预约演示
更新于:2026-03-07

Indian Association for the Cultivation of Science
更新于:2026-03-07
概览
标签
肿瘤
皮肤和肌肉骨骼疾病
感染
小分子化药
合成多肽
siRNA
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 小分子化药 | 4 |
| 合成多肽 | 3 |
| siRNA | 1 |
关联
8
项与 Indian Association for the Cultivation of Science 相关的药物靶点 |
作用机制 KRAS调节剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 KRAS调节剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 G4抑制剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
100 项与 Indian Association for the Cultivation of Science 相关的临床结果
登录后查看更多信息
0 项与 Indian Association for the Cultivation of Science 相关的专利(医药)
登录后查看更多信息
5,532
项与 Indian Association for the Cultivation of Science 相关的文献(医药)2026-03-01·ARCHIVES OF MICROBIOLOGY
Bioremediation of oil contaminants from the marine ecosystem by Alcanivorax borkumensis: an overview
Review
作者: Sachan, Garima ; Sharma, Swati ; Chauhan, Shikha ; Sarkar, Saptak ; Gupta, Shreya
2026-03-01·JOURNAL OF MOLECULAR BIOLOGY
Post-Translational Modifications Orchestrate Repair of Trapped Topoisomerase-Induced DNA Breaks via TDP1 and TDP2
Review
作者: Sengupta, Abhik ; Das, Benu Brata ; Bhattacharyya, Arpan ; Basu, Saini
DNA topoisomerases are critical for maintaining DNA topology and facilitating replication, transcription, and chromatin organization in both nuclear and mitochondrial genomes. When covalently trapped on DNA as topoisomerase cleavage complexes (Topcc's), notably Top1ccs and Top2ccs, these enzymes generate cytotoxic DNA lesions that disrupt genomic integrity and threaten cell viability. Tyrosyl-DNA phosphodiesterase (TDP1 and TDP2) has emerged as key player in the resolution of these lesions, with broader roles in the repair of diverse DNA end structures. Post-translational modifications (PTMs) dynamically regulate the DNA damage response by modulating the activity, localization, and interactions of repair factors. This review provides a comprehensive overview of the mechanisms by which PTMs modulate the activity of Top1 and Top2, and the repair of their covalently trapped complexes. We further delineate how PTMs fine-tune the functional networks of TDP1 and TDP2, enhancing their efficiency in resolving Topccs and preserving genome stability. Together, these insights highlight the multilayered regulatory mechanisms that safeguard genomic integrity and offer potential avenues for therapeutic intervention.
2026-02-17·LANGMUIR
Ion-Induced Morphological Plasticity in a Self-Assembling Peptide Hydrogel
Article
作者: Roy, Susmita ; Mondal, Tanushree ; Banerjee, Arindam ; Hamley, Ian W. ; Sinha, Anushree ; Mondal, Biplab
A hydrogel formed by a short peptide is presented that exhibits remarkable stimuli-responsiveness and plasticity, undergoing a morphological transformation from nanofibers to nanospheres in the presence of monovalent (Li+, Na+, K+) or trivalent (Al3+, Fe3+) metal ions under physiological conditions. The nanofibrillar structure of the hydrogel was examined using transmission electron microscopy (TEM), atomic force microscopy (AFM), small-angle X-ray scattering (SAXS), and X-ray diffraction (XRD) studies and atomistic molecular dynamics simulations, in complement, to explain the nanostructural transitions at the microscopic level. Interestingly, exposure to divalent metal ions (Mg2+, Ca2+, Co2+, Ni2+) induces a unique shrinking (syneresis) behavior, accompanied by a morphological shift to nanoribbons. Both simulations and SAXS analysis confirm that these ions cause a contraction in the packing of gelator peptides, significantly reducing the interpeptide distance. This ion-specific adaptability confers tunable physicochemical properties and morphological plasticity. Hydrogels incorporating mono- or trivalent ions exhibit enhanced thermal stability and mechanical strength relative to ion-free counterparts, underscoring the reinforcing role of metal coordination. Strikingly, shrunken gels formed in the presence of divalent ions display even greater stiffness than freshly prepared gels in the absence of any metal ions, suggesting that syneresis acts as a postassembly strengthening mechanism. These findings highlight a versatile, stimuli-responsive soft material in which ion-peptide interactions orchestrate nanoscale morphology, mesoscale network architecture, and macroscopic mechanical performance-opening avenues for adaptive hydrogel systems in targeted biomedical, sensing, and controlled-release applications.
