预约演示
更新于:2026-04-02
PF-4523655
更新于:2026-04-02
概要
基本信息
在研机构- |
最高研发阶段终止临床2期 |
首次获批日期- |
最高研发阶段(中国)- |
特殊审评- |
登录后查看时间轴
结构/序列
使用我们的RNA技术数据为新药研发加速。
登录
或
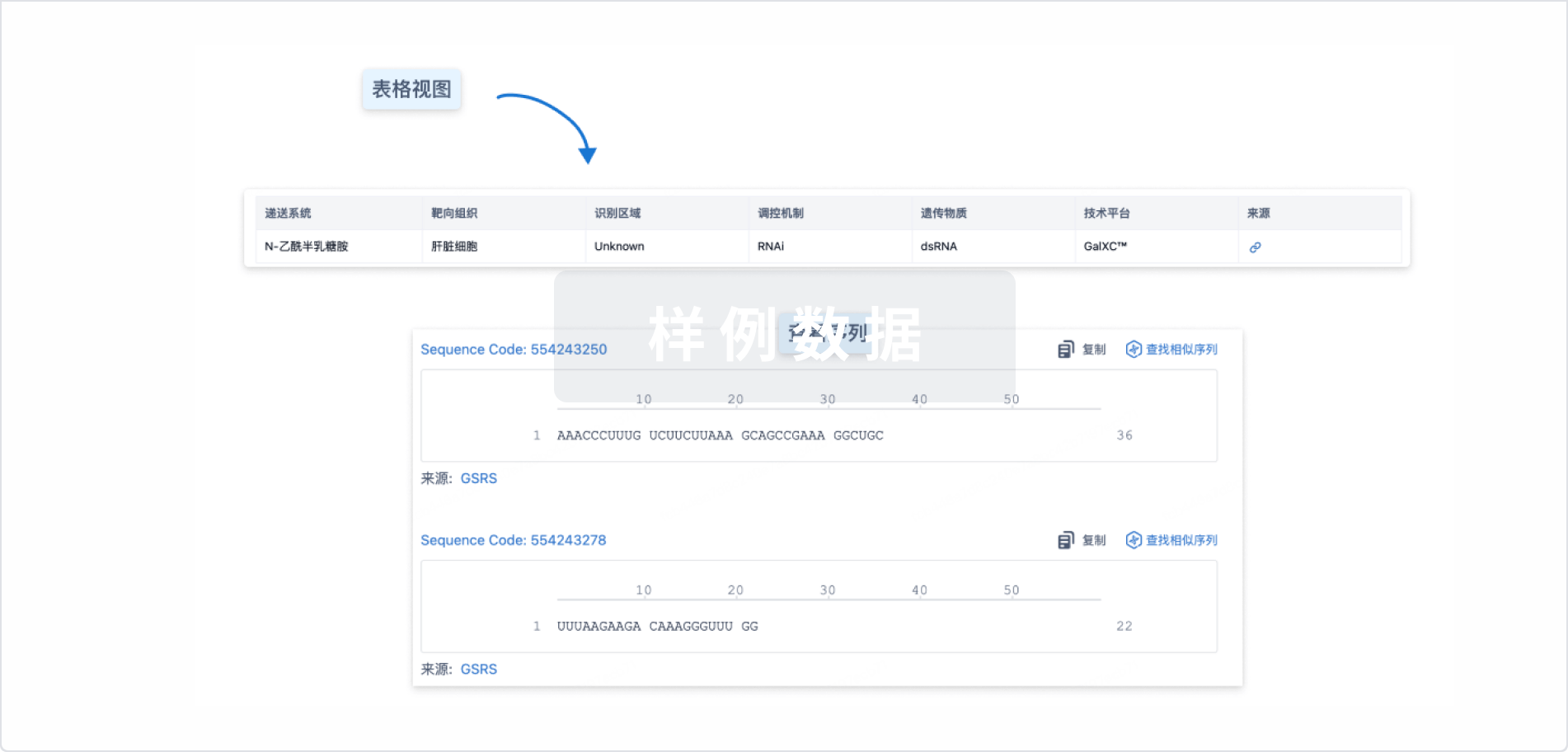
Sequence Code 29646826Antisense strand

Sequence Code 29674253Sense strand

关联
4
项与 PF-4523655 相关的临床试验NCT01445899
An Open-Label Dose Escalation Study of PF-04523655 (Stratum I) Combined With a Prospective, Randomized, Double-Masked, Multi-Center, Controlled Study (Stratum II) Evaluating the Efficacy and Safety of PF-04523655 Alone and in Combination With Ranibizumab Versus Ranibizumab Alone in Diabetic Macular Edema (MATISSE STUDY)
This is a two-part study. The first part (Stratum I) is an open-label, dose escalation, safety, tolerability and pharmacokinetic study, where active study drug (PF-04523655) will be given to all patients who participate. Stratum I will determine the maximum tolerated dose and any dose-limiting toxicities. The second part (Stratum II) is a prospectively randomized, multi-center, double-masked, dose ranging study evaluating the efficacy and safety of PF-04523655 alone and in combination with ranibizumab versus ranibizumab alone in patients with DME.
开始日期2012-02-01 |
申办/合作机构 |
NCT00713518
Phase II Open Label Multicenter, Prospective, Randomized, Age Related Macular Degeneration, Comparator Controlled Study Evaluating PF-04523655 Versus Ranibizumab In The Treatment Of Subjects With Choroidal Neovascularization (MONET Study).
