预约演示
更新于:2025-08-21

Genomics Ltd.
更新于:2025-08-21
概览
标签
肿瘤
ADC
关联
1
项与 Genomics Ltd. 相关的药物作用机制 CD276抑制剂 [+1] |
最高研发阶段临床2/3期 |
首次获批国家/地区- |
首次获批日期- |
1
项与 Genomics Ltd. 相关的临床试验NCT05294419
The Healthcare Evaluation of Absolute Risk Testing Study: A Multi-centre, Single Arm, Pragmatic Study in Primary Care Setting
The aim of this study is to demonstrate the integration and use of cardiovascular disease (CVD) integrated risk tool (IRT) in an environment as close to real-world as possible.
This study will recruit participants of both biological sexes and any ancestry or background who require and are eligible for a CVD risk assessment as part of the NHS Health Check. Those aged 45-64 years are most likely to benefit from CVD IRT and will be included in the study, as they are more likely to be asymptomatic but also derive most benefit from preventative measures.
The study will be conducted in GP surgeries as the CVD IRT will have its greatest impact if incorporated into primary care practice for early identification of patients at highest risk.
This study is a device performance evaluation.
This study will recruit participants of both biological sexes and any ancestry or background who require and are eligible for a CVD risk assessment as part of the NHS Health Check. Those aged 45-64 years are most likely to benefit from CVD IRT and will be included in the study, as they are more likely to be asymptomatic but also derive most benefit from preventative measures.
The study will be conducted in GP surgeries as the CVD IRT will have its greatest impact if incorporated into primary care practice for early identification of patients at highest risk.
This study is a device performance evaluation.
开始日期2021-11-04 |
申办/合作机构 |
100 项与 Genomics Ltd. 相关的临床结果
登录后查看更多信息
0 项与 Genomics Ltd. 相关的专利(医药)
登录后查看更多信息
175
项与 Genomics Ltd. 相关的新闻(医药)2025-08-11
·循因缉药
全球视野,深度视角
各位亲爱的股东,大家早上好中午好晚上好!
更快更全的更新在星球:<循因缉药 Elite>,加入方法如下。另外读者群继续开放,欢迎加微信。
微信号!
星球在这里!
各位,小提问:
2025年8月7日这一天是不是犯了天条了?
特么怎么这么多我们覆盖的公司在这一天发财报?让不让人活了...
再加上8.6和8.8发的...只能一个个看了...今天就“宠幸”10x Genomics吧。
顺便提一嘴,咱们单细胞&空间群还有位置,速来。
#01
“重回巅峰”
根据10x Genomics发布的2025Q2财报,本季度营收1.729亿美元,同比增长12.9%,环比增长11.6%。
回顾过去2年的营收情况,我们发现这已经接近这两年10x Genomics营收的极值,仅次于2023Q4的水平。
更为关键的是,10x Genomics竟然扭亏为盈了!
虽说我们预估10x Genomics即将扭亏为盈,但是2025Q2的3453.8万美元净利润着实吓到了我们。
那么,这意味着10x Genomics的经营出现本质的大逆转吗?
并没有。
此前,10x Genomics与Bruker就NanoString相关产品的专利侵权诉讼达成了和解。
本季度,10x Genomics将这6800万美元分别将2730万美元计入授权与版税收入和4070万美元计入和解金收入(我怎么记得要2025Q3才到账呢?)。
如果剔除这两部分,10x Genomics的亏损将达到3346.2万美元,营收也将下滑至1.456亿美元。
即,营收同比下滑4.9%,环比下滑6.0%。
当然,拿到手了就死拿到手了,这么算也有点对10x Genomics太苛刻了。
但是,当我们回看其业务本身的时候,出问题了。
#02
“有点小问题”
我们先看仪器设备的销售情况。
从图上可以看出,不论是来自单细胞Chromium的572.7万美元,还是空间生物学的877万美元,都呈现出一种持续下滑的态势。
这反映出市场对于采购新设备意愿缺失,甚至对于目前强推的空间生物学设备都兴趣缺缺。
好在,从试剂耗材上来看单细胞业务和空间生物学业务都在增长。
尤其是空间生物学试剂耗材,一扫2025Q1的颓势,达到3639.7万美元,同比增长24.4%,环比增长。
但是,这一增长也有一定的水分。
在这之前,我们先看看全球各区的表现。
大头来自美国的1.035亿美元,同比增长15.4%,环比增长19.2%。
但...但是,这里面包含了Bruker给的专利版税。
所以,最靓的仔应该是中国大区。
本季度,中国区营收2317万美元,同比增长68.7%,环比增长37.2%。
简直逆天!
