预约演示
更新于:2026-05-30
4-Phenylbutyrate
4-苯基丁酸酯
更新于:2026-05-30
概要
基本信息
原研机构 |
权益机构- |
最高研发阶段临床前 |
首次获批日期- |
最高研发阶段(中国)临床前 |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C10H12O2 |
InChIKeyOBKXEAXTFZPCHS-UHFFFAOYSA-N |
CAS号1821-12-1 |
关联
9
项与 4-苯基丁酸酯 相关的临床试验NCT04531878
Jian-She Wang of Children's Hospital of Fudan University
The purpose of the study is to improve the prognosis of inhereditary cholestasis caused by ABCB11 gene mutations by using BSEP function rescue drugs
开始日期2023-02-08 |
申办/合作机构 |
NCT04421677
Safety and Tolerability of Phenylbutyrate in Inclusion Body Myositis
This is a pilot study (phase 1 clinical trial) to evaluate the safety and tolerability of phenylbutyrate in IBM. In this open label study, 10 patients with sporadic inclusion body myositis will be treated with phenylbutyrate (3 gm twice daily) for 3 months. There will be a run-in period, during which certain biomarkers will be measured at baseline and at the end of the run-in period in addition to final measurement at the end of the treatment period.
开始日期2020-08-20 |
申办/合作机构 |
JPRN-UMIN000012782
Efficacy and safety of 4-phenylbutyrate in refractory cholestatic disease including progressive familial intrahepatic cholestasis, primary biliary cirrhosis, primary sclerosing cholangitis and Alagille syndrome. - Efficacy and safety of 4-phenylbutyrate in refractory cholestatic disease including progressive familial intrahepatic cholestasis, primary biliary cirrhosis, primary sclerosing cholangitis and Alagille syndrome.
开始日期2014-02-01 |
申办/合作机构 |
100 项与 4-苯基丁酸酯 相关的临床结果
登录后查看更多信息
100 项与 4-苯基丁酸酯 相关的转化医学
登录后查看更多信息
100 项与 4-苯基丁酸酯 相关的专利(医药)
登录后查看更多信息
1,715
项与 4-苯基丁酸酯 相关的文献(医药)2026-07-01·CHEMICO-BIOLOGICAL INTERACTIONS
Isopsoralen-induced hepatotoxicity: The regulatory role of endoplasmic reticulum stress-triggered mitochondrial dysfunction and apoptosis
Article
作者: Yu, Yingli ; Lou, Pengshuo ; Wang, Chen ; Yuan, Mingxin ; Zhang, Yue ; Zhou, Kun ; Han, Yiping
OBJECTIVE:
Fructus Psoraleae is a commonly used herb in traditional Chinese medicine, widely applied in the treatment of osteoporosis, vitiligo, and other diseases. Isopsoralen, as the core active component of Fructus Psoraleae, possesses both pharmacological effects and toxic risks, among which hepatotoxicity is a key issue limiting its safe clinical application. Currently, the molecular mechanism underlying isopsoralen-induced liver injury has not been fully elucidated. This study investigates isopsoralen-induced liver injury, focusing on the interaction between endoplasmic reticulum stress (ERS) and mitochondrial dysfunction.
RESULTS:
In HepG2 cells, isopsoralen dose- and time-dependently reduced survival and increased lactate dehydrogenase release. In mice, isopsoralen elevated serum aminotransferases and creatine kinase. RNA sequencing revealed differential gene expression related to ERS and mitochondria. Isopsoralen increased the ERS markers glucose-regulated protein 78 and C/EBP homologous protein in vitro. Co-treatment with the ERS inhibitor 4-phenylbutyrate reduced caspase expression and apoptotic proteins while increasing the anti-apoptotic protein Bcl-2 in mouse liver tissue. 4-phenylbutyric acid also restored the mitochondrial fusion/fission balance, increased adenosine triphosphate content, and attenuated oxidative stress.
CONCLUSION:
Isopsoralen induces ERS, activating downstream apoptotic pathways that lead to mitochondrial dysfunction and subsequent liver injury. Inhibiting ERS alleviates this hepatotoxicity by improving mitochondrial dynamics and reducing oxidative stress.
2026-06-01·COMPUTATIONAL BIOLOGY AND CHEMISTRY
Biomarker discovery and drug repurposing in hepatocellular carcinoma through transcriptomics, machine learning, network pharmacology, and molecular dynamics
Article
作者: Khan, Najeeb Ullah ; Unar, Ahsanullah ; Alfaifi, Mohammed ; Kamli, Hossam
This study employed an integrative computational and systems biology framework to define a diagnostic gene signature for hepatocellular carcinoma (HCC) and to explore its potential translational relevance in a hypothesis-generating manner. Differential expression analysis of transcriptomic data from 230 samples identified 2748 significantly differentially expressed genes (DEGs), including 2283 upregulated and 465 downregulated genes, with FGF4 (log2FC = 10.08) and REG1B (log2FC = 10.02) among the top hits. Four machine learning classifiers were trained using this signature and demonstrated consistently high predictive performance, with XGBoost emerging as the top-performing model (accuracy = 0.97, F1-score = 0.96, ROC-AUC = 0.981). Logistic Regression (L1) and Random Forest achieved comparable performance (ROC-AUC = 0.980 and 0.979, respectively), while SVM-linear also showed high robustness (ROC-AUC = 0.978). All models showed good calibration, with low Brier scores (<0.04) and precision consistently exceeding 0.90 across most recall thresholds, indicating strong but not perfect classification performance. SHAP-based explainability analysis was used to rank and prioritise the most influential predictors, refining the biomarker panel to 81 genes that collectively accounted for approximately 50 % of the model's explanatory contribution, and highlighting key downregulated predictors in HCC, including GDF2, COLEC10, BMP10, LRAT, and DNASE1L3. Protein-protein interaction and functional enrichment analyses revealed five major molecular clusters and provided systems-level insights into dysregulated biological processes associated with HCC. Drug-gene interaction mining mapped 78 target proteins to clinically relevant compounds, including tolrestat, alcuronium, metyrosine, and 4-phenylbutyric acid. Molecular docking suggested favorable binding propensities for several complexes, including alcuronium-3UON (-8.5 kcal/mol), tolrestat-1ZUA (-8.3 kcal/mol), metyrosine-2XSN (-6.7 kcal/mol), and 4-phenylbutyric acid-2NZ2 (-5.9 kcal/mol). A 100 ns molecular dynamics simulation of the tolrestat-AKR1B10 (1ZUA) complex indicated structural stability, with protein backbone RMSD stabilising at 1.5-3.0 Å, ligand RMSD at 0.6-1.4 Å, and persistent interactions involving Trp22, His110, Glu111, and Phe122. Physicochemical and pharmacokinetic profiling further prioritised tolrestat as a computationally favourable candidate (MW = 357.35, LogP = 3.64, TPSA = 81.86 Ų), exhibiting acceptable drug-likeness, high predicted gastrointestinal absorption, and low synthetic complexity (SA = 2.34), in contrast to alcuronium (MW = 666.89, SA = 7.86), which showed multiple rule violations. Collectively, this in silico study proposes a robust diagnostic gene signature for HCC and identifies tolrestat as a promising repurposing candidate that warrants experimental validation, demonstrating the utility of integrating machine learning, network biology, and molecular simulation in translational cancer research.