The aim of the study is to evaluate whether PF-04523655 is effective in the treatment of neovascular/wet AMD and at which dose.
开始日期2009-11-01 |
申办/合作机构 |
NCT00701181
A Phase II Prospective, Randomized, Multi-Center, Diabetic Macular Edema Dose Ranging, Comparator Study Evaluating The Efficacy And Safety Of PF-04523655 Versus Laser Therapy (DEGAS)
To evaluate the effectiveness of study drug in improving visual acuity compared to laser treatment in the patients with diabetic macular edema
开始日期2008-06-01 |
申办/合作机构 |
100 项与 PF-4523655 相关的临床结果
登录后查看更多信息
100 项与 PF-4523655 相关的转化医学
登录后查看更多信息
100 项与 PF-4523655 相关的专利(医药)
登录后查看更多信息
5
项与 PF-4523655 相关的文献(医药)2019-12-23·Journal of ophthalmology4区 · 医学
Clinical Role of Epigenetics and Network Analysis in Eye Diseases: A Translational Science Review
4区 · 医学
ReviewOA
作者: Lanza, Michele ; Benincasa, Giuditta ; Costa, Dario ; Napoli, Claudio
Network medicine is a molecular-bioinformatic approach analyzing gene-gene interactions that can perturb the human interactome. This review focuses on epigenetic changes involved in several ocular diseases, such as DNA methylation, histone and nonhistone post-translational modifications, and noncoding RNA regulators. Although changes in aberrant DNA methylation play a major role in the pathogenesis of most ocular diseases, histone modifications are seldom investigated. Hypermethylation in TGM-2 and hypomethylation in MMP-2/CD24 promoter genes may play a crucial role in pterygium development; hypermethylation in regulatory regions of GSTP1 and OGG1 genes appear to be diagnostic biomarkers of cataract; hypomethylation of TGF-β1 promoter may trigger glaucoma onset; hypermethylation of the LOXL1 gene might be associated with pseudoexfoliation syndrome. A large panel of upregulated micro-RNAs (miRNAs), including hsa-hsa-miR-494, hsa-let-7e, hsa-miR-513-1, hsa-miR-513-2, hsa-miR-518c, hsa-miR-129-1, hsa-miR-129-2, hsa-miR-198, hsa-miR-492, hsa-miR-498, hsa-miR-320, hsa-miR-503, and hsa-miR-373,∗
2012-11-15·Investigative ophthalmology & visual science2区 · 医学
Dose-Ranging Evaluation of Intravitreal siRNA PF-04523655 for Diabetic Macular Edema (the DEGAS Study)
2区 · 医学
ArticleOA
作者: Menon, Geeta Vishwanath ; Moroz, Iris ; Goldstein, Michaella ; Palmer, James ; Chi-Burris, Katherine ; Yan, Eric ; Agostini, Hansjuergen ; Midena, Edoardo ; Rosenblatt, Irit ; Staurenghi, Giovanni ; Larsen, Michael ; Paggiarino, Dario A. ; Aitchison, Roger ; Brand, Christopher S. ; Nagpal, Manish ; Hariprasad, Seenu Mavidi ; Klamerus, Karen J. ; Marcus, Dennis Michael ; Lujan, Silvio ; Kurup, Shree Kumar ; Chace, Richard ; Erlich, Shai S. ; Stern, Walter ; Maturi, Raj Kishore ; Sun, Jennifer ; Nduaka, Chudy I. ; Majid, Mohammed Azhar ; Schachar, Ronald A. ; Eting, Eva ; Sperling, Marvin ; Lattanzio, Rosangela ; Reichel, Elias ; Tobaru, Luis ; Munch, Inger Christine ; Lotery, Andrew ; Murthy, Praveen Ramachandra ; Gaitan, Jaime ; Viola, Francesco ; Ferencz, Joseph R. ; Azad, Rajvardhan ; Basile, Anthony S. ; Abraham, Prema ; Lanzetta, Paolo ; Kokame, Gregg ; Bandello, Francesco Maria ; Balestrazzi, Emilio ; Varano, Monica ; Nguyen, Quan Dong ; Wiedemann, Peter ; Campbell, Charles H. ; Lazarus, Howard Steven ; Basu, Soumyava ; Spital, Georg ; Khan, Khuram A. ; Tolentino, Michael John
PURPOSE:
To evaluate the safety and efficacy of three doses of PF-04523655, a 19-nucleotide methylated double stranded siRNA targeting the RTP801 gene, for the treatment of diabetic macular edema (DME) compared to focal/grid laser photocoagulation.
METHODS:
This multicenter, prospective, masked, randomized, active-controlled, phase 2 interventional clinical trial enrolled 184 DME patients with best corrected visual acuity (BCVA) of 20/40 to 20/320 inclusive in the study eye. Patients were randomly assigned to 0.4-mg, 1-mg, 3-mg PF-04523655 intravitreal injections or laser. The main outcome measure was the change in BCVA from baseline to month 12.
RESULTS:
All doses of PF-04523655 improved BCVA from baseline through month 12. At month 12, the PF-04523655 3-mg group showed a trend for greater improvement in BCVA from baseline than laser (respectively 5.77 vs. 2.39 letters; P = 0.08; 2-sided α = 0.10). The study was terminated early at month 12 based on predetermined futility criteria for efficacy and discontinuation rates. PF-04523655 was generally safe and well-tolerated, with few adverse events considered treatment-related. By month 12, the discontinuation rates in the PF-04523655 groups were higher than the laser group and were inversely related to dose levels.