接下来,你能想到的就是但是要来了。
其实,这一增长部分是包含由于中美关税战造成的提前备货引起的。
按照10x Genomics的估算提前备货的规模大概在400万美元左右,如果剔除这部分,中国区仍然增长,但是就没这么夸张了。
再加上中国区华大智造、新格元、寻因生物、M20们的围追堵截,美国方面Illumina收购Fluent BioSciences加入战团。
我要是10x Genomics,我肯定急了。
#03
“急了”
急了的体现,就在于10x Genomics以3000万美元对价股票+现金的形式收购Scale Biosciences。
后续还有里程碑付款,咱们就不展开了。
我们得先看Scale Bio的技术是什么,再来看10x Genomics收购他的逻辑。
公司创始人有3个,Jay Shendure,Cole Trapnell和Garry Nolan。
(创始人Jay Schendure也是单细胞方面造诣颇深,大家可以搜索下很有意思)
Scale Bio原先并不是叫这个名字,他原名是创立于2011年的Apprise。
2017年被罗氏诊断收购,但是后来又分拆出来,这时候就是Scale Bio了。
公司创始人Garry Nolan,是个牛人。
他是Stanford的Rachford and Carlota A. Harris 讲席教授,创立了Rigel(上市制药公司,主要覆盖血液瘤、实体瘤和免疫疾病)、BINA(生信公司,被罗氏诊断收购)、Ionpath、Akoya(对,就是那个Akoya,被Quanterix并购)。
Scale Bio的技术被称之为Quantum Barcoding Technology,抛开牛X的量子编码技术这个名字,其核心就是Split-seq的变种。
简单的说,就是通过一管细胞,重复拆分+添加不同的sub-barcode来达到足够的区分单细胞的水平。
原理重要,但也没有那么重要。
重要的是,这玩意不需要什么特殊的设备!
大家还记得Illumina收购的Fluent bio的技术是什么特点么?不需要特殊设备!
没错了,这就是冲着Illumina去的。
当然,不仅仅是Illumina一家。
由于不需要特殊设备,10x Genomics可以将单细胞测序技术推向更多的实验室,更加丝滑的切入市场。
甚至,有可能挑战更低的价格底线。
#04
最后-代价呢?
那么,代价是什么呢?
Scale Bio的技术如果得以大规模推广,那么不仅仅是带来此前无法触达市场的增量,还有侵蚀原有10x Genomics产品线的减量。
单细胞测序的设备必然再受冲击,而原先使用的10x Genomics固有试剂耗材也有可能转换门庭使用Scale Bio试剂耗材。
没办法,留给10x Genomics的选项和时间不多了。
财报发布后,10x Genomics股价反应平淡。
10x说预计收购Scale Biosciences不会对2025年剩余时间的收入或运营费用产生重大影响。
未来,计划推出Visium HD XL、Xenium RNA Plus蛋白等,以增强多组学空间分析能力。
咱们继续边走边看。
END
小编在这里!
星球在这里!
各位股东看了么?赞了么?转发了么?别忘了点赞。谢谢!
所有内容均不作为投资建议,信息均来自公开资料,星球更新更快。
近期原创:
PacBio:走出泥潭
Myriad凭什么暴涨46%??!!
Exact Sciences:一个季度营收8.11亿,调高全年指引!投资人:逃!
她启动了曼哈顿计划
Personalis 2025Q2:丸辣!Twist:狂喜!