2026-06-01·TISSUE & CELL
Mechanism of endoplasmic reticulum stress: Insights for potential therapeutic benefits in burns
Review
作者: Ren, Zhiyang ; Tang, Shuhan ; Li, Ke ; Zhu, Bin
Burns are injuries to the skin and deep tissues caused by thermal effects, which can trigger inflammation, necrosis, and systemic pathological reactions, often leading to life-threatening complications such as shock, infection, and multi-organ failure. Endoplasmic reticulum stress (ERS) is a key driving factor for tissue damage after burns. Burns are proposed to induce ERS through multiple mechanisms, including protein denaturation, inflammatory cytokine storms, and metabolic disturbances, Under stress conditions, cells can activate three key sensors, IRE1, PERK, and ATF6, to fully activate the main signaling pathway of unfolded protein response (UPR). In severe burns, sustained ERS promotes cell death via CHOP-mediated apoptosis, programmed necrosis, and ferroptosis (activated by the PERK-eIF2α-ATF4 pathway), exacerbating tissue injury. The progressive elevation of endoplasmic reticulum stress markers such as GRP78 and CHOP during burn course has been proven to be an important biological indicator for evaluating the degree of secondary organ damage. Consequently, therapeutic interventions such as propranolol (to counteract catecholamine effects) and 4-PBA (to stabilize protein folding) are under investigation as potential approaches for improving burn outcomes. This review systematically examines the pathological mechanisms of burn-induced ERS, associated modes of cell death, clinical manifestations of organ dysfunction, potential drug therapies, and other burn-related stress responses.
35
项与 4-苯基丁酸酯 相关的新闻(医药)2026-05-14
HONG KONG, May 13, 2026 /PRNewswire/ -- METiS TechBio (7666.HK), a tech-bio company focused on AI-powered nanodelivery innovation, today successfully listed on the Hong Kong Stock Exchange (HKEX), becoming the world's first publicly listed AI-powered drug delivery company and the first AI-powered large-molecule biopharmaceutical company listed on the Hong Kong market. With the goal of becoming the "SpaceX of pharmaceuticals," METiS TechBio has, within six years of its founding, become the fastest unicorn in China's AI pharmaceutical sector to reach the IPO milestone.
Continue Reading
From left to right: Jin Li, Independent Non-Executive Director; Serena Shao, Head of Capital Markets & External Relation; Wenshou Wang, Co-Founder, Executive Director and COO; Dr. Chris Lai, Co-Founder, Chairman and CEO; Hongming Chen, Co-Founder, Executive Director and CRDO; Alan Fu, CFO; Peter Edward Lobie, Independent Non-Executive Director; and Frank Yee Chon Lyn, Independent Non-Executive Director, all of METiS TechBio.
METiS TechBio's IPO was jointly sponsored by Jefferies, Deutsche Bank Securities Asia and CITIC Securities (Hong Kong). A total of 201,229,000 H shares were offered globally, raising over HKD 2.1 billion in proceeds and marking the largest healthcare IPO fundraising in Hong Kong so far in 2026. The Hong Kong public offering was oversubscribed by more than 6,900 times, locking up over HKD 730 billion in subscription funds, representing the highest subscription amount for the Hong Kong public offering among healthcare IPOs in Hong Kong so far in 2026. For the international placing, the Company received orders from more than 280 institutional investors and recorded 82 times oversubscription for the allocable tranche, also marking the highest international placing oversubscription multiple among healthcare IPOs in Hong Kong so far in 2026.
METiS TechBio's Hong Kong IPO once again set a record for AI pharmaceutical IPOs in Hong Kong with a top-tier lineup of 18 cornerstone investors subscribing for a total of USD 148 million, securing comprehensive support from "global asset managers + specialized healthcare funds + AI technology funds + state-level funds + Chinese public fund managers." Among them, BlackRock, the world's largest asset manager, led the subscription with USD 50 million. Global asset management giants and ultra-long-term funds including UBS Asset Management Singapore, Mirae Asset of South Korea, and ORIX Corporation of Japan collectively made allocations. Guofengtou Innovation Investment Fund, a state-level fund, made its first investment in the AI pharmaceutical sector. Internationally renowned specialist healthcare funds, including Deerfield, RTW and Lake Bleu Capital, participated in the investment, while leading technology investment institutions, including Walden International, Hillhouse Capital and IDG Capital, also expanded their presence into the sector. Top Chinese public fund managers including GF Fund, ICBC UBS Asset Management, China Asset Management, and Fullgoal Fund also made a rare joint appearance as subscribers with lock-up commitments. This fully demonstrates that AI nanodelivery technology has risen to the level of a national strategic priority and key area of support, and has become a critical technology driving future growth and breakthroughs in global biopharmaceuticals.
Dr. Chris Lai, Co-Founder, Chairman and Chief Executive Officer of METiS TechBio, stated:
"The future of biomedicine will no longer be simply about 'taking medicine when one falls ill.' METiS TechBio's ambition is to harness AI to build nano-rockets that can navigate with precision through the inner space of the human body's 30 trillion cells, write the code of nucleic acids and proteins into cells, and reprogram diseased and aging cells into healthy cells. This was our founding aspiration, and it is the mission to which we will dedicate our lives. The IPO marks a new starting point for us to accelerate forward, and we will strive to live up to the support and trust we have received from all sectors."
Proprietary World-First AI Nanodelivery Platform, Creating the First-Ever Data, Algorithms, and Models for AI for Nano
METiS TechBio's core competitiveness stems from NanoForge, its proprietary world-first AI nanodelivery platform. The Company has built the world's largest ionizable lipid library at the scale of tens of millions, independently developed the world's only algorithm for de novo lipid generation and lipid language model, established the world's first end-to-end lipid and lipid nanoparticle (LNP) screening platform, and created the world's first AI-powered multiscale simulation platform for small-molecule formulation development.
NanoForge encompasses AI foundation models, METiS AI agents, quantum chemistry and molecular dynamics simulations, and an AI-driven high-throughput screening platform, serving as the foundation of the Company's proprietary AI-driven nanotechnology innovation system. Building upon NanoForge, the Company has developed three technology solutions – AiTEM, AiLNP, and AiRNA – which can not only significantly shorten drug development timelines but also enhance drug safety and efficacy.
Differentiated Innovative Pipeline Covering Major Disease Areas Including Immunology, Oncology and Metabolic Diseases, Supported by a World-First In Vivo Targeted Delivery System
Based on the NanoForge platform, METiS TechBio has taken the lead in the industry in achieving precision targeted delivery to eight key organs and tissues: the liver, lungs, heart, muscles, tumor tissues, immune system, central nervous system, and gastrointestinal tract. Its pipeline products cover oncology, immunology, central nervous system disorders, metabolic diseases, and other areas.
