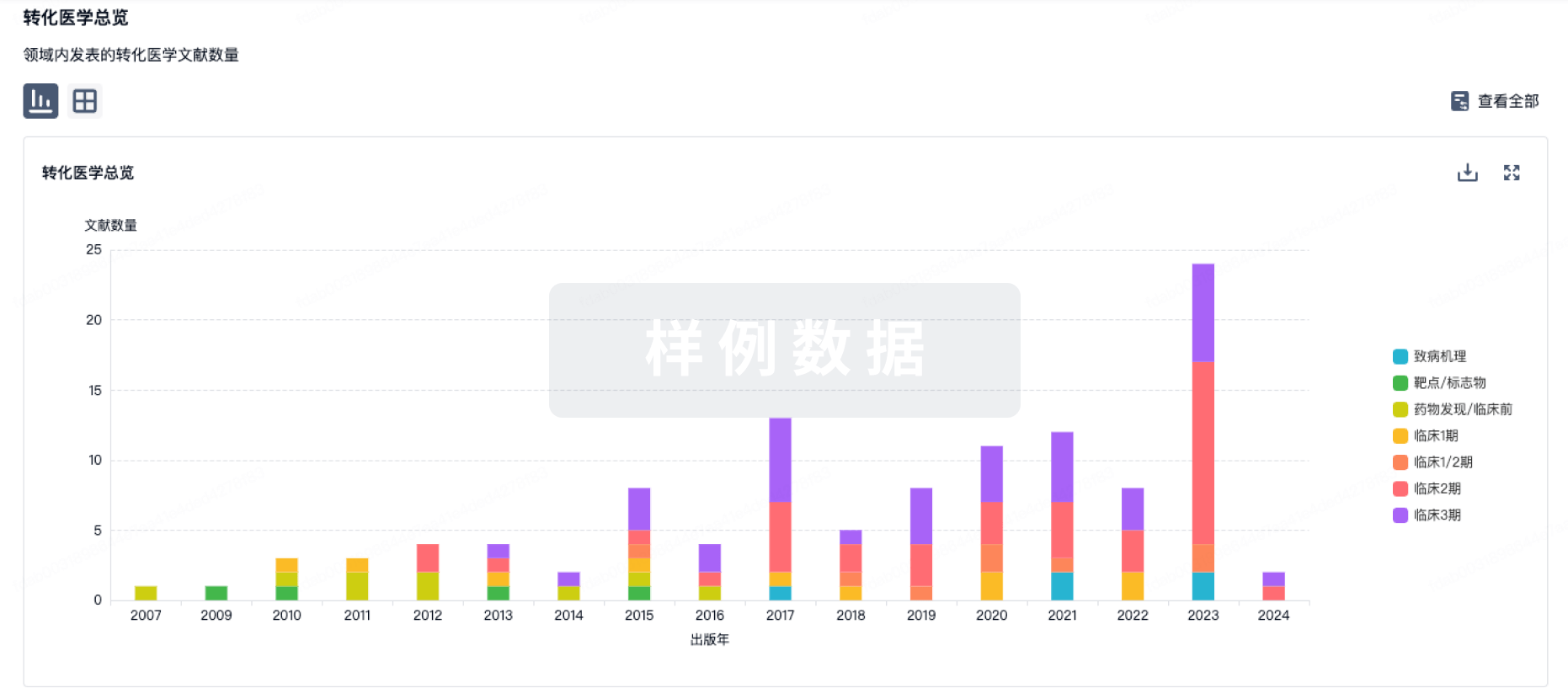
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
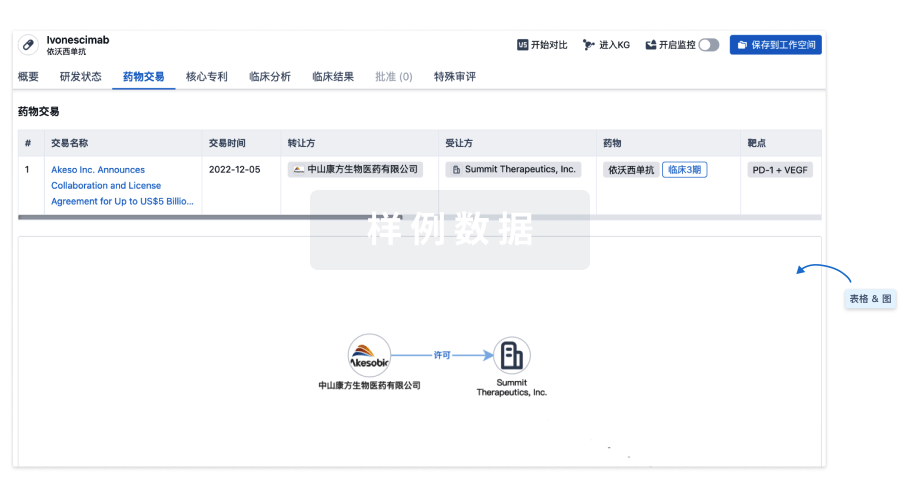
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
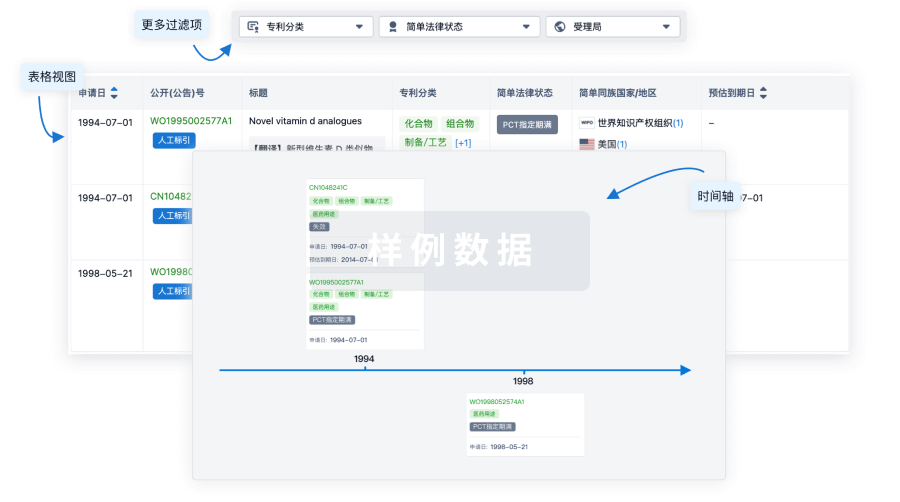
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