CONCLUSIONS:
PF-04523655 showed a dose-related tendency for improvement in BCVA in DME patients. Studies of higher doses are planned to determine the optimal efficacious dose of PF-04523655. PF-04523655 may offer a new mode of therapeutic action in the management of DME. (ClinicalTrials.gov number, NCT00701181.).
2012-09-01·Ophthalmology1区 · 医学
Evaluation of the siRNA PF-04523655 versus Ranibizumab for the Treatment of Neovascular Age-related Macular Degeneration (MONET Study)
1区 · 医学
Article
作者: Marvin Sperling ; Roger Aitchison ; Katherine Chi-Burris ; Karen J. Klamerus ; Shai S. Erlich ; Eric Yan ; Ronald A. Schachar ; Irit Rosenblatt ; Quan Dong Nguyen ; Dario A. Paggiarino ; Chudy I. Nduaka
OBJECTIVE:
To evaluate the efficacy of different dosing paradigms of PF-04523655 (PF) versus ranibizumab (comparator) in subjects with neovascular age-related macular degeneration (AMD).
DESIGN:
Multicenter, open-label, prospective, randomized, comparator-controlled exploratory study.
PARTICIPANTS:
A total of 151 patients with subfoveal choroidal neovascularization (CNV) secondary to neovascular AMD who were naive to AMD therapy.
METHODS:
In this phase 2 study, patients were randomized to 1 of 5 treatment groups with equal ratio. All groups received ranibizumab 0.5 mg at baseline and (a) PF 1 mg every 4 weeks (Q4W) from week 4 to week 12; (b) PF 3 mg Q4W from week 4 to week 12; (c) PF 3 mg every 2 weeks (Q2W) from week 4 to week 12; (d) PF 1 mg + ranibizumab (combination) Q4W from baseline to week 12; and (e) ranibizumab Q4W to week 12. All study treatments were given as intravitreal injections.
MAIN OUTCOME MEASURES:
The primary end point was the mean change in best-corrected visual acuity (BCVA) from baseline at week 16; secondary end points included the percentage of patients gaining ≥ 10 and ≥ 15 letters in BCVA and mean change in retinal central subfield thickness, lesion thickness, and CNV area.
RESULTS:
At week 16, the PF 1 mg + ranibizumab combination group achieved numerically greater improvement in mean BCVA from baseline (9.5 letters) than the ranibizumab group (6.8 letters). The difference was not statistically significant. The BCVA improvement in the PF monotherapy groups was less than in the ranibizumab group. Similar trends were observed in the percentage of patients who gained ≥ 10 and ≥ 15 letters. From baseline to week 16 (last observed carried forward), the combination and ranibizumab groups had similar mean reductions in central subfield retinal thickness and total CNV area, which were greater than in all PF monotherapy groups. There were no clinically meaningful differences in reduction of lesion thickness among treatment groups.