The Company has more than 10 pipeline products, including several candidates at the discovery stage, four preclinical candidates, three clinical-stage products, one pre-NDA product, and two animal health products. As of the Latest Practicable Date, the Company had filed a total of 224 patent applications and had been granted 52 patents.
METiS TechBio has developed MTS-004, China's first AI-enabled formulation drug to complete a Phase III clinical trial. MTS-004 is the first and only PBA (pseudobulbar affect) drug in China to have completed clinical trials, and is expected to fill the gap in PBA drug treatment in China. The MTS-004 program took 38 months from project initiation to completion of the Phase III clinical trial. Through AiTEM's predictive analytics and advanced modeling technologies, the Company successfully reduced preclinical formulation development time from one to two years to less than three months.
The "rocket + satellite" next-generation in vivo immunotherapy paradigm is a core capability through which METiS TechBio advances the development of its differentiated innovation pipeline. MTS-105 is a benchmark pipeline product designed to efficiently activate anti-tumor immunity within specific organs in vivo, with the potential to become the world's first in vivo mRNA-encoded TCE therapy for solid tumors, for the treatment of liver cancer and other advanced solid tumors with liver metastases. Leveraging its AiLNP and AiRNA platforms, METiS TechBio enables highly efficient expression of TCE in hepatocytes and liver cancer cells through specific liver-targeted LNP delivery, activating potent intratumoral immunity and thereby achieving powerful tumor cell killing. MTS-105 is currently in the IIT stage and has received Orphan Drug Designation from the U.S. FDA.
Dual Engines of Platform Collaborations and Product Partnerships, Reinforcing Barriers Through a Closed Loop of
Technology
Iteration
, Commercial Application, and Real-World Data Feedback
In terms of commercialization strategy, METiS TechBio adopts a "platform collaboration + product partnership" dual-engine business model, forming a synergistic and recycling ecosystem of "technology iteration – commercial application – real-world data feedback." To date, METiS TechBio has more than 30 pharmaceutical and biotechnology partners worldwide, with customers primarily including global top-tier pharmaceutical companies, innovative biotechnology companies, and medical research institutions.
