预约演示
更新于:2026-03-18

Istituto Italiano di Tecnologia
更新于:2026-03-18
概览
标签
肿瘤
其他疾病
神经系统疾病
小分子化药
化学药
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 小分子化药 | 28 |
| 化学药 | 1 |
关联
29
项与 Istituto Italiano di Tecnologia 相关的药物靶点 |
作用机制 TYMS抑制剂 |
原研机构- |
在研适应症 |
最高研发阶段临床前 |
首次获批国家/地区 中国 |
首次获批日期1997-01-01 |
作用机制 COX-1抑制剂 [+2] |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区 美国 |
首次获批日期1987-12-31 |
靶点 |
作用机制 CDC42 抑制剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
16
项与 Istituto Italiano di Tecnologia 相关的临床试验NCT06928857
Pilot Study on the Effects Induced by an Electromyographic-controlled Functional Electrical Stimulator (FitFES) for Upper Limb Rehabilitation in Post-stroke Patients
Upper limb disabilities are among the most debilitating issues after a cerebral stroke. One promising approach in motor rehabilitation is the use of functional electrical stimulation (FES). This technique can be integrated into daily therapy to follow an adaptive approach, exploiting the residual capacities of patients. FES can help to stimulate the affected muscles, improve coordination and strengthen the weakened muscles, thus supporting the rehabilitation process.
开始日期2025-03-03 |
申办/合作机构 |
NCT07235111
Omics Sciences for the Identification of Pathogenetic Mechanisms and Biomarkers in Neurodegenerative Diseases
The study aims to use 'omics' sciences, employing the most advanced technologies currently available, in order to identify pathogenic genomic variants, proteins and/or altered molecular pathways in neurodegenerative diseases and to obtain a new and more complete characterisation of subjects affected by the neurodegenerative diseases under study. Thanks to the integration of genomic, gene expression (transcriptomic and epigenomic), protein and metabolic data and clinical data, the study also aims to identify new markers for the diagnosis, prognosis, also in terms of response to therapy, and monitoring of neurodegenerative diseases.
The study involves the enrolment of at least 1.200 individuals with neurodegenerative disease.
The study involves the enrolment of at least 1.200 individuals with neurodegenerative disease.
开始日期2025-02-28 |
申办/合作机构 |
NCT06763874
Pilot Study of Usability and Acceptability for the Optimization of a Functional Electrical Stimulator Controlled by Electromyographic Signal for the Rehabilitation of the Upper Limb in Persons Post-stroke
Functional recovery of the upper limb after a cerebral stroke is one of the major critical issues in rehabilitation. The advent of innovative technologies can be helpful to rehabilitators and intervene where there is no other solution but a clear therapeutic indication. The use of functional electrostimulators can actively and functionally support movements, helping people affected by stroke to complete a motor gesture taking into account their residual capacities.
开始日期2025-01-14 |
申办/合作机构 |
100 项与 Istituto Italiano di Tecnologia 相关的临床结果
登录后查看更多信息
0 项与 Istituto Italiano di Tecnologia 相关的专利(医药)
登录后查看更多信息
6,322
项与 Istituto Italiano di Tecnologia 相关的文献(医药)2026-06-01·COGNITION
Emotional egocentricity bias is modulated by implicit expectations of interpersonal emotional contingencies and perceptual noise
Article
作者: Bolis, Dimitris ; Kotsaris, Vassilis ; Azevedo, Ruben
Emotional Egocentricity Bias (EEB) refers to the tendency to project one's own emotional state onto others. While previous research has demonstrated EEB in multiple paradigms, its underlying mechanisms remain unclear. Across two studies, we used a novel dual-task paradigm to examine how fluctuations in expected interpersonal emotional contingencies (IEC) and perceptual ambiguity shape EEB. In each trial, participants underwent an implicit emotion induction through a roulette game and subsequently categorized an ambiguous facial expression. Experimental blocks varied in the probability of emotion congruency between self and other. Behavioural results showed that implicit congruency expectations modulated EEB as accuracy was highest for congruent trials in neutral and congruent blocks but reversed in incongruent blocks, indicating implicit adaptation to IEC. Interestingly, higher perceptual noise improved performance and amplified contextual effects, suggesting that EEB is jointly shaped by interpersonal predictive processing and sensory noise. To model individual learning of IEC, we employed a Hierarchical Gaussian Filters (HGF) computational model, revealing that participants updated their beliefs about IEC in a volatility-sensitive manner and that decisions were primarily based on posterior beliefs. Heart rate acceleration following outcomes was linked to belief updates, suggesting that arousal influences socio-emotional learning. These findings show that EEB reflects context-sensitive inferences shaped by internal states and perceived uncertainty and highlight the role of interoception in adaptive emotion recognition. This work adds to our understanding of EEB offering insights for future studies on embodied emotion perception in dynamic social contexts.