相关资料:注1:https://investors.10xgenomics.com/news/news-details/2025/10x-Genomics-Reports-Second-Quarter-2025-Financial-Results/default.aspx注2:https://investors.10xgenomics.com/news/news-details/2025/10x-Genomics-to-Acquire-Scale-Biosciences/default.aspx注3:https://d18rn0p25nwr6d.cloudfront.net/CIK-0001770787/ac12fb62-217a-47a6-91b4-43b3fdd4f910.pdf注4:https://s28.q4cdn.com/592666581/files/doc_financials/2025/q2/TXG_Q2-2025_Earnings-Prepared-Remarks-08-07-25.pdf
财报专利侵权
2025-08-08
·今日头条
2025年8月7日——单细胞测序领域再迎重磅整合!10x Genomics宣布,以3000万美元现金+股票首付收购单细胞技术公司Scale Biosciences(以下简称“Scale”),并承诺根据里程碑额外支付金额(未公布)。这笔交易不仅标志着10x在单细胞实验规模化赛道上的关键落子,更被视为单细胞测序从“实验室尖端”迈向“全局普及”的战略转折点。
3000万美元“技术并购”:10x的“规模化野心”与Scale的“终极归宿”
根据10x Genomics首席执行官Serge Saxonov博士透露的交易细节,
此次收购的核心是Scale的两大价值:技术能力与市场潜力。
Scale成立于2018年(前身为罗氏旗下平台技术分支),凭借其专利的“分裂池组合条形码技术”(split-pool combinatorial barcoding),在单细胞测序领域迅速崛起。其核心产品矩阵包括:
QuantumScale单细胞RNA试剂盒:
无需仪器即可完成单细胞条形码标记,大幅降低实验门槛;
ScalePlex:
允许样本在洗涤前混合,减少操作步骤并提升通量;
单细胞甲基化检测试剂盒:
全球首个商业化单细胞甲基化状态检测方案。
“我们对Scale的技术觊觎已久,”Saxonov直言,