METiS TechBio recorded revenue of RMB 105 million in 2025, primarily from the upfront payment for the MTS-004 product partnership. The total milestones for MTS-004 in the PBA indication amount to RMB 1.845 billion, with additional potential milestone payments of up to RMB 100 million for potential indication expansion. Meanwhile, a single-target contract in existing platform collaborations is valued at up to USD 109 million. The dual-engine business model has been preliminarily validated by the market.
Core Senior Executives at the Founding-Team Level Joining Forces Across Disciplines, with Sustained Heavy Investment in R&D and Talent Development
METiS TechBio's R&D team is led by its three co-founders. Building upon the Company's R&D centers in Beijing and Hangzhou, the team brings together multidisciplinary expertise spanning nanomaterials, chemistry, biology, physics, computational science, medicine, and other fields, providing strong support for innovation in AI nanodelivery and cross-disciplinary breakthrough research. The R&D team comprises more than 100 scientists and technical personnel, including approximately 40 Ph.D. holders. The majority of the Company's operating expenses are R&D-related, primarily including employee benefits for scientists and engineers, share-based compensation expenses, procurement of laboratory materials and equipment, and professional service fees (including fees paid to CROs). From 2023 to 2025, the Company maintained sustained high R&D expenditure of approximately RMB 270 million.
METiS TechBio will always prioritize R&D as a key direction for strategic investment. Approximately 50.0% of the net proceeds from the IPO will be used to support core technology research, development, and advancement of its AI infrastructure and AI-driven nanomaterials platform; approximately 20.0% will be used for ongoing and planned clinical trials in its AI-developed product pipeline, advancing candidate drugs across multiple therapeutic areas and modalities; and approximately 10.0% will be used to develop animal health and anti-aging solutions and to expand AI-enabled solutions into these high-growth areas.
SOURCE METiS TechBio
21%
more press release views with
Request a Demo
临床3期孤儿药
2026-05-07
·华津生物
点击蓝字 关注“华津生物”
足细胞损伤是多种肾小球疾病的核心病理机制,然而传统体外模型在模拟足细胞三维微环境及成熟表型方面存在明显局限。近日,美国西奈山伊坎医学院肾脏病学部 Kristin Meliambro 教授团队 在 《Nephrology Dialysis Transplantation》 上发表综述,系统整合了肾脏类器官在足细胞病研究中的最新进展。
区别于既往以肾小管为主的类器官综述,该文聚焦足细胞损伤建模,涵盖药物诱导损伤、遗传性肾病、复杂肾小球疾病及自身免疫性足细胞病等多个维度,并在此基础上评价了当前模型的优势与不足,提出了提升类器官保真度、成熟度和功能整合性的关键方向,为肾脏精准医学研究提供了重要参考。
(点击可放大)
表1. 足细胞病研究中不同体外系统的比较,涵盖永生化足细胞系、原代足细胞培养、干细胞来源足细胞及肾脏类器官。
(点击可放大)
图1. 肾脏类器官与体内肾小球结构对比示意图。
药物诱导性足细胞损伤:
从概念验证到药物筛选