2026-03-04·NANO LETTERS
Plasmonic Nanocavity-Assisted Long-Range Dipole–Dipole Interactions for Rare-Earth Ions
Article
作者: Kang, Ke ; Chen, Huan ; Zhang, Zhenglong ; Xie, Xin ; Gou, Chengxiang ; Li, Wangze ; Zhang, Min ; Kang, Bowen ; Guo, Lei ; Hu, Huatian
Trivalent rare-earth ions (RE3+) exhibit sharp 4f-4f transitions that are highly sensitive to dipole-dipole interactions (DDIs), yet the intrinsic short-range of DDIs restricts their application in long-distance quantum coupling. Here, we demonstrate a plasmonic nanocavity pair that simultaneously couples to upconversion nanoparticles (UCNPs) and quantum dots (QDs), enabling enhanced emission and inter-emitter interaction. The combined effects of Purcell enhancement and surface plasmon polariton (SPP) propagation extend the DDI-mediated energy transfer over a record distance exceeding 7.5 μm, 2 orders of magnitude longer than conventional Förster resonance systems. Moreover, the polarization-controlled SPP propagation directs the in-plane energy redistribution, achieving anisotropic and tunable coupling. This work establishes a viable route toward controllable, long-range DDIs among rare-earth emitters, advancing integrated quantum photonic architectures.
2026-03-01·NEUROPSYCHOLOGIA
Investigating the role of sensorimotor versus contextual cues in the sense of joint agency: a human-human and human-robot study
Article
作者: De Tommaso, Davide ; Navare, Uma Prashant ; Wykowska, Agnieszka ; Kompatsiari, Kyveli ; Ciardo, Francesca ; Hobbelink, Veerle
Sense of Joint Agency (SoJA), is the feeling of control experienced by humans for their own, as well as their partner's actions, when acting in joint action with others. SoJA is ubiquitous in human-human interaction. Therefore, it is both interesting as well as relevant to understand the factors that affect the formation of SoJA both in human-human and in human-robot interaction. On the one hand, previous work suggests that sensorimotor cues may be the main determinant of "lower-level" implicit SoJA in human-human joint action. On the other hand, recent work shows that contextual factors, such as perceived intentionality, can impact the formation of implicit SoJA with a humanoid robot. In the current study, we aimed to investigate, using behavioral and electroencephalography (EEG) measures, whether endowing a humanoid robot with a precisely human sensorimotor pattern would be sufficient to elicit SoJA with the robot, even without manipulating the perceived intentionality of the robot. Participants completed a joint task with another human, and with a robot that was controlled by another human, thus endowing the robot with a precisely human sensorimotor repertoire. Importantly, participants completed two sessions with this controlled robot. In one session, they were (factually) told that the robot was controlled by another human. In another session, they were told that the robot was pre-programmed. We expected that participants may attribute less intentionality to the robot they believed was pre-programmed. In either session, participants' perception regarding the robot's intentionality was not explicitly manipulated. Interval estimates for self- and other-generated actions were used to estimate SoJA. In addition, we also measured participants' readiness potential (RP) for self- and partner actions, and their N100 responses for self- and partner-generated sensory outcomes in the task. The results show that temporal estimates, and ERPs, did not differ between self- and partner-generated action-outcome conditions, in both human-human and human-robot sessions. Thus, these results suggest that endowing a humanoid robot with a human sensorimotor pattern may be sufficient to elicit SoJA in joint action with the robot, regardless of the intentionality attributed to the robot. Furthermore, an exploratory spectral analysis of movement-related beta activity suggested that people may nevertheless disengage earlier from the joint action when interacting with a robot partner, as compared to a human partner. Together, our results contribute to understanding the mechanisms that underlie the emergence of SoJA in joint action, as well as the extent to which artificial agents, such as humanoid robots, can be integrated into our teamwork as "full" interaction partners.