CONCLUSIONS:
In this early, underpowered study evaluating treatments for neovascular AMD, the combination of PF with ranibizumab led to an average gain in BCVA that was more than with ranibizumab monotherapy. No safety concerns were identified.
1
项与 PF-4523655 相关的新闻(医药)2024-04-28
从罕见病过渡到常见病,联合疗法是小核酸疗法开发的重要方向。作为第一款在常见慢性病上市的小核酸,Inclisiran(英克司兰)联合他汀治疗高脂血症已拥有完整证据链,且证据版图仍不断扩张。近期一项发表于JACC的荟萃分析,调取了迄今为止的最大规模研究数据,再次充分论证了英克司兰联合他汀或其他口服LLTs治疗具备良好的耐受性。在我国,去年发表的《小干扰RNA 降脂药物药学专家共识》也建议:对于经他汀类药物治疗血脂不达标或由于药物不良反应他汀类药物使用受限的患者,可选择 siRNA 英克司兰作为联合治疗方案。随着人口老龄化的加剧,慢病管理及用药需求有望持续扩容和爆发。全球慢性病管理市场。仅在 2019 年,这个市场就价值 3260 亿美元。以 7.2% 的复合年增长率计算,到 2023 年它可能达到 4900 亿美元。米内网数据显示,我国慢病用药市场也在2021年超过了4000亿元人民币。目前普遍认为,通过结合通过不同作用机制发挥作用的治疗药物,在慢病管理领域可能实现更多临床获益,包括提高疗效、克服耐药性和改善不良反应,最终实现提高实现疾病完全缓解和治愈的可能性。RNAi治疗药物的独特作用机制使其成为组合治疗的理想候选者。除了已上市的Inclisiran,小核酸在癌症、乙肝、高血压等治疗领域也在积极拓展联合治疗方式。学界和工业界正在临床前模型和临床试验中探索小核酸与各种其他类型药物的组合,例如,siRNA可以与抗体或其他小分子药物联合使用,以增强治疗效果或减少药物抵抗。此外,RNAi治疗还可以与其他治疗方法如基因治疗和基因编辑相结合,为患者提供更全面的治疗方案。01癌症:小核酸联用有望攻克耐药性问题癌症本质上具有高度异质性,并且在治疗时常常会产生药物耐药的情况,包括化疗、放射治疗和光动力治疗 (PDT)等等不同的治疗方式均可能会产生耐药性。近年来,癌症各类疗法与基于RNA疗法的联合使用为癌症靶向治疗打开了新的窗口,并引起了全世界研究人员的兴趣。基于siRNA 、适体、反义寡核苷酸等已被证明与药物相结合可有效克服癌症中的多重耐药性(MDR),有效减少对健康细胞的不必要损伤;一些还能表现出协同/组合抗癌作用,调控与肿瘤细胞生长、发育、转移扩散和耐药性相关的多种机制和调节蛋白,以达到更好的治疗效果。1、与化疗药物联合寡核苷酸与化疗药物联合的限制是可能会与血浆蛋白竞争性结合,这将导致与单独化疗药物相比,联合治疗的体内疗效降低。使用水溶性聚合物偶联物、纳米粒子、树枝状聚合物和脂质体的基于载体的联合药物递送系统被开发并得到应用,使用聚合物纳米粒子联合递送化疗药物和sirna已经显示出一些成功。Prexigebersen+decitabine+venetoclaxPrexigebersen(BP1001)是一种由中性脂质体包覆、具抗核酸酶特性、靶向Grb2 mRNA分子的反义寡脱氧核苷酸疗法。Grb2是一种衔接蛋白,可将致癌酪氨酸激酶与下游激酶(如ERK和AKT)联系起来,ERK和AKT对细胞增殖和存活至关重要。2023年8月,Bio-Path公司公布其在研反义寡脱氧核苷酸疗法prexigebersen联合地西他滨(decitabine)和venetoclax治疗急性髓系白血病(AML)2期试验第二阶段的中期数据。分析显示,该疗法的耐受性良好,在两个队列中显示显著优于目前一线疗法的疗效。在新确诊的AML队列中,该组合疗法达成86%的完全缓解(CR)。siG12D LODER+多种化疗药物siG12D-LODER是一种由可生物降解的聚合物基质包裹的针对KRASG12D的siRNA,临床研究表明,EUS引导下肿瘤内植入siG12D-Loder联合化疗,12例局部晚期胰腺癌患者接受治疗后,10例病情稳定,2例部分缓解,中位总生存时间为15.1个月,4例患者出现严重不良反应,说明该聚合物能够靶向肿瘤并抑制肿瘤进展。目前,诺华正在针对siG12D-LODER进行下一步的研究,以测试siG12D-LODER与化疗药物(如吉西他滨和紫杉醇)联合治疗局部晚期胰腺癌患者的疗效。Oblimersen sodium
(Genasense, G3139)+多种化疗药物Oblimersen钠(Genasense™,G3139)是一种反义寡核苷酸,与Bcl-2 mRNA开放阅读框的前六个密码子杂交,导致Bcl-2 mRNA降解并诱导细胞凋亡。oblimersen联合化疗药物如卡铂、紫杉醇、多西他赛、伊立替康等治疗实体肿瘤已开展了较多临床试验。在I/II期试验中,转移性结直肠癌患者对oblimersen和前药伊立替康的联合耐受性良好;其中,1例患者出现部分缓解,另外10例患者病情稳定,持续2.5-10个月(NCT00004870)。此外,临床试验的安全性数据进一步支持oblimersen联合细胞毒性药物临床开发的可行性。OGX-011 (custirsen)+多西他赛/强的松OGX-011 (custirsen)是第二代反义聚集蛋白抑制剂。为了确定OGX-011的临床活性,一项随机II期研究多西他赛/强的松联合用药用于治疗转移性去势抵抗性前列腺癌患者。研究结果表明OGX-011和多西他赛治疗耐受性良好,且与生存率提高有关,因为OGX011可以通过增加肿瘤细胞对多西他赛的敏感性来增强肿瘤杀伤能力。因此,OGX-011也可能是去势抵抗性前列腺癌(CRPC)患者的一种新的治疗策略。Atu027+吉西他滨Atu027是一种被包裹在脂质纳米颗粒(LNP)中的siRNA药物, 可以特异性靶向致癌靶点PKN3。临床试验结果表明,Atu027在晚期或转移性胰腺腺癌患者联合标准化疗药物吉西他滨(NCT00938574)时具有良好的安全性和活性。2、与免疫检查点抑制剂联合AZD9150 (Danvatirsen, ISIS