肾脏类器官最早被用于复现已知的肾小球损伤模型。Zhou et al在类器官中成功重现了嘌呤霉素氨基核苷(PAN)引起的足细胞损伤,观察到nephrin、podocin和synaptopodin的表达均出现下调,并进一步证明TRPC5抑制剂AC1903能够有效改善上述变化。Nguyen et al的研究则证实了PAN所造成的足细胞损伤具有剂量依赖性。此外,脂多糖诱导(LPS)的足细胞损伤还伴随着促炎因子释放和氧化应激通路的激活,而甲泼尼龙可部分减轻这些损伤效应。
其他研究利用肾脏类器官来研究临床相关药物的时间进程和肾毒性潜力。以阿霉素为例,该化疗药物在啮齿类中可稳定诱发FSGS,但在人类中致病程度较低。为明确其对足细胞的直接毒性,多项研究在类器官中进行了系统验证。Hale et al发现低剂量阿霉素即可导致MAFB丢失、caspase 3激活及肾小球体积缩小;Kumaret al进一步证实微型类器官中足细胞TUNEL阳性,NPHS1与SYNPO表达降低;Bajajet al证明阿霉素选择性损伤肾小球而不影响肾小管标志物;Dilzet al则通过多维度量化确认了上述效应。这些研究共同表明,肾脏类器官能以生理相关方式有效评估药物的足细胞毒性。
遗传性肾小球疾病:
从机制解析到基因治疗
肾脏类器官已成为研究遗传性肾小球疾病的重要平台。Tanigawa et al针对NPHS1基因,利用患者来源诱导多功能干细胞(iPSC)揭示了不同突变类型的差异化表型:错义突变(E725D)导致nephrin虽能表达,但无法正常定位至足细胞膜表面;截断突变则使nephrin完全无法被检测到,而基因校正可逆转上述缺陷。Hale et al在携带NPHS1复合杂合突变的类器官中进一步证实nephrin与podocin蛋白水平均降低。Dorison et al则针对NPHS2突变,展现了变异特异性的podocin表达减少、运输障碍及与nephrin相互作用异常。此外,Freedman et al发现PODXL功能丧失突变可在类器官中诱导显著的足细胞结构与分子缺陷,与后续关于罕见家族中突变PODXL相关FSGS的报告一致。
类器官在法布里病和Alport综合征(AS)的建模中也展现出重要价值。在法布里病方面,Kim et al利用CRISPR编辑的GLA突变类器官重现了足细胞中三己糖酰基鞘脂醇(Gb3)积聚、氧化应激及凋亡等特征,且酶替代及抗氧化治疗可部分缓解。在Alport综合征方面,研究从建模优化逐步推进至治疗验证:Morais et al首先证实AS类器官虽基底膜不成熟但仍保留组装时序;Hirayama et al通过振荡培养获得更成熟的肾小球基底膜(GBM)结构,成功再现疾病特异性胶原缺陷,并证明化学伴侣4-苯基丁酸(4-PBA)可部分恢复错义突变类器官中Ⅴ型胶原α5链表达。在此基础上,Yabuuchi et al与Saei et al进一步探索外显子跳跃和剪接调控等基因治疗策略,体外均实现了COL4A5表达的部分恢复,支持了肾脏类器官系统用于评估遗传性肾小球疾病基因治疗效用的潜力。
对于致病机制尚存争议的INF2突变,Subramanian et al联合肾脏类器官、小鼠模型和原代培养体系,证明其通过功能获得效应驱动足细胞损伤。类器官中表现为nephrin核周错位、线粒体结构异常及细胞-基质粘附减弱,且在PAN刺激后进一步加剧,与小鼠模型中的发现一致,体现了类器官在解析突变致病机制中的整合价值。
复杂疾病建模:
遗传背景与环境因素的整合
肾脏类器官为研究需要“二次打击”的复杂遗传风险因素提供了理想系统。APOL1高风险变异是典型范例:该基因在小鼠中不表达,长期限制体内研究。Liu et al利用CRISPR-Cas9构建了首个APOL1 G1类器官模型,单细胞转录组分析揭示IFN-γ可诱导APOL1表达,但仅在G1背景联合内质网应激后才出现明显细胞应激与去分化。Chun et al与Song et al进一步阐明了脂滴积聚、氧化磷酸化下降及线粒体功能障碍等下游机制。在治疗层面,Nystrom et al证明JAK1/2抑制剂巴瑞替尼可阻断该信号轴、保护足细胞活力;Juliar et al则将视角拓展至内皮,发现IFN-γ可提前于足细胞损伤启动内皮焦亡与网络崩解,而JAK/STAT抑制剂可阻断上述过程。
COVID-19新冠疫情期间,肾脏类器官被迅速用于探究SARS-CoV-2是否能直接感染足细胞。Helms et al报告足细胞仅被最低限度感染,可能与ACE2低表达有关;而Jansen et al与Garreta et al则提供了直接感染证据,并发现促纤维化通路激活可能增加远期慢性肾病风险。Garreta et al进一步证明高糖环境可上调ACE2表达并重编程足细胞代谢,为糖尿病患者中观察到的COVID-19相关肾脏并发症高发提供了机制解释。这些研究体现了肾脏类器官在公共卫生危机中快速响应、机制性解析复杂疾病的能力。
肾脏类器官还被用于模拟慢性代谢性疾病背景下的肾小球损伤。Garreta et al首次将类器官暴露于振荡高糖条件,成功诱导出细胞外基质沉积、胶原与纤连蛋白上调及线粒体损伤等早期糖尿病肾病特征。Veloso Pereira et al则从衰老角度切入,证明急性足细胞损伤可启动CDKN2A等衰老相关基因表达,驱动肾小球老化进程。这些研究提示,肾脏类器官具备整合多重应激源模拟慢性全身性疾病的潜力,但能否稳定再现体内肾小球损伤特征仍需进一步验证。
自身免疫性足细胞病:
新兴方向的探索
在此期间,动物研究可能从主要的证据基础,逐步演变为校准NAMs预测性能的参考系统,最终成为仅在必要时使用的补充信息。在自身免疫性足细胞病这一新兴领域,Gupta et al利用肾脏类器官建立了原发性FSGS(pFSGS)循环致病因子的功能读数平台。研究发现,pFSGS患者血浆可直接诱导类器官足细胞出现凋亡、损伤标志物上调、基质沉积增加及炎症因子分泌增强。值得注意的是,无论是移植后复发FSGS还是pFSGS患者,其血浆均可诱发类器官足细胞损伤,而后者经血浆置换后即丧失诱导类器官足细胞损伤的能力,这与临床观察高度一致,进一步验证了该模型的转化应用价值。
(点击可放大)
图2. 已发表的肾脏类器官应用于足细胞损伤建模研究示意图汇总。
优势与局限并存:
肾脏类器官在足细胞病建模
中的进展与突破路径