1
项与 Indian Association for the Cultivation of Science 相关的新闻(医药)2024-05-19
导读“我们只关注癌症,并见过各种疑难杂症。我们的医生一天内治疗的罕见癌症比大多数医生一生看到的还要多”。 MD安德森癌症中心在官网介绍自己时有这样一段话。 如果是别的研究所这么说多少有点说大话的嫌疑,但如果是MD安德森,那似乎就很合理了。 在今年新鲜出炉的“美国最佳医院”排名中,MD安德森再一次在肿瘤医院排名中位列第一,而在近十年里,它几乎年年第一。 MD安德森logo也非常有趣,那就是它将“cancer”一词用斜线划掉,可见MD安德森对战癌症的勃勃雄心。 2016年奥巴马政府发起了癌症登月计划,而MD安德森早在2012年就启动了这项计划。为何MD安德森总能在癌症研究方面快人一步? 01癌症疫苗IMGS-001开启临床试验 2023年10月底,德克萨斯大学MD安德森正式开展了IMGS-001对局部晚期或转移性实体肿瘤患者的1a/1b期试验。 IMGS-001是由美国生物技术公司ImmunoGenesis研发的抗癌新药,是一种双特异性PD-L1/PD-L2抗体药物,旨在针对现有免疫疗法具有抵抗力、难以治愈、排斥免疫的肿瘤。 为何IMGS-001的药物试验会选择在MD安德森进行?这或许与MD安德森在癌症领域的绝对权威有关。在《美国新闻与世界报道》的肿瘤排名名单中,MD安德森常年霸榜第一。 2022年,MD安德森开启了1680项临床试验,22种药物获得了FDA批准。02MD安德森雄心勃勃的创建史MD安德森位于休斯顿的德克萨斯医疗中心,而德克萨斯医疗中心是世界上最大的医疗中心。MD安德森的创建与一个人密切相关,那就是梦露·邓纳威·安德森(Monroe Dunaway Anderson)。 安德森出生于1873年,他的父亲是一位银行行长。长大后,安德森和家族成员共同从事棉花生意,并很快积攒了一大笔财富。1936年,安德森创建了以他的名字命名的慈善基金会,并为基金会提供了约30万美元的启动资金。 1939年,安德森去世,基金会分得了一笔1900万美元的巨款。事实上,对于这笔巨额基金该如何处理,MD安德森基金会没有明确的章程。不过这笔基金的受托人有一个强烈的愿景,希望用于医疗保健。 