2
项与 Istituto Italiano di Tecnologia 相关的新闻(医药)2021-11-12
CrestOptics manufactures high-end microscopy instruments under its own brand and as an original equipment manufacturer (OEM)
CrestOptics is a manufacturer of high-end microscopy solutions and advanced systems for fluorescence microscopy and diagnostic applications. (Credit: Konstantin Kolosov from Pixabay)
CrestOptics S.p.A., a manufacturer of high-end microscopy solutions and advanced systems for fluorescence microscopy and diagnostic applications, today announced it has been acquired through a majority shareholding by Apposite Capital LLP, the healthcare specialist private equity investor.
The investment will support CrestOptics’ international commercial expansion and boost product development and innovation, establishing the company as an attractive commercial partner and employer in the life science industry.
CrestOptics represents the fourth investment of Apposite’s new fund, Apposite Healthcare III. Apposite forms a close partnership with its companies to provide them with the strategic, financial and human resources required to accelerate growth through expansion into new product lines and geographies, both organically and through synergistic acquisitions.
As part of the agreement, Dr David Martyr will join the CrestOptics’ Board of Directors as Chair. Previously Group President of Leica Microsystems, and CEO of Tecan, David brings extensive industry experience. He will be focused on supporting the rapid expansion of the Company.
CrestOptics manufactures high-end microscopy instruments under its own brand and as an original equipment manufacturer (OEM). The Company’s instrumentation can be integrated with other high-end imaging systems used in life science research to enhance detection and imaging performance.
“This agreement demonstrates that CrestOptics is an established brand and a strong player in the worldwide microscopy market. Strategic collaborations with external partners such as Apposite Capital LLP will help us further enforce our market positioning and strengthen our product portfolio with other innovative technologies. Apposite and CrestOptics share the same vision for the future and see huge potential for growth” said Renato Giacobbo Scavo, CEO of CrestOptics.
“CrestOptics has already started an ambitious process of internationalization and this new partnership will boost our efforts to enable us to reach a wider and more diversified market at the global level,” said Alessandra Scarpellini, PhD, Head of Sales and Marketing at CrestOptics.
“CrestOptics, a leader in the development and manufacture of advanced imaging systems for life science research, is exactly the type of differentiated company Apposite seeks to invest in and we are excited to support rapid growth by accelerating further innovation and product development, and facilitate geographical expansion into new markets by leveraging our industry knowledge and contacts,” said David Porter, Founding Partner of Apposite Capital LLP. “This partnership will cement CrestOptics’ position as a pioneering and reliable imaging innovator and provider on a global scale.”
“CrestOptics has already demonstrated that it is a highly capable and innovative group, offering very high performance spinning disc imaging systems at affordable prices. With an exciting pipeline of technology and products in development, including expanded super-resolution capabilities, I am delighted to be joining the CrestOptics team.” said Dr David Martyr, Chair of Board of Directors of CrestOptics.
The agreement is supported by existing shareholder, Xyence, an Italian life sciences investor and partner of CrestOptics since 2018, which supports the early-stage diagnostics research programmes the Company is conducting with Istituto Italiano di Tecnologia (IIT). These research programmes in the fields of cancer, Alzheimer’s disease and point of care virus detection will be carried on by the spin off D-TAILS, led by Vincenzo Ricco, founder of CrestOptics, who will also continue to bring value to the Company in a non-executive capacity.
Financial terms of the acquisition were not disclosed.
Advisers to CrestOptics and Xyence: Chiomenti, Leading Law, Madlex (legal), AKRAN Intellectual Property (IP).
Advisers to Apposite Capital: Deloitte (financial advisory, accounting and tax), McDermott Will & Emery (legal), PMSI (commercial), Peter Call (specialist microscopy consultant), Botti & Ferrari (IP), Lockton (insurance)
Source: Company Press Release
并购合作
2012-10-30
Novartis Venture Funds, along with Novo Ventures, led a $