“单细胞实验的规模化(scale)是当前行业的最大瓶颈——研究人员渴望一次实验分析数百万甚至上亿细胞,但现有技术的成本、复杂度和通量限制了这一可能。Scale的技术恰好能突破这一限制,与我们Chromium平台的单细胞多组学能力形成完美互补。”
而对Scale创始人兼首席执行官Giovanna Prout来说,这场收购是“理想照进现实”的选择。“作为一家不足50人的初创公司,
我们曾梦想将技术推向全球,但宏观经济寒冬让独立上市的路径愈发艰难。
”Prout坦言,“10x的全球市场覆盖、成熟的商业化能力和技术整合经验,能让Scale的技术更快触达更多研究人员,真正改变单细胞研究的格局。”
从罗氏到10x:Scale的“技术传承”与单细胞领域的“生态融合”
Scale的技术底色深厚,其核心团队的履历堪称“单细胞测序界的‘复仇者联盟’”:
联合创始人Garry Nolan博士是斯坦福大学教授、单细胞测序先驱,曾发明量子条形码技术(2011年专利),并主导了Apprise(后被罗氏收购)和空间转录组公司Akoya Biosciences的创立;
Prout则拥有12年Illumina、4年10x Genomics的行业经验,深度参与过CZI“十亿细胞计划”等行业标杆项目。
这种背景让Scale的技术自带“生态基因”。Nolan博士透露,其2011年发明的量子条形码技术,最初目标是推动单细胞测序的规模化,但受限于罗氏的研发优先级,未能充分释放潜力。“如今加入10x,相当于‘回家’——这里有全球最成熟的单细胞技术平台(如Chromium)和空间转录组技术(如Xenium),能将我们的条形码技术与空间、表观遗传数据无缝整合。”
这种整合正是单细胞领域的下一个“兵家必争之地”。当前,单细胞研究已从“单组学”(仅测基因表达)迈向“多组学”(同时测基因、甲基化、空间位置等),但不同组学数据的整合仍面临技术壁垒。Scale的条形码技术若与10x的空间转录组、表观测序工具结合,有望实现“一个样本、多维度解析”的全场景覆盖,大幅提升研究效率。
行业震荡:单细胞测序从“精英游戏”到“普惠工具”
此次收购的深层背景,是单细胞测序领域的“冰火两重天”:
一方面,技术突破推动其在癌症、神经科学等领域的应用爆发(2024年全球单细胞测序市场规模已达45亿美元,年增长率超30%);另一方面,实验成本高、操作复杂等问题仍限制了其普及——据行业调研,仅15%的实验室能独立完成百万级细胞的单细胞实验。