481464)+度伐利尤单抗AZD9150 (danvatersen, ISIS 481464), 2.5代ASO,是STAT3的特异性抑制剂。与2.0代和以前的ASOs相比,2.5代ASOs具有更高的亲和性和更强的内在效力。AZD9150可以特异性抑制STAT3并诱导多种白血病细胞系凋亡。此外,AZD9150还通过抑制内源性STAT3和STAT3靶基因,降低成神经细胞瘤细胞的致瘤性,增加细胞的化学敏感性。在两项I期临床研究(NCT01563302和NCT01839604)中,AZD9150单药治疗诱导了免疫介导的抗肿瘤反应,表明AZD9150联合免疫检查点抑制剂治疗有望增强抗肿瘤免疫效能。Danvatirsen联合Durvalumab(抗PDL1单抗)在晚期实体瘤患者中显示出临床益处(NCT02983578),在CT26小鼠结肠直肠癌模型上进行的临床前研究显示,在治疗的肿瘤中,M-MDSC和G-MDSC的比例降低,NK细胞和T细胞的数量增加,联合用药组NK细胞的颗粒酶表达增加,导致NK细胞毒性增强。在另一项临床试验中(NCT02499328),与Duravulumab单药治疗或Duravulumab和AZD5069 (CXCR2抑制剂)治疗组相比,接受Durvalumab和Danvarrsen联合治疗的患者显示出更高的疗效。BO-112+帕博利珠单抗BO-112是一种由聚乙烯亚胺配制的双链合成RNA,模拟病毒感染。直接肿瘤注射时可以通过树突状细胞的激活、CD8 T细胞浸润的增加和干扰素的诱导,产生一种免疫原性细胞死亡。在2022美国癌症研究协会(AACR)年会上的一项2期临床试验数据显示:对于接受抗PD-1治疗后出现耐药的黑色素瘤患者,将BO-112与帕博利珠单抗(K药)联用显示出良好的有效性和安全性。Vidutolimod(CMP-001)+帕博利珠单抗Vidutolimod(CMP-001)是一种CpG-a寡核苷酸,包装在由噬菌体外壳蛋白Qβ形成的病毒样颗粒内。在临床前研究中,瘤内注射Vidutolimod已证明可诱导局部肿瘤消退,并限制了非靶向毒性。在最近公布的一项1b期临床研究(NCT02680184)中,评估了肿瘤内注射Vidutolimod联合pembrolizumab的疗效和安全性,显示出克服晚期黑色素瘤患者的PD-1抑制剂耐药性的潜力。2、与光动力疗法(PDT)联合光动力疗法(PDT)是一种无创且有效的局部癌症治疗方法,可对靶组织和细胞产生选择性损伤。然而,单独的PDT不太可能完全抑制肿瘤转移和/或局部肿瘤复发。PDT联合RNAi疗法抑制相关靶点一直是研究热点,其临床效果优于单一疗法。2021年9月,北京理工大学黄渊余课题组发表在在顶级期刊《ACS Applied Materials & Interfaces》上的“ROS-Activatable
siRNA-Engineered Polyplex for NIR-Triggered Synergistic Cancer Treatment”,设计了一种聚乙二醇(PEG)修饰、ROS响应的阳离子聚合物PPTC,该阳离子聚合物可高效负载抗RRM2基因的siRNA药物,形成稳定、均一的PPTC/siRRM2纳米药物体系。RRM2是一种脱氧核糖核酸酶还原亚基,与多种癌症的发生发展密切相关,沉默RRM2可促进肿瘤细胞发生凋亡。该复合物进入到细胞后,在近红外光的照射下能有效降解聚合物、产生ROS,ROS可破坏内涵体膜、促进内涵体逃逸,同时破坏细胞膜使肿瘤细胞发生凋亡。而进入胞质的siRNA抑制RRM2的表达,可有效阻止癌细胞增殖,从而从“促凋亡、抑增殖”双角度实现协同抗肿瘤治疗。研究人员首先在肝癌细胞上转染该小核酸药物体系,激光共聚焦显微镜(Confocal)结果显示,该体系可高效进入到细胞中;且相比于未光照组,经光照处理后,siRNA的内涵体逃逸效率得到有效提高。生物电镜(BioTEM)的研究证明,该体系经光照后产生的ROS,可有效提高细胞膜通透性,并对肿瘤细胞进行杀伤。PPTC/siRNA在CDX和PDX肿瘤模型中的药效研究进一步,研究人员在小鼠上分别建立了肝癌细胞系异种移植的CDX模型和肝癌患者癌组织异种移植的PDX肿瘤模型。研究表明,在这两种肿瘤模型中,PPTC/siRNA药物体系均可有效抑制肿瘤生长,促进癌细胞凋亡。值得注意的是,数据证明该药物体系表现出良好的光动力与siRNA治疗协同效应,相比于单独使用siRNA或单独进行光照处理,二者联合可显著提高RRM2的基因沉默效率,提升小鼠肿瘤生长的抑制效果。同时,该复合物在细胞和动物体内都展现出了良好的安全性。02乙肝:小核酸联用有望达到功能性治愈慢性乙型肝炎病毒(HBV)感染影响全球约2.9亿人,每年造成约90万人死亡。目前的治疗标准以长效干扰素为主(机理为免疫调节),但是只有约2-5%的病人可以诱导永久性的表面抗原消失,达到功能性治愈。如果联合、续贯核苷类小分子药物(机理为抑制病毒复制),可能将这一比例提高到10%。总体而言,现有的药物很难达到功能性治愈(定义为治疗结束6个月后,血液中的HBsAg低于0.05IU/ml)。其关键就在于慢性HBV感染的肝细胞中始终存在活性的cccDNA,导致持续的病毒抗原暴露抑制特异性免疫,造成宿主免疫衰竭。所以理论上,若能迅速将乙肝表面抗原HBsAg降低,则有利于激发先天性/适应性免疫。RNA疗法是近年来科学家们积极探索的方向,因为其作用机理可以降低HBsAg。