1941年,德克萨斯州政府批准了德克萨斯大学在该州某地建立一家癌症研究和治疗医院的请求,并为这项计划拨款了50万美元,但没有具体说明建设地点。 得克萨斯大学与安德森基金会协商后达成一致:如果这所医院建在休斯顿,并以基金会的捐助者名字来命名,那么安德森基金会可以再提供50万美元的资金。之后,德克萨斯大学在德克萨斯州立癌症医院和癌症研究部的基础上,成立了MD安德森癌症研究医院。 在那之后,MD安德森癌症研究医院经历了多次名字的变迁: 1941年:由德克萨斯州立法机构成立,前身为德克萨斯州立癌症医院和癌症研究部。1942年:更名为德克萨斯大学MD安德森癌症研究医院,以表彰MD安德森基金会的支持。1955年:更名为德克萨斯大学MD安德森医院和休斯顿肿瘤研究所。1972年:德克萨斯大学系统重组导致德克萨斯大学系统癌症中心的成立,包括休斯顿的医院和研究机构以及史密斯维尔的科学园。 1946年,研究医院迎来第一位全职校长——伦道夫·李·克拉克(Randolph Lee Clark)。 乔纳森·罗兹与李·克拉克(右) 彼时,MD安德森癌症研究医院在当时才刚刚起步,正值战乱年代,美国研究所的费用和各类用品都极为短缺,因此研究医院的发展一度受限,举步维艰。克拉克上任后立志用有限的资源,将研究医院发展为“美国大学系统内的第一家癌症医院”。他当时给好友伦道夫·菲尔兹(Randolph Fields)写信时提到建设癌症医院的思路: “有三个主要目标:与癌症疾病相关的教育、治疗和研究活动。这些目标是整个企业的主题,也是每个部门职能的基调”。 可以说,克拉克一生都在兢兢业业为实现这三个目标而奋斗,他在这里工作了32年,直到1978年退休;之后由查尔斯·A·莱梅斯特(Charles A. LeMaistre)和约翰·门德尔森(John Mendelsohn)陆续接任。 直到2011年,第四任校长罗纳德·德皮尼奥(Ronald DePinho)的上任又将MD安德森的癌症研究带向了全新的方向,在他的带领下,MD安德森启动了一项雄心勃勃的癌症“登月”(Moon Shots)计划。03MD安德森何以成为全球癌症研究的先行者?目前,MD安德森如今有23040名员工,包括1870名教职员工。