Scale的“规模化”技术恰好瞄准这一痛点。以其QuantumScale试剂盒为例,研究人员无需昂贵的单细胞分选设备,即可在普通实验室完成高通量单细胞标记,成本较传统方法降低60%。而10x的Chromium平台已覆盖全球超1万家实验室,整合Scale技术后,这一数字有望在2年内翻倍。
“单细胞测序不应是‘少数实验室的奢侈品’,”Prout强调,“通过10x的全球渠道,Scale的技术将进入更多发展中国家、中小型实验室,推动癌症早筛、药物反应预测等临床应用的落地。”
未来已来:多组学整合或成“下一个爆点”
对于收购后的计划,Saxonov表示,10x将分两步走:
短期:
继续支持Scale现有产品的客户,确保技术过渡平稳;
长期:
将Scale的条形码技术与Chromium平台深度融合,开发新一代单细胞多组学解决方案(如“单细胞基因表达+甲基化+空间位置”三联检测)。
Nolan博士则展望了更远的未来:
“想象一下,未来一个肿瘤样本,既能通过单细胞测序解析癌细胞的异质性,又能通过空间技术定位其微环境,还能通过甲基化检测追踪表观遗传变异——这些数据将在10x的平台上无缝整合,为精准治疗提供‘全维度地图’。Scale的加入,让这张地图的绘制速度至少提升10倍。”
从罗氏的研发分支到独立初创,再到融入10x的生态,Scale的故事折射出单细胞测序领域从“技术验证”到“规模化落地”的进化轨迹。而对10x而言,此次收购不仅是扩大市场份额的“战术动作”,更是其“用技术普惠推动生命科学进步”愿景的关键拼图。
【关于投稿】
转化医学网(360zhyx.com)是转化医学核心门户,旨在推动基础研究、临床诊疗和产业的发展,核心内容涵盖组学、检验、免疫、肿瘤、心血管、糖尿病等。如您有最新的研究内容发表,欢迎联系我们进行免费报道(公众号菜单栏-在线客服联系),我们的理念:内容创造价值,转化铸就未来!
转化医学网(360zhyx.com)发布的文章旨在介绍前沿医学研究进展,不能作为治疗方案使用;如需获得健康指导,请至正规医院就诊。
责任声明:本稿件如有错误之处,敬请联系转化医学网客服进行修改事宜!
微信号:zhuanhuayixue
热门推荐活动 点击免费报名
线上|2025年8月12日
▶ “破卷之战:新修饰战队重燃科研创新力”线上交流会
线上|2025年8月20日
▶ 极限突破,先机必争:单细胞全转录组空间多组学技术 CosMx SMI 的前沿解析
北京|2025年9月19-20日
▶ 第六届单细胞技术应用研讨会暨空间组学前沿研讨会