早期类器官研究主要复现PAN、LPS及阿霉素等损伤因素对足细胞的毒性,为药物筛选提供了概念验证,但这些模型多依赖人工损伤条件,转化相关性尚不明确。随着分析方法与分化方案不断改进,人源化高通量肾毒性筛选平台已初具雏形,其临床价值仍待进一步验证。
相比之下,遗传性足细胞病是肾脏类器官最具优势的应用领域。通过患者来源iPSC与CRISPR基因编辑技术相结合,研究者已在单基因足细胞病、Alport综合征和法布里病中成功实现了可重复、可量化的疾病表型建模,并在此基础上测试了多种治疗策略。由于类器官内源性表达足细胞关键基因,其在机制解析方面具有天然优势。以APOL1特定风险变异型肾病(AMKD)相关研究为例,类器官不仅验证了内皮损伤假说,所揭示的信号通路更直接推动了巴瑞替尼进入2期临床试验,充分体现了类器官平台在转化效率和可扩展性方面相较传统动物模型的独特价值。
然而,类器官系统仍存在一些固有局限,在模拟糖尿病等全身性疾病以及衰老过程时尤为明显。由于缺乏循环免疫细胞、激素和代谢输入,加之血管网络发育不全,类器官尚无法模拟肾小球滤过功能,也难以研究血流动力学因素和肾小球内细胞间的相互串扰。不过,这些局限在某些领域亦可转化为优势。在自身免疫性足细胞病研究中,免疫系统的缺失使得研究者能够独立评估自身抗体和循环因子对足细胞的直接致病作用,Gupta et al已初步验证了这一思路的可行性。此外,类器官保留了nephrin等裂孔隔膜蛋白的表达,这一优势恰恰是永生化足细胞系所不具备的。
(点击可放大)
表2. 肾脏类器官在肾脏疾病(包括肾小球与非肾小球疾病)建模中的优势与局限概览。
针对上述局限,研究者正从多个方向探索改进策略。振荡培养有助于促进肾小球基底膜成熟,补充血管内皮生长因子可增强血管化程度,内皮细胞共培养与3D打印支架则能改善血管网络的形成,而微流控芯片可将生理剪切力引入培养体系。更为前沿的探索是,将肾单位祖细胞与输尿管芽祖细胞组合生成的肾脏“组装体”,在体内移植后不仅能够形成有功能的血管网络,还可展现出滤过与重吸收等初步的肾功能特征,为构建更高保真度的疾病模型开辟了新路径。
从更宏观的视角来看,肾脏类器官的战略价值已获得国际广泛认可。欧盟HCA|Organoid项目与美国国立卫生研究院标准化类器官建模中心正着力推进方法学的统一与规范,FDA现代化法案2.0亦从政策层面明确支持其作为动物模型的替代方案。随着技术不断迭代,肾脏类器官有望成为解析足细胞病机制、加速靶向疗法开发的核心工具,但从平台成熟走向临床转化,仍需经历严格的验证过程。
总 结
肾脏类器官已在药物毒性评估、遗传性肾病机制解析、复杂疾病建模及自身免疫研究等多个层面展现出独特价值,正逐步从概念验证走向转化应用。随着血管化、成熟度及功能整合性等关键瓶颈被逐一突破,加之国际标准化项目的推进和政策层面的支持,肾脏类器官有望在未来成为连接基础研究与临床精准医学的核心桥梁,为足细胞病的靶向治疗开辟新的路径。
参考文献:
de Cos M, Zeghal M, Meliambro K. Modeling Podocytopathies with Kidney Organoids. Nephrol Dial Transplant. 2026 Apr 21:gfag091
END
关于我们
深圳华津生物科技有限公司致力于提升肾脏疾病研究和肾药开发创新水平。公司与全球最顶尖的干细胞再生医学领域科学家合作,聚焦iPSC来源肾脏细胞、肾类器官、肾类器官芯片及高仿生模拟物平台的打造,为肾脏疾病科研和肾脏药物发现提供最前沿的技术服务和整体解决方案。
公司目前已经建立了Kidnioid®技术平台,其中包括iPSC细胞库、肾类器官及心脏类器官分化技术平台、肾脏类器官疾病模型库,肾类器官芯片平台、动物肾脏疾病模型、分子/蛋白平台、病理平台以及肾类器官培养、分化试剂盒产品等一整套为肾脏疾病研究和肾药开发的完善体系。目前已经为国内众多临床肾病科、大学、科研院所及药企提供产品和技术服务,赢得了客户的信任,已经成为国内肾脏类器官技术平台领头羊。
华津肾类器官平台解决方案
Our Solutions
高度模拟人体生理环境:能够高度贴近人体的真实生理反应,更准确的药物毒性预测。
显著降本增效:大幅削减了药物开发的实验成本与研发周期。
契合伦理趋势:顺应全球减少动物实验的大势。
(点击查看大图)
联系我们
Contact Us
欢迎与我们联系,了解更多肾类器官在药物研发与肾毒性测试中的应用!
地址
广东省深圳市龙华区观澜银星智界2期A座11楼
官方网址
www.nephromedicine.com
联系电话
商务支持
17276665437(李先生)
技术支持
18843183823(田先生)
入群/投稿
13686113433(谭女士)
商务客服1
商务客服2
技术客服
添加客服小助理,了解更多内容
扫码关注华津生物
的官方微信
求点赞
求分享
求喜欢
2026-04-27
Neuropharmacology and Therapy
ACADEMIC LECTURE
NPT锐潮新声
前沿对话系列讲座
2026 年 4 月 19 日,由Neuropharmacology and Therapy(NPT)期刊、逻辑神经科学公众号联合主办的 NPT 锐潮新声·前沿对话学术讲座第11期顺利举办。本次讲座以阿片类药物耐受机制与疼痛管理为核心议题,邀请刘通教授、张俊明教授等领域权威学者参与研讨,线上累计6000 余名同行参会,围绕阿片耐受与阿片诱导性痛觉过敏(OIH)的核心机制展开解析,探讨基础研究向临床镇痛转化的创新路径,为慢性疼痛精准诊疗、阿片类药物安全应用提供新的理论与实践依据。
主讲嘉宾简介
刘通教授
张俊明教授、医学博士
本次近两小时学术交流,从分子机制、临床痛点、药物研发延伸至青年学者科研成长,逐层解析阿片类药物长期应用所致耐受与痛敏的核心问题,搭建基础研究与临床实践深度融合的学术交流平台。
一、主题报告:阿片耐受与痛敏的机制解析 —— 星形胶质细胞与外泌体的核心调控作用
阿片类药物为临床镇痛一线用药,长期应用易诱发药物耐受和阿片诱导性痛觉过敏,是慢性疼痛治疗与临床安全用药的关键瓶颈。刘通教授以科学问题为导向,从星形胶质细胞、外泌体两个研究维度,报告阿片类药物耐受与痛敏的突破性研究成果,为临床镇痛干预提供新靶点与理论支撑。
1. 星形胶质细胞内质网应激通路:介导吗啡耐受与痛敏的关键通路