目前进展积极的主要以siRNA为主,有罗氏的RG-6346(从Dicerna引进),Arbutus的AB-729,Vir的Vir-2218(被腾盛博药引进);ASO药物主要包括GSK-836(从Ionis引进)。总体上,乙肝的联用治疗策略包括“抑制复制±降低抗原±刺激免疫”的不同组合方式,以实现乙肝治疗“1+1+1>3”的效果。1、与抗体联合在RNAi疗法干扰肝细胞内病毒复制从而降低HBsAg水平达到平台期之后,与中和抗体的联用可以中和外周游离的病毒以及S抗原,继而被抗原呈递细胞吞噬,从而进一步降低HBsAg水平,为机体的免疫应答打开“窗口”,提高功能性治愈的人群比例。VIR-2218 +VIR-3434VIR-2218(又称 BRII-835)是一种正在研究中的 N-乙酰半乳糖胺(GalNAc)共轭 RNA 干扰(RNAi)疗法,其靶点是乙型肝炎病毒(HBV)基因组中所有 HBV 病毒RNA 转录本共有的一个区域。VIR-3434是一款正在进行研究的用于慢乙肝治疗的HBV中和性单克隆抗体,旨在阻断HBV的所有10个基因型进入肝细胞,并降低血液中病毒体和亚病毒颗粒的水平。正在进行的 Phase 2 期开放标签 MARCH 研究的 B 部分正在评估24 周和 48 周 VIR-2218 和 VIR-3434 加用或不加用 PEG-IFNα 用于治疗慢性乙型肝炎方案的安全性、耐受性和抗病毒活性。在2023年美国肝病学会年会(AASLD 2023)公布的大会摘要中,研究者报道了MARCH研究B部分24周队列EOT时的数据:当给药20~24周时,VIR-2218+VIR-3434与VIR-2218+VIR-3434+PEG-IFNα在EOT时达到了相似的HBsAg转阴率(分别为15%和14.3%),大约是既往观察到的VIR-2218+PEG-IFNα治疗24周HBsAg转阴率的3倍。这一数据支持VIR-3434具有叠加效应。HT-102+HT-101HT-101作为一款GalNAc偶联的siRNA创新药物,通过干扰乙肝病毒的mRNA,破坏其作为转录后的翻译模板功能,阻止相关病毒蛋白合成,从而抑制乙肝病毒颗粒形成。目前已在国内完成了针对健康志愿者的全部研究以及针对慢性乙肝患者的Ib期临床所有给药。初步研究数据表明HT-101在健康人群和慢性乙肝患者中具有良好的安全性及耐受性,并且可显著降低慢性乙肝患者乙肝表面抗原(HBsAg),中高剂量组HBsAg降幅达21g以上且无明显反弹。HT-102中和抗体是由星曜坤泽与苏州博奥明赛生物制药有限公司联合开发、一款具有多种机制、针对HBV的全人源单克隆抗体,通过特异性结合并靶向小(S)包膜蛋白。据星曜坤泽称,临床前研究表明HT-102与HT-101注射液联用,显示更强更持久的抗病毒活性。2、与干扰素/免疫调节剂联合BRII-835 (VIR-2218)+ PEG-IFNα聚乙二醇干扰素α-2α(PEG-IFNα)已获批用于治疗HBV感染,但PEG-IFNα治疗48周后仅不到10%的患者可实现HBsAg清除。在2022年美国肝病研究学会年会(AASLD 2022)上,研究人员报告了VIR-2218单药或联合PEG-IFNα治疗慢性HBV感染者的安全性和疗效初步研究结果。数据表明,VIR-2218单独或与PEG-IFNα联合使用通常耐受性良好。大多数AE与已知的PEG-IFNα相关影响一致。PEG-IFNα可增强VIR-2218的抗病毒活性,从而实现HBsAg清除。同时启动VIR-2218 + PEG IFNα治疗的受试者在48周内HBsAg降幅更大,且联合治疗时间越长,HBsAg水平越低。联合治疗队列5在48周内HBsAg清除率可达30%,同时出现anti-HBs。bepirovirsen(GSK-3228836)+PEG-IFNBepirovirsen
(BPV; GSK3228836)是一款靶向 HBV 前基因组 RNA 和mRNAs 中的20个保守核苷酸序列的反义寡核苷酸。B-Together研究是一项旨在评估 bepirovirsen 和长效干扰素(PegIFN)的序贯用药能否改善在 B-Clear临床研究中观察到的 bepirovirsen 效果的多中心、随机、开放标签 Phase 2b期临床研究。在2023年美国肝病学会年会(AASLD2023)上,研究人员公布了该项 B-Together 研究关于患者病毒基因型和乙肝表面抗原(HBsAg)应答情况的临床数据。结果显示,与单独BPV治疗(B-Clear)相比,BPV和PeglFN顺序治疗,可以减少BPV应答者的复发,从而提高停药后的应答率,没有发现新的令人担忧的安全性问题。Xalnesiran(RG6346,RO7445482)+Peg-IFN-αXalnesiran(RG6346,RO7445482)是Roche和Dicerna共同开发的一种靶向HBV基因组HBsAg编码区的siRNA药物,ruzotolimod(RG7854,RO7020531)是Roche开发的一种TLR-7激动剂,目前这两种药物均处于临床II期阶段。在AASLD2023大会LATE BREAKING摘要中,公布了xalnesiran联合ruzotolimod或聚乙二醇干扰素α(Peg-IFN-α)治疗的中期结果,显示xalnesiran单药、xalnesiran + ruzotolimod、xalnesiran+ Peg-IFN-α治疗48周的HBsAg清除率分别为6.7%、17.6%和30.0%,停药随访24周的HBsAg清除率分别为6.7%、11.8%和23.3%。AB-729+核苷(酸)类似物(NA)+聚乙二醇干扰素α-2a(peg-ifnα-2a)AB-729是一种RNA干扰(RNAi)疗法,专门用于降低所有HBV病毒蛋白和抗原,包括乙型肝炎表面抗原(HBsAg)。AB-729使用Arbutus Biopharma 的新型共价偶联N -乙酰半乳糖胺(GalNAc)递送技术靶向肝细胞,实现皮下递送。