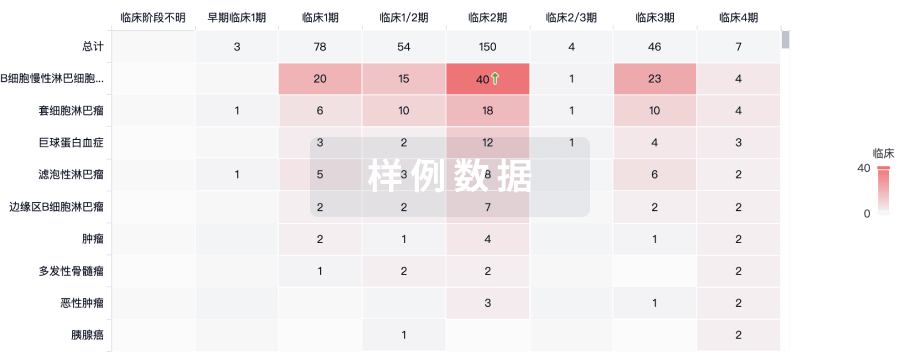
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
批准
利用最新的监管批准信息加速您的研究。
登录
或
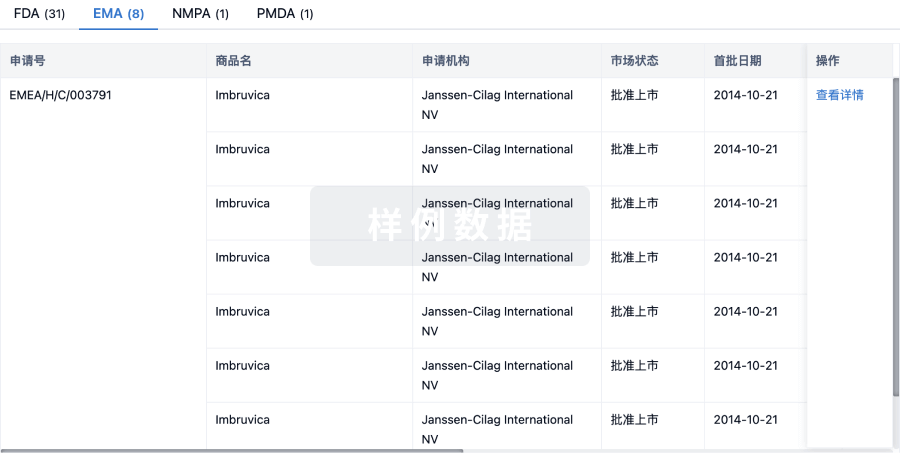
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
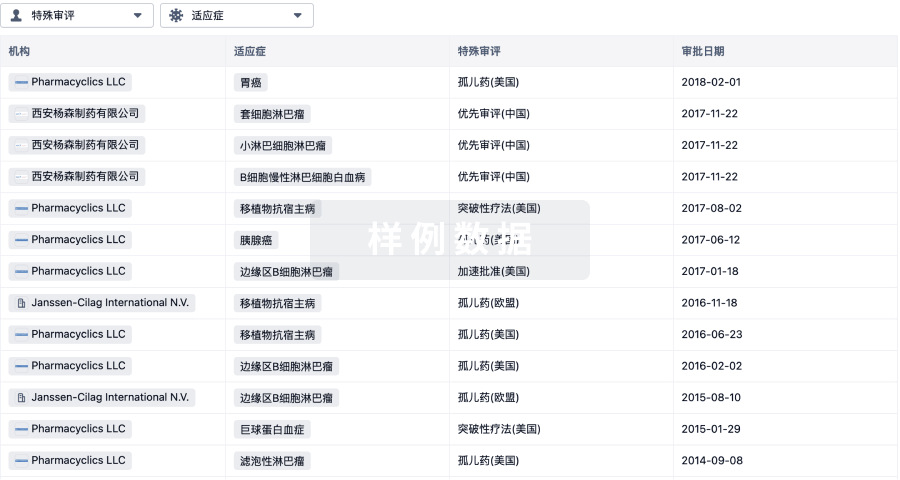
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用