长期吗啡暴露可特异性激活脊髓星形胶质细胞内质网应激信号,通过PERK/ATF4 通路上调脂质运载蛋白 2(LCN2)与 NLRP3 炎症小体的表达。
• 下游效应:LCN2 结合黑皮质素 4 受体(MC4R),增强背根神经节(DRG)与脊髓神经元兴奋性;NLRP3 炎症小体激活加剧神经炎症反应,二者共同参与吗啡耐受与痛敏的发生发展。
• 干预验证:在糖尿病性疼痛和坐骨神经分支选择性损伤(SNI)慢性疼痛模型中,内质网应激抑制剂(4-PBA、TUDCA)或 NLRP3 抑制剂(β-hydroxybutyrate)处理,可显著提升吗啡的镇痛效应。
2. 外泌体介导的神经元 - 胶质细胞通讯:跨细胞神经炎症传导新途径
慢性吗啡处理导致神经元ADAR1 表达下调,引发双链 RNA(dsRNA)异常蓄积并通过外泌体释放至脑脊液;外泌体携带的 dsRNA 被小胶质细胞内体 TLR3 受体识别,激活 TRIF 信号通路并促进 IL-6 表达,诱发神经炎症。
• 干预验证:抑制外泌体释放(GW4869)或阻断 TLR3 信号通路,可有效恢复吗啡镇痛作用,缓解痛敏症状。
二、临床对话:镇痛临床困境与基础 - 临床转化路径
张俊明教授结合临床实践,聚焦阿片类药物镇痛效能与不良反应并存、耐受与痛敏鉴别困难、临床管理精准度不足等核心问题,与刘通教授围绕疼痛流行病学、临床管理策略、创新药物研发展开对话,明确基础研究成果的临床转化方向。
1. 中国疼痛临床现状与核心挑战
• 疾病负担:中国慢性疼痛患病人数约 3 亿,发病与人口老龄化、骨关节退行性病变密切相关。
• 临床矛盾:阿片类药物严格管控下,镇痛不足与药物滥用风险的平衡成为临床难点。
• 核心鉴别:阿片耐受需增加剂量以维持镇痛效果,阿片诱导性痛敏需降低剂量缓解疼痛,二者临床处置策略相反,精准鉴别是临床诊疗的关键。
2. 创新药物研发与多模式镇痛策略
• 非阿片类镇痛靶点:聚焦 Nav1.7/1.8 通道、α2A 肾上腺素能受体、Mrgpr 受体,研发无成瘾性镇痛药物。
• 阿片类药物优化:开发 μ 阿片受体激动与 NMDA 受体拮抗双功能分子,提升镇痛效能、降低耐受发生风险。
• 多模式镇痛:联合神经调控、物理治疗等手段,减少阿片类药物用量,实现安全镇痛。
张俊明教授指出,吗啡仍是急性疼痛、癌痛镇痛的标准用药,但其成瘾、耐受、痛敏等不良反应与公共卫生风险限制了临床应用;刘通团队的研究阐明了相关不良反应的核心机制,为保留阿片镇痛优势、降低用药风险提供了转化依据。
三、研讨问答:临床与科研核心问题解析
1. 科研选题:聚焦临床需求与机制创新
刘通教授分享研究思路:以阿片类药物临床应用难题、国家战略需求为选题导向,围绕神经炎症与疼痛敏化核心机制,通过无偏组学筛选发现新靶点,再开展机制验证;外泌体 - dsRNA-TLR3 通路研究为反向筛选、正向验证的典型案例。张俊明教授补充,选题应锚定未解决的临床关键问题,契合科研资助方向,同时强化系统科研训练,形成独立研究思维。
2. 核心概念鉴别:阿片耐受与阿片诱导性痛敏
• 阿片耐受:药理学现象,需增加药物剂量维持镇痛效果,不直接诱发疼痛。
• 阿片诱导性痛敏:临床疼痛症状,需降低药物剂量方可缓解,二者机制部分重叠但通路偏好存在差异,精准鉴别对临床处置至关重要。
3. 镇痛药物研发方向:非阿片靶点与阿片优化并行
非阿片类药物聚焦 Nav1.7/1.8、α2A 受体、Mrgpr 受体等靶点;阿片类药物以双功能分子为优化方向;同时采用多模式镇痛方案,降低阿片类药物依赖。
4. 慢性疼痛流行病学:中国 3 亿患者的成因
核心诱因为人口老龄化相关的退行性疾病高发;社会心理因素如慢性应激、生活压力亦推动疼痛发生;阿片类药物严格管控有效降低滥用风险,领域仍需完善流行病学数据体系。
5. 机制与转化答疑
• dsRNA:以结构与长度依赖性方式激活 TLR3,序列特异性较低,呈蓄积性激活特征。
• 吗啡 - 胶质细胞受体互作:可通过单细胞测序、药理学阻断、条件性基因敲除等技术验证。
• 外泌体:来源决定生物学功能,干细胞、年轻来源外泌体具备镇痛潜力,临床转化前景明确。
6. 期刊投稿方向(NPT)
NPT 期刊编委张俊明教授指出,期刊优先接收机制创新、具备临床转化价值的研究,涵盖阿片类药物优化、非阿片镇痛新药、胶质细胞 - 神经元互作等方向;支持神经免疫、神经血管、神经肿瘤等交叉学科研究,以及 AI、类器官、人源组织等新技术应用研究;欢迎疼痛、成瘾神经机制、神经退行性疾病等领域的高质量成果投稿。
四、青年学者科研指引:选题与成长路径
本次讲座结合领域前沿与实践经验,为青年学者提供科研选题、研究设计、成果转化的实操性建议,助力疼痛医学与神经药理学领域青年研究者突破发展瓶颈。
本期总结
本次 NPT 锐潮新声讲座以阿片类药物耐受与疼痛管理为核心,覆盖分子机制、临床转化、药物研发、青年科研成长全链条,明确星形胶质细胞内质网应激、外泌体介导的细胞通讯在阿片耐受与痛敏中的调控作用,确立非阿片靶点研发、多模式镇痛的临床发展方向。
本次交流反映了领域对阿片类药物安全应用、慢性疼痛精准诊疗的迫切需求。期待领域内研究者持续聚焦该方向,深化机制研究、推动协同创新,加速创新靶点与药物的临床转化,为中国慢性疼痛管理、阿片类药物合理应用提供科学方案,推动神经药理学与疼痛医学领域的持续发展。
关于期刊
Neuropharmacology and Therapy(简称NPT、神经药理学与治疗、ISSN:2990-8779)是一本开放获取(Open Access)国际学术期刊。该期刊以神经药理学方法阐明重要神经系统疾病的发病机制,推动领域内转化研究与学术交流,重点关注神经疾病药物作用机理及神经系统功能障碍的治疗相关研究。NPT定位 JCR 一区,预期影响因子 ≥ 10。
期刊荣誉主编为中国科学院院士、复旦大学脑科学研究院杨雄里教授;主编为国家杰出青年科学基金获得者、河北大学基础医学院吉永华教授,及国家高层次人才计划领军人才、南昌大学药学院张汉霆教授。编委团队由多本SCI期刊主编/副主编,及全球神经药理学领域的杰出学者组成,为期刊学术质量把控与国际影响力建设提供坚实支撑。
期刊重点关注的研究方向(包括但不限于):
• 神经疾病新型治疗策略的研发与临床转化