MD安德森现任领导,即第五任总裁彼得·W·T·皮斯特斯(Peter WT Pisters),他以癌症外科医生、研究员、教授和医院管理人员而闻名。 MD安德森的财政来源也很丰富,最主要的一个来源是德克萨斯州癌症预防研究所(CPRIT)。2022财年,MD安德森从CPRIT获得了超过3200万美元的资金支持,用于研究、招聘和培训等事务。 此外,MD安德森获得了国家癌症研究所卓越研究专业计划(SPORE)资助,用于子宫内膜癌、前列腺癌、黑色素瘤、胃肠道癌、脑癌、白血病癌、肺癌、卵巢癌和肝细胞癌等9个方面的研究。 “对抗癌症”是MD安德森一个非常显著的标签,可以说,MD安德森是个不折不扣的癌症研究和治疗的先行者,甚至在Logo设计上,MD安德森特意将“癌症”一词划掉了。 1971,美国颁布《国家癌症法》,指定的美国最早的三个综合癌症中心,MD安德森是其中之一;目前,美国国家癌症研究所指定了53个综合癌症中心,MD安德森亦是其中之一。 另外,在《美国新闻与世界报道》“2023年-2024年最佳医院”评选中,MD安德森在癌症护理方面排名第一。自1990年有这项排名以来,该机构每年都被评为全国癌症护理排名前两位,近十多年来,几乎年年第一。 之所以能够在癌症领域获得如此高的声誉,离不开MD安德森在癌症方面的开拓精神。 2016年,奥巴马政府启动癌症登月计划,主张将1亿9500万美元用于癌症研究,2022年,拜登政府紧随其后,承诺在未来25年将癌症死亡率降低50%。 而早在2012年的时候,有着先见之明的MD安德森第四任校长罗纳德·德皮尼奥(Ronald DePinho)带领MD安德森开启了“登月”(Moon Shots)计划。 2016年12月,德皮尼奥(中)在华盛顿特区签署的《21世纪治愈法案》 这项计划旨在降低癌症的死亡率和发病率。在最开始的时候,MD安德森计划为该项目投入30亿美元,并提出口号:“癌症在此终结”(Cancer ends here)。让癌症可预防、可看见、可治疗,甚至被遗忘。 事实上,这项计划也取得了不错的成果:2014年10月,研究人员提出新算法将卵巢肿瘤切除成功率提高到80%;2016年8月,早期治疗活检预测黑色素瘤免疫治疗的反应;2016年10月,首个由IACS开发的饥饿癌细胞药物进入临床试验;2020年2月,CD19 CAR NK细胞在血液系统癌症中取得高反应。 