上海|2025年11月20-23日
▶ 第八届上海国际肿瘤内科学论坛(8th SIMOS)
上海|2025年12月10日
▶ 第五届现代临床分子诊断研讨会
点击对应文字 查看详情
并购专利到期
2025-06-28
·生物谷
6 月 28 日,由生物谷、梅斯医学主办,上海市免疫学研究所联合主办的 “2025(第八届)多组学研究与应用前沿论坛”聚焦“技术突破→机制解析→临床落地”全链条,由七位产学研嘉宾分别从代谢组学早筛、肿瘤微环境解析到空间技术应用,勾勒多组学临床转化的清晰路径,为论坛画上圆满句点。多组学临床研究与产业化上海百趣生物医学科技有限公司邓君亮副主任以《新一代代谢组学 NGM 在结直肠癌早筛中的应用》为切入点,揭秘新一代代谢组学技术(NGM)在标准品数据库以及临床检测上的 “双突破” 。复旦大学附属肿瘤医院王建华教授以《单细胞与空间多组学技术在前列腺癌中的应用》为题,系统阐述了通过单细胞转录组联合空间转录组解析前列腺癌微环境异质性的研究体系。北京大学第一医院冀培丰教授作《全组织高分辨空间蛋白组检测方法开发及其应用》报告,重点讲解了基于 CT 成像原理的全组织、高分辨、高通量蛋白检测新技术。上海交通大学医学院单细胞组学与疾病研究中心刘宁宁副主任带来《从群体到单细胞:揭秘瘤内真菌与宿主相互作用致癌的分子机制》报告,通过ITS1 rDNA 扩增子测序、宏基因组测序及单细胞组学技术(SAHMI),解析了瘤内真菌与宿主互作致癌机制。深圳华大生命科学研究院张银博士以《乳腺癌空间图谱及小静脉对免疫浸润的调节作用》为题,分享了单细胞与空间转录组联合构建的乳腺癌组织空间细胞图谱 ,为乳腺癌免疫治疗提供新靶点。10x Genomics Xenium 市场拓展经理赖登攀作《Complementary Spatial Technologies Built to Tackle Biology's Biggest Challenges》报告,介绍了 Xenium 空间分析技术与多组学技术的互补应用方案。M20 Genomics 学术及创新业务总监严青带来《VITA 单细胞转录组:在临床样本和微生物领域的技术突破与科研应用》报告,介绍了VITA 单细胞转录组技术在临床冻存 / FFPE 样本及微生物单细胞测序中的核心突破。当日论坛以 “技术落地为核心,临床价值为导向” ,串联代谢组学早筛、肿瘤微环境解析、空间技术应用等核心议题,既展现 “从 0 到 1” 的技术突破,也凸显 “从 1 到 N” 的产业潜力。