• 神经系统疾病治疗靶点的结构解析与功能验证
• 神经疾病潜在药物分子(含天然产物、小分子化合物)的发现、设计与合成
• 神经药理学领域的多组学研究与应用
• 神经药物相关生物数据库构建与分子设计软件开发
• 具神经疾病治疗潜力新分子的生物活性及作用模式研究
• 神经系统疾病治疗的临床研究、临床试验与荟萃分析
• 神经药理学研发的管理、商业转化与监管研究
• 神经药理学领域的其他前沿研究方向
期刊接收的稿件类型:
• 研究论文(Research Articles)
• 综述文章 (Review Articles)
• 社论 (Editorials)
• 读者来信 (Letters)
• 研究亮点 (Research Highlights)
• 前沿视角 (Perspectives)
• 病例报告 (Brief Cases Report)
• 研究方法 (Study Methods)
选择NPT,赋能神经药理学研究国际传播
1. 开放获取模式:全球读者可通过期刊官网、ScienceOpen平台免费直接下载论文,有效拓展研究成果的传播范围,提升学术影响力。
2. 无版面限制:作为纯电子期刊,突破传统出版的版面约束,对论文数量与篇幅均不设限,保障研究成果的完整呈现,避免因内容表达需求导致的稿件退审。
3. 零费用发表:不向作者收取投稿费、文章处理费(APC)等任何出版相关费用。
4. 版权归属作者:作者可完整保留所发表文章的全部版权。
5. 高效同行评审:采用优化的快速同行评审流程,大幅缩短稿件评审周期,提升发表效率。
6. 即时在线发布:稿件录用后即刻在线首发,加速研究成果的传播与学术转化。
7. 免费学术支持:为作者提供免费的语言润色、查重协助等学术服务,助力稿件质量提升。
8. 全球化成果推广:通过多渠道开展论文精准推广,包括向领域内临床医师与研究者定向发送邮件、在微信及X(Twitter)向期刊粉丝推送、发布新闻稿进行全球媒体宣传等。
9. 国际化投审稿系统:依托ScholarOne在线投审稿平台(https://mc04.manuscriptcentral.com/nptherapy),实现稿件注册、提交、评审全流程线上化操作,便捷高效。
期刊官网
如需了解NPT更多详情,包括稿件提交指南、免费内容提醒服务注册流程等,可访问期刊官方网站:https://npt-journal.org/,获取最新的期刊动态与标准化操作指引。
国际数据库收录情况
NPT已被CAS、Google Scholar、WorldCat、Primo Central、Sherpa Romeo、Summon等国际知名学术数据库收录。稿件一经上线,即可在Google Scholar实现检索,保障已发表论文被全球科研机构与研究者高效获取,提升成果的学术可见度与传播效率。后续数据库收录进展,将通过期刊官网实时更新。
编辑部联系方式
1. 稿件咨询:请致编辑部邮箱editorial@npt-journal.org
2. X(Twitter):@NTherapy22372
3. 微信公众号:欢迎关注NPT期刊官方微信公众号,及时获取期刊资讯~
NPT期刊服务号
NPT学术订阅号
版权声明:
本文由Neuropharmacology and Therapy(NPT)期刊编辑部编撰。如需转载请通过公众号后台留言。
我们期待与学界同仁携手,共同维护学术内容的规范传播与知识共享,助力神经药理学领域的高质量学术交流与发展。欢迎关注我们,及时获取最新学术动态与出版资讯。
☞欢迎【分享,点赞,转发】
100 项与 4-苯基丁酸酯 相关的药物交易
登录后查看更多信息
外链
| KEGG | Wiki | ATC | Drug Bank |
|---|---|---|---|
| - | - | - |
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 肝内胆汁淤积 | 临床3期 | - | 2023-02-08 | |
| 遗传病 | 临床3期 | - | 2023-02-08 | |
| α1-抗胰蛋白酶缺乏症 | 临床2期 | 美国 | 2001-11-01 | |
| 阿尔茨海默症 | 临床2期 | - | - | |
| 阿尔茨海默症 | 临床2期 | - | - | |
| 包涵体肌炎 | 临床1期 | 美国 | 2020-08-20 | |
| 炎症 | 临床前 | 德国 | 2025-07-07 | |
| 急性肺损伤 | 临床前 | 中国 | 2025-05-16 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
临床2期 | - | 醖憲顧鑰鹽網鬱艱蓋製(鹽廠願選衊構構網繭範): P-Value = 0.01 更多 | - | 2018-09-01 | |||
Placebo |
登录后查看更多信息
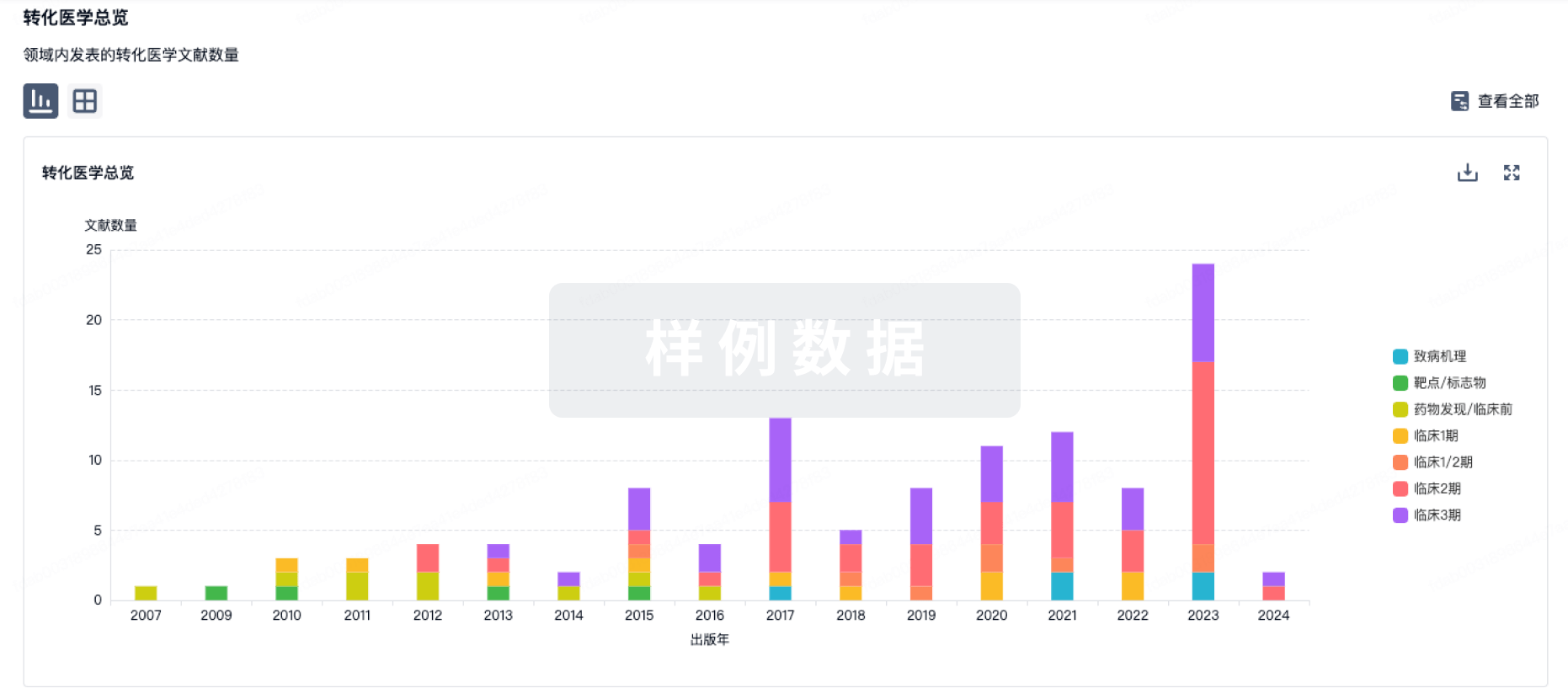
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
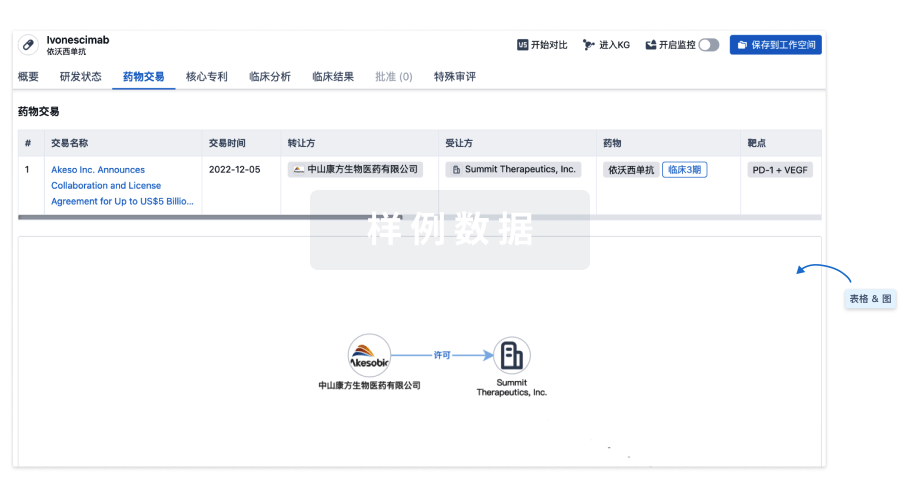
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
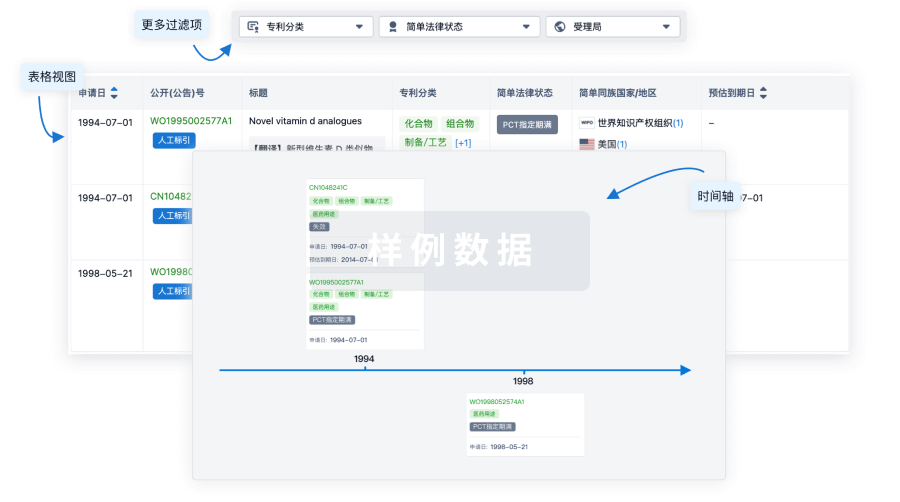
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
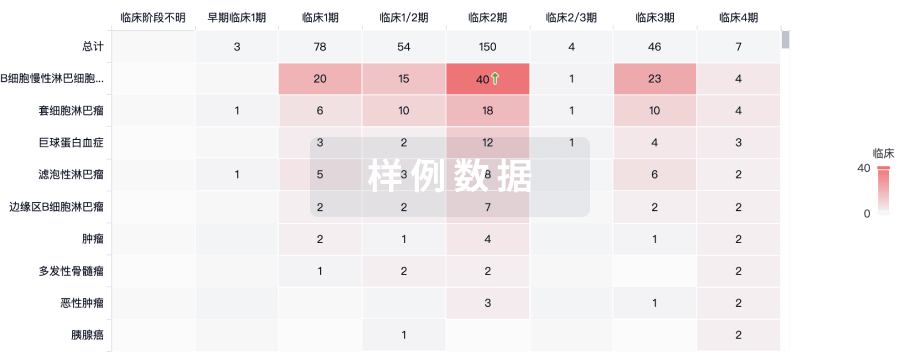
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
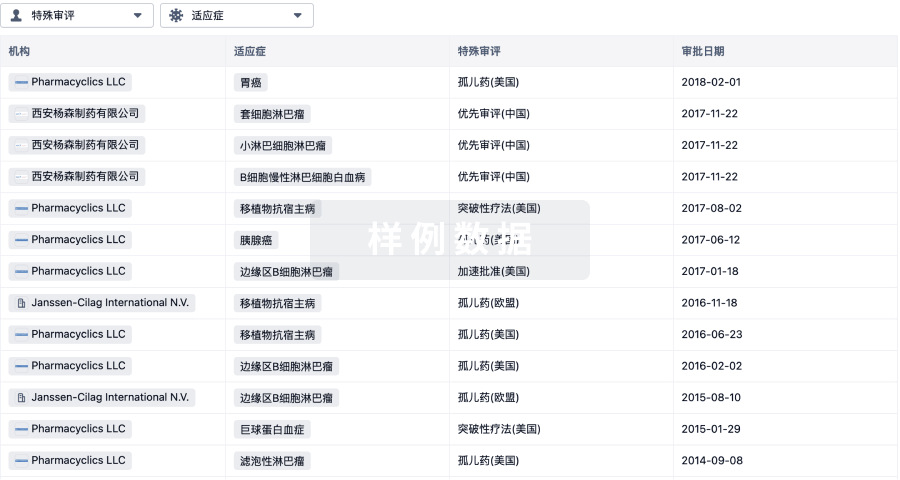
批准
利用最新的监管批准信息加速您的研究。
登录
或
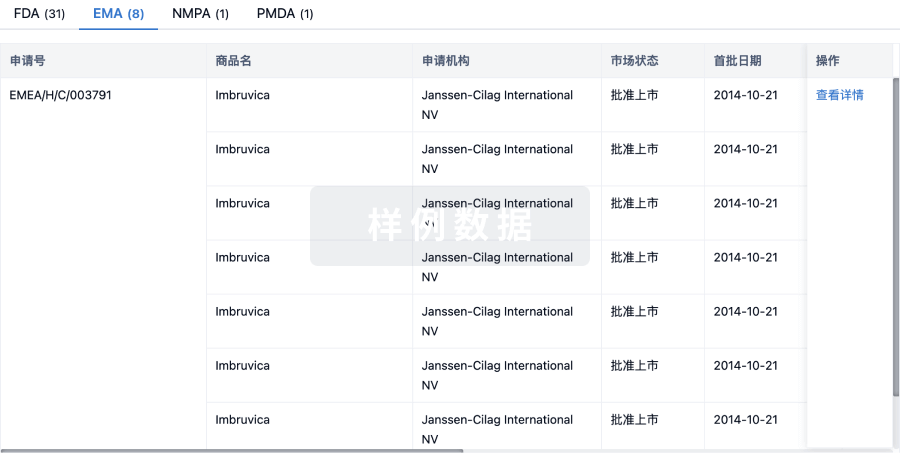
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用