在医疗试验之外,MD安德森还实现了从法律、提案等角度提升公众健康指数:2019年5月,德克萨斯州将烟草销售年龄提高至21岁,这同样是MD安德森癌症登月计划的一项重要成果。 尽管30亿美元的资金投入不足以让MD安德森未实现“消除癌症”的终极目标,但这项计划无疑推动了癌症领域的研究,更在社会范围内促成了对癌症的关注。后续拜登政府再次提出癌症登月计划,或许多少受到MD安德森该计划的指引。 在临床试验方面,2022年这年,MD安德森进行了1680项临床试验,临床试验共有10074名患者参与,获得了116项专利,研究的经费高达11亿美元,在该中心测试的22种药物获得了FDA批准。 可以说,“癌症登月计划”是MD安德森癌症研究和治疗的一个缩影,正是这种开拓精神以及对人类健康和安全的关切,才会铸就MD安德森在癌症领域的高峰。 参考资料 1、Best Hospitals | U.S. News Hospital Rankings and Ratings.U.S. News & World Report.2、Dosing Begins in Phase 1a/1b Study of IMGS in R/R Solid Tumors..targeted Oncology.3、Randolph Lee Clark.wikipedia.4、MD Anderson Cancer Center.wikipedia.识别微信二维码,添加生物制品圈小编,符合条件者即可加入生物制品微信群!请注明:姓名+研究方向!版权声明本公众号所有转载文章系出于传递更多信息之目的,且明确注明来源和作者,不希望被转载的媒体或个人可与我们联系(cbplib@163.com),我们将立即进行删除处理。所有文章仅代表作者观点,不代表本站立场。
疫苗免疫疗法临床研究
100 项与 Indian Association for the Cultivation of Science 相关的药物交易
登录后查看更多信息
100 项与 Indian Association for the Cultivation of Science 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年06月10日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