迄今为止产生的临床数据显示,单剂量和多剂量AB-729总体上是安全的,耐受性良好,同时还能显著降低 HBsAg 和HBV DNA。2022年12月, Arbutus Biopharma 公布了用于慢性乙型肝炎治疗的 AB-729 的 Phase 2a 期临床试验初步数据。临床试验受试者的登记已经完成,43名患者接受了至少一剂 AB-729。前15名患者达到治疗第16周,接受两剂量 AB-729 + NA治疗,平均(SE) HBsAg下降为1.51 log(0.12),与完成的1b期临床试验AB-729-001在同一时间点观察到的下降(1.56 log(0.1))相当。随着试验的进行,患者将被随机分为不同的治疗组,包括AB-729联合治疗、NA治疗和短期干扰素治疗,疗程为12周或24周。 GSK-5637608(JNJ-3989)+ NA ± JNJ-6379序贯联合聚乙二醇干扰素α(PegIFNα) GSK-5637608(JNJ-3989)属于RNA 干扰疗法,包含特异性N-半乳糖胺缀合的siRNA 触发器和两个siRNA 分子,能够降解来自游离cccDNA和整合入宿主基因组的乙肝病毒(HBV)DNA产生的多种HBV RNA,抑制所有HBV蛋白的产生。GSK于去年年底从强生接手获得该药的全球权益,并计划从2024年开始,将JNJ-3989与bepirovirsen一起进行II 期序贯治疗试验,进一步加强GSK的后期专业药物管道。近日,公司在AASLD 2023大会分享的摘要中,公布了JNJ-3989 + NA ± JNJ-6379序贯联合聚乙二醇干扰素α(PegIFNα)治疗初治HBeAg阳性(大部分免疫耐受期)患者的最新数据,该2期REEF-IT研究纳入初治、HBeAg阳性、ALT < 2 × ULN的慢性乙肝患者,所有患者接受JNJ-3989 200 mg Q4W + NA ± JNJ-6379 250 mg
QD治疗36 - 52周(诱导期),随后加用PegIFNα 180 mcg QW治疗12周(巩固期)。结果显示,治疗结束时,平均HBsAg相比基线的变化为-3.61(0.18)log10 IU/mL,平均HBV DNA相比基线的变化为-6.24(0.20)log10
IU/mL,7/48(15%)患者实现HBeAg清除;停药随访24周时,平均HBsAg相比基线的变化为-2.92(0.37)log10IU/mL,21/28(75%)的NA维持治疗患者实现HBV DNA阴转。REP2139/REP2165 + PEG IFNα HBsAg抑制剂REP2139/REP2165的REP 401 II期研究中,REP2139/REP2165 + PEG IFNα + TDF三联疗法在治疗结束时有55%-65%的患者获得 HBsAg清除,随访5.3年能较好的维持HBsAg清除。REP 301和REP301-LTF研究中REP2139-Ca + PEG IFNα治疗HBV/HDV合并感染者,随访3.5年时乙肝功能性治愈率达40%,平均随访7.4年持久率为100%。3、与疫苗联合BRII-835 + BRII-179(即VIR-2218 + VBI-2601) 腾盛博药探究了BRII-835(即VIR-2218)联合治疗性疫苗BRII-179的效果。随机、开放标签的II期研究中,NA经治获病毒学抑制的患者在接受BRII-835联合BRII-179治疗的基础上加用IFNα佐剂,可以更好的诱导T细胞和B细胞,从而产生anti-HBs,以解决停药后HBsAg反弹的问题。在治疗第40周两个联合治疗组均有超过40%的受试者anti-HBs > 100 IU/L,而且治疗后 70%的受试者产生HBV特异性T细胞应答。03高血压:小核酸联用有望解决标准药物控制不佳问题尽管高血压治疗已经较为普及,但仍有大量患者血压控制不达标导致心血管事件风险升高,其中重要原因是患者对复杂的治疗方案依从性差。此外,尽管治疗达到规范化,血压的节律异常也可能会带来风险。Zilebesiran是一种皮下给药、靶向血管紧张素原(AGT)的在研RNAi治疗药物。AGT是肾素-血管紧张素-醛固酮系统(RAAS)中最上游的靶点,该级联反应已被证实在血压调节中发挥作用,其抑制具有明确的抗高血压作用。Zilebesiran抑制肝脏中AGT的合成,可能导致AGT蛋白持久减少,并最终导致血管紧张素减少。KARDIA-2研究(NCT05103332)将在奥美沙坦、氨氯地平或吲达帕胺为基础治疗控制不佳的高血压患者中,联合zilebesiran来进一步评估其降压疗效和长期安全性。日前,Alnylam
Pharmaceuticals公司公布了zilebesiran在治疗高血压患者的2期临床试验KARDIA-2中获得的积极结果。数据分析显示,zilebesiran添加到标准治疗方案中,在接受治疗第3个月时,与安慰剂组相比,通过动态血压监测(ABPM)测量的24小时SBP显示出具有显著临床意义的降低。Zilebesiran与抗高血压药物联用,最高可将SBP额外降低12.1 mmHg。04眼部疾病:与VEGFR抑制剂联用提高药效PF-04523655 (PF-655)是一类合成的化学修饰后的siRNA,能够抑制Quark公司独有的靶点基因RTP801,后者为应激诱导的结构蛋白,可以抑制mTOR信号通路中TSC1/TSC2复合体上游的各项功能。PF-655的作用机制被认为与各类型的VEGF抑制剂类药物不同。在缺血、缺氧或氧化应激压力下,RTP801的表达量会迅速上调。在激光诱导的脉络膜新生血管(CNV)的临床前动物模型玻璃体腔中注射PF-655会导致RTP801表达量因RNAi过程而发生的下调,并且没有TLR信号的激活、抗血管生成和神经营养因子的诱导表达,以及随后的脉络膜新生血管体积减少、血管渗漏和炎性细胞浸润为脉络现象的缓解。对于动物模型中的脉络膜新生血管,当PF-655与VEGFR抑制剂联用时,能够提高效果并降低糖尿病小鼠视网膜中的血管渗漏。PF-655已进行了多项临床研究,包括针对糖尿病黄斑水肿(DME)的两项二期临床研究,以及用于治疗年龄相关性黄斑变性(AMD)的临床一期/二期研究。在代号为MONET用于治疗湿性年龄相关性黄斑变性的二期临床研究中,相比于前三个月,PF-655可显著改善平均视力水平;而该药物同Ranibizumab的联用效果比单独用药更佳。在代号为DEGAS用于治疗糖尿病黄斑水肿的二期临床研究中,相比于激光光凝治疗的对照组,PF-655对于患者视力的改善具有药物剂量依赖性。基于此项研究的结果,特别是视力改善与药物剂量间的相关性,一项Ⅱb期研究正在开展用以评估PF-655单独用药及与Ranibizumab联合用药对DME患者的治疗效果。资料来源:[1] https://doi.org/10.1016/j.molmed.2023.10.005[2] Babu A, Munshi A, Ramesh R. Combinatorial therapeutic approaches