随着多组学技术与临床需求深度耦合,这场论坛不仅是前沿成果 “展示窗”,更成为产学研协同转化的 “加速器” ,为精准医疗未来注入持续动能!让我们一起期待下一届多组学会议~未来,随着单细胞时空组学、AI 算法与临床队列的深度结合,多组学技术将在疾病早筛、疗效预测、个体化治疗等领域持续突破,为人类健康事业开启全新篇章。让我们一起期待下一届多组学会议~扫码获取照片直播以及电子会刊,回顾更多精彩报告细节!
100 项与 Genomics Ltd. 相关的药物交易
登录后查看更多信息
100 项与 Genomics Ltd. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年03月21日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
临床前
1
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
Vobramitamab Duocarmazine ( CD276 x DNA ) | CD276阳性实体瘤 更多 | 临床前 |
登录后查看更多信息
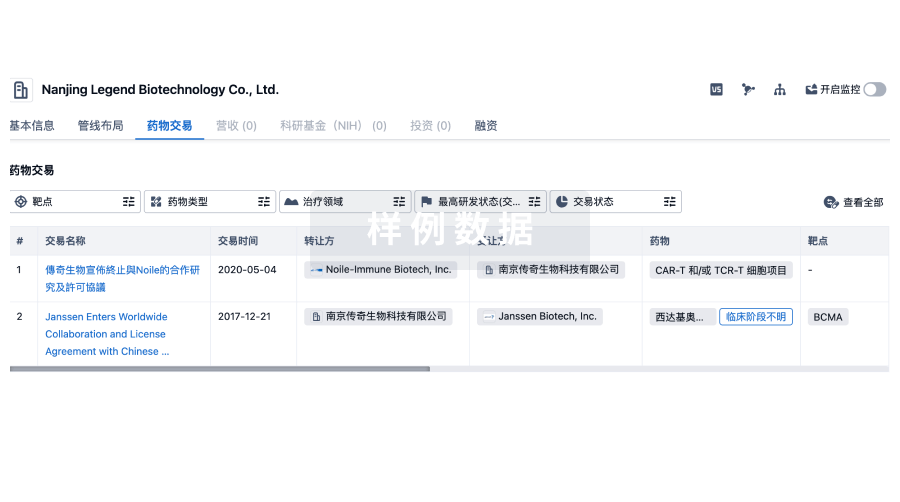
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
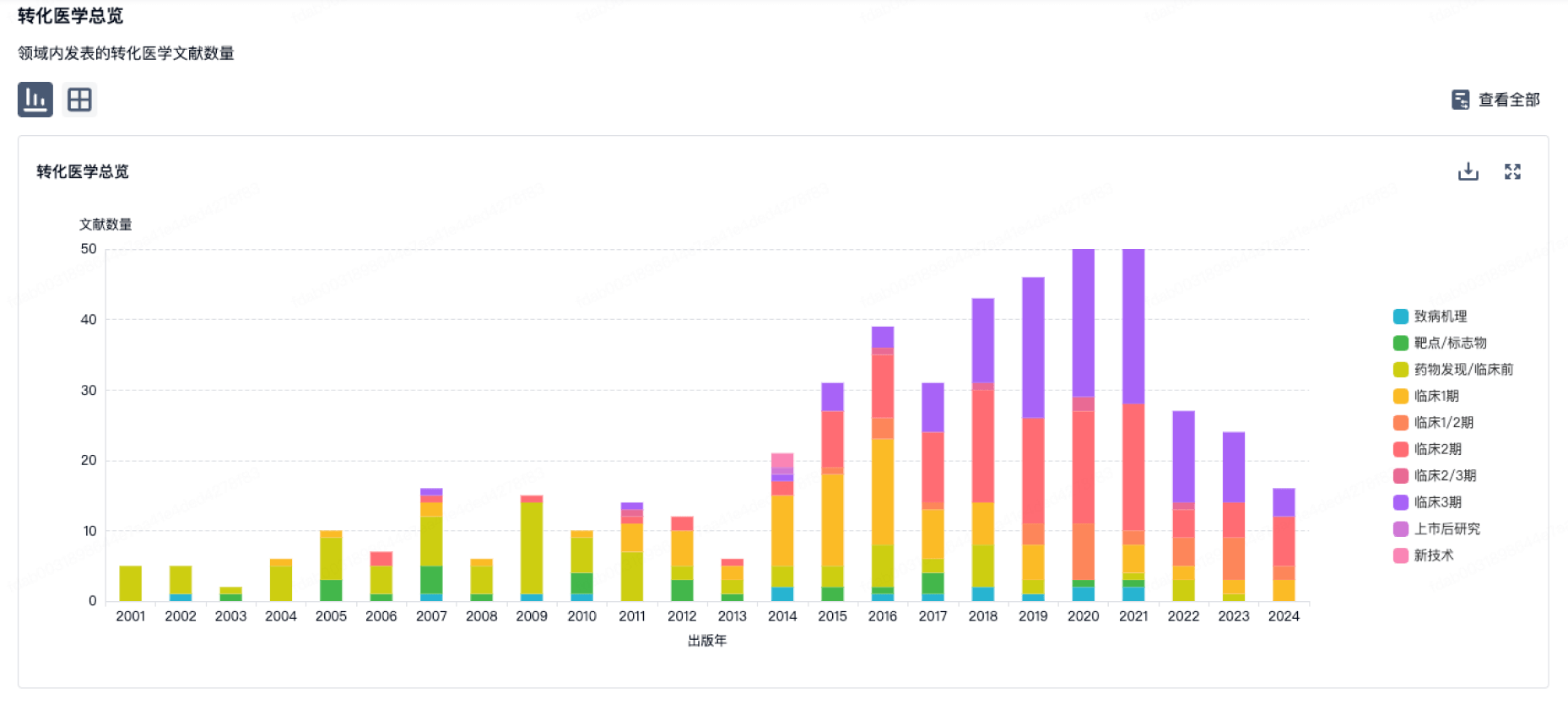
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
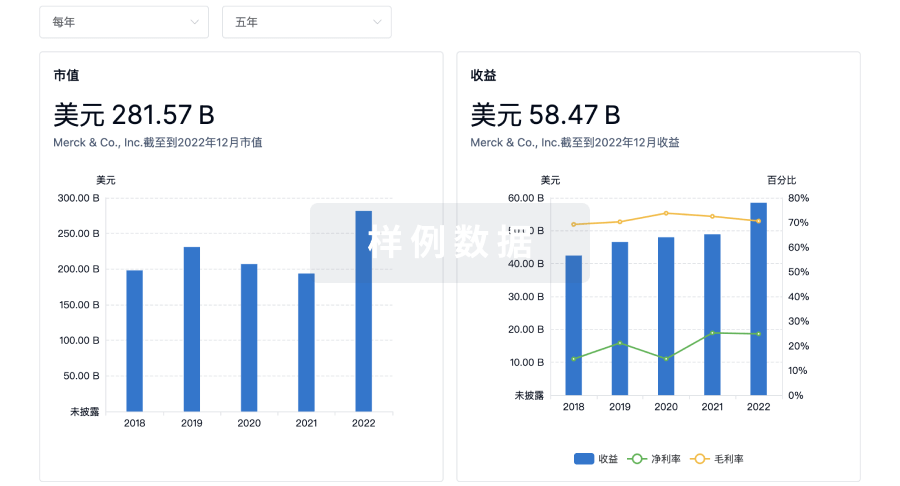
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
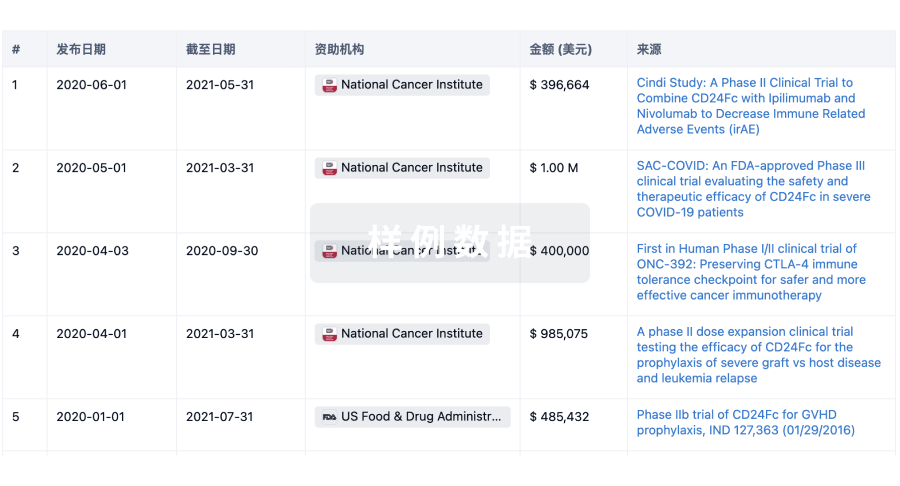
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
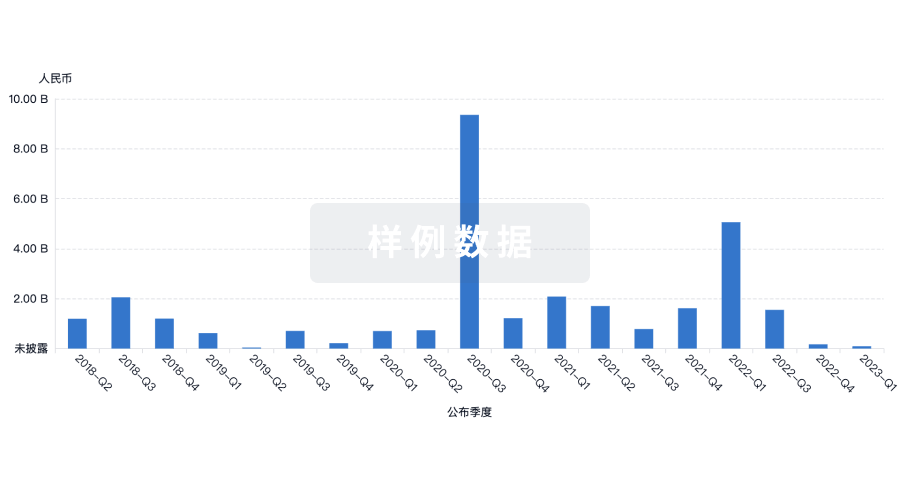
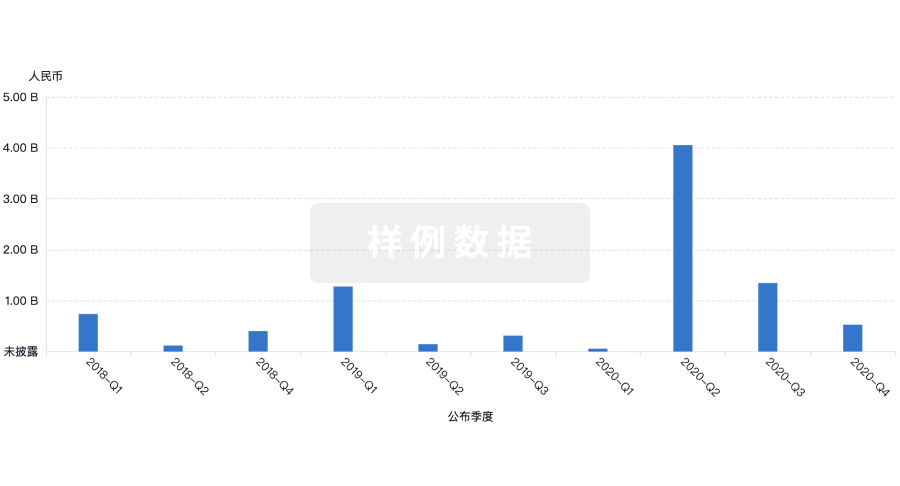
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