2
6
临床前
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
Sunshinamide ( GPX4 x TrxR1 ) | 乳腺癌 更多 | 临床前 |
C17 (Indian Association for the Cultivation of Science) ( Top I ) | 利什曼病 更多 | 临床前 |
TTh1 ( KRAS ) | 肿瘤 更多 | 临床前 |
TI12 ( G4 ) | 白血病 更多 | 临床前 |
IGT递送Nanog siRNA ( NANOG ) | 前列腺癌 更多 | 临床前 |
登录后查看更多信息
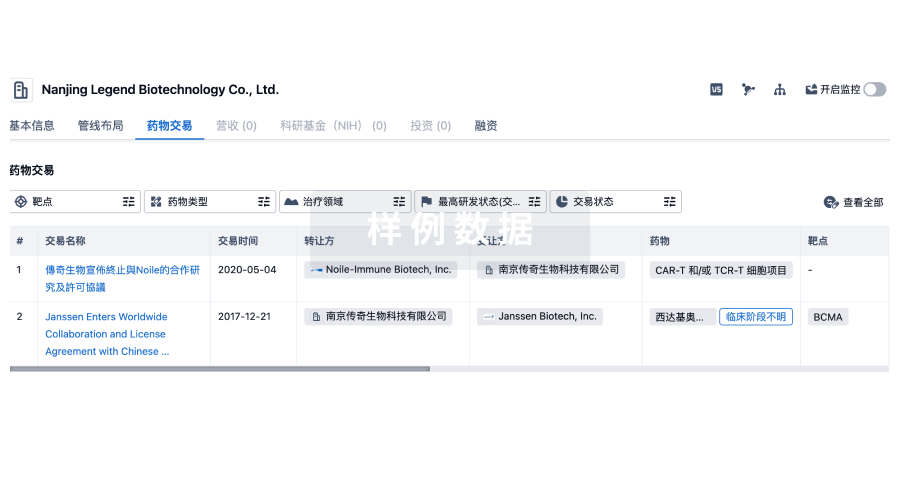
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
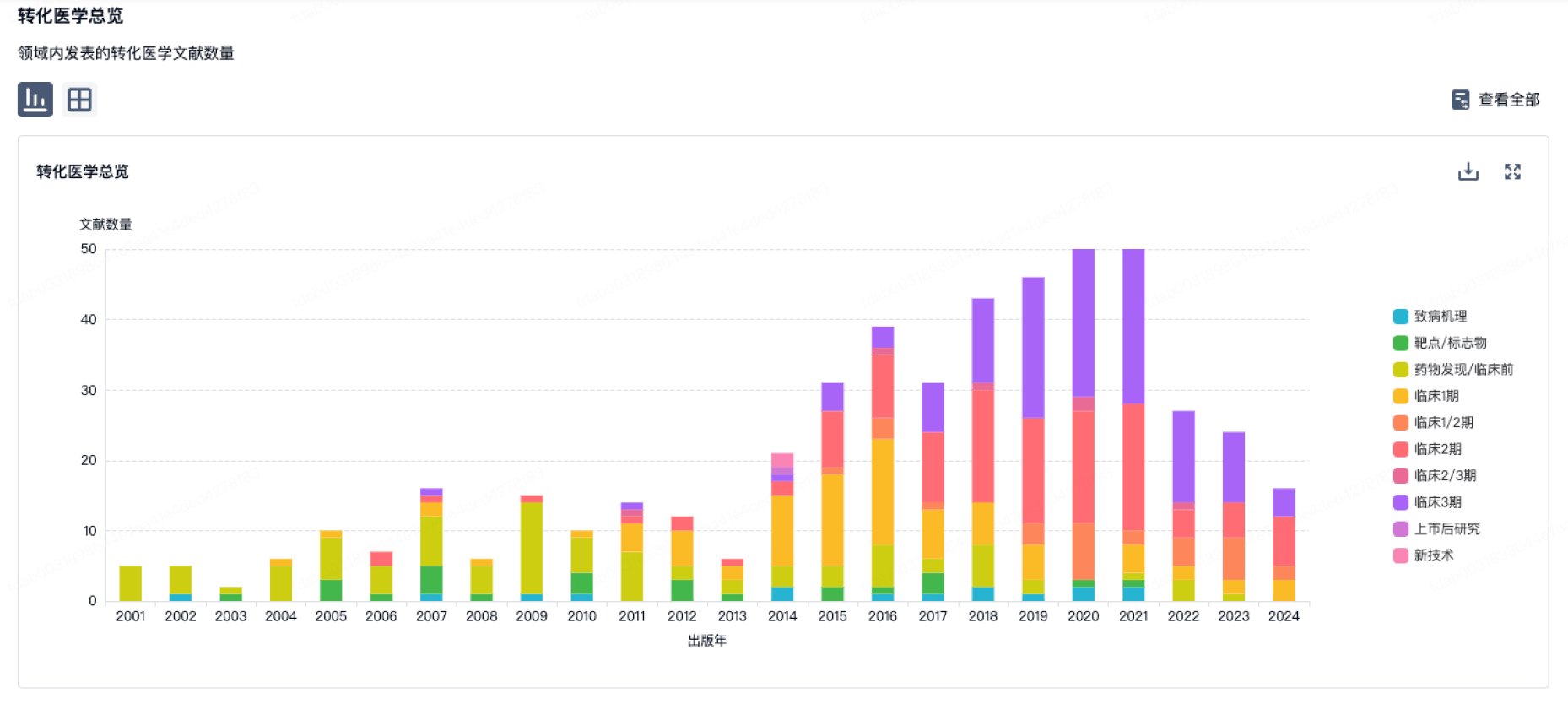
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
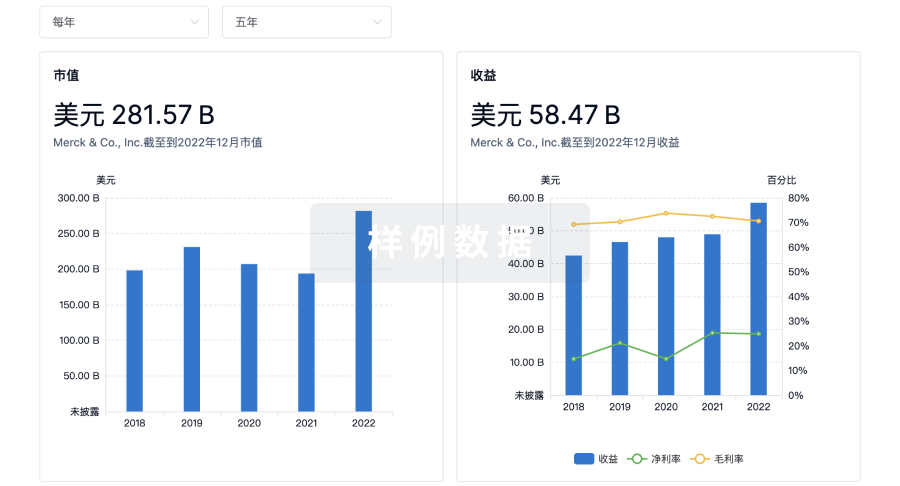
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
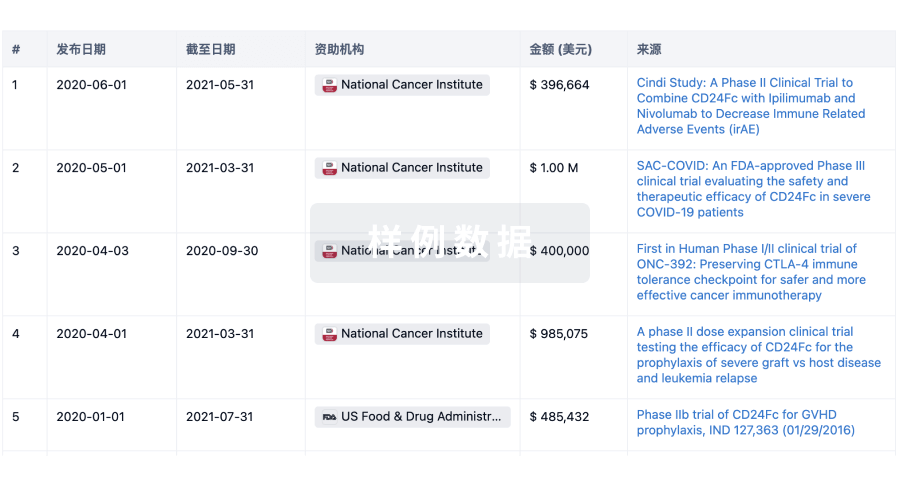
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
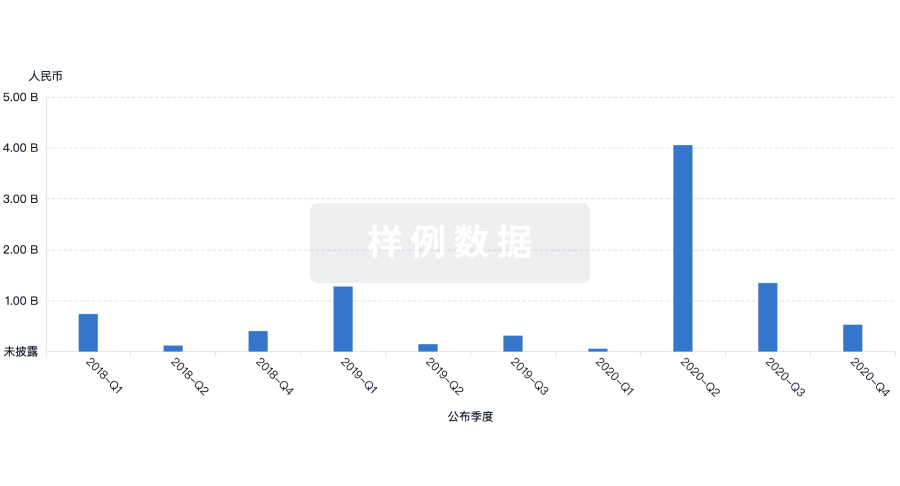
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
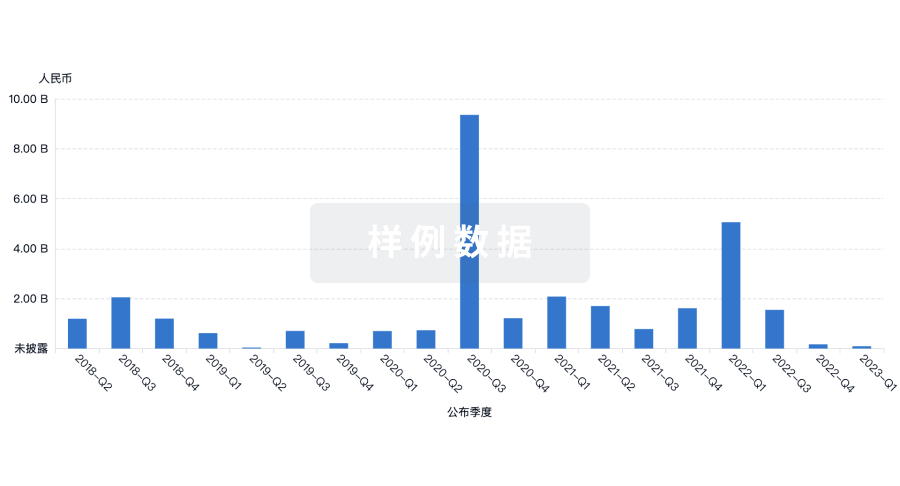
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用