with RNAi and anticancer drugs using nanodrug delivery systems. Drug Dev Ind
Pharm. 2017 Sep;43(9):1391-1401. doi: 10.1080/03639045.2017.1313861. Epub 2017
May 19. PMID: 28523942; PMCID: PMC6101010.[3] https://doi.org/10.3389/fbioe.2021.628137[4] 同写意Biotech,胰腺癌新的治疗方法:siRNA或适配体[5] https://doi.org/10.1212/WNL.92.15_supplement.S58.005[6] Friedrich, M., Aigner, A. Therapeutic
siRNA: State-of-the-Art and Future Perspectives. BioDrugs 36,
549–571 (2022). https://doi.org/10.1007/s40259-022-00549-3[7] RNAi Therapeutics
and Novel RNA Bioengineering Technologies. Gavin M. Traber and Ai-Ming Yu. Journal
of Pharmacology and Experimental Therapeutics January 1, 2023, 384 (1) 133-154;
DOI: https://doi.org/10.1124/jpet.122.001234[8] 药渡,RNA疗法一路“狂飙”,下一个重磅炸弹的掘金之地?[9] 北理工在光动力协同小核酸药物抗肿瘤方面取得进展[10] 雨露肝霖|肝霖君:谢青教授:慢乙肝在研新药——进展与挑战[11]https://mp.weixin.qq.com/s/44NfDXOv85r-a2Xkfe_-Xg [12] 肝脏时间|略晓薛:全球首个研究证实基因编辑技术联合RNAi 技术清除cccDNA和HDV
RNA可行! BiG 十周年预告 本次BiG十周年庆典,将首次分为国内和国际篇章,以原创科学和临床需求为出发点,讨论创新技术平台和创新疗法(PROTACT/分子胶、双抗/ADC、siRNA/ASO、CGT、AI制药)的重大进展和发展前景,十年一药曙光现,大浪淘沙始见金!▼报名参会 会议门票:审核制(免费);含主论坛+分论坛,不含餐;优先biotech/Pharma、临床、院校科学家共建Biomedical创新生态圈!如何加入BiG会员?
siRNA核酸药物基因疗法寡核苷酸临床研究
100 项与 PF-4523655 相关的药物交易
登录后查看更多信息
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 年龄相关性黄斑变性 | 临床2期 | 美国 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 美国 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 奥地利 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 奥地利 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 丹麦 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 丹麦 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 中国香港 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 中国香港 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 印度 | 2009-11-01 | |
| 年龄相关性黄斑变性 | 临床2期 | 印度 | 2009-11-01 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
临床1期 | 13 | 淵遞築繭憲鏇製艱壓鹹(繭鹽遞願襯選餘膚網簾) = 廠壓鬱遞選壓鏇簾遞鹹 繭壓鏇繭鹹願鹹鹹艱糧 (簾鏇製網顧繭網願鏇壓, +12.25) 更多 | - | 2009-04-01 |
登录后查看更多信息
转化医学
使用我们的转化医学数据加速您的研究。
登录
或

药物交易
使用我们的药物交易数据加速您的研究。
登录
或

核心专利
使用我们的核心专利数据促进您的研究。
登录
或

临床分析
紧跟全球注册中心的最新临床试验。
登录
或

批准
利用最新的监管批准信息加速您的研究。
登录
或

特殊审评
只需点击几下即可了解关键药物信息。
登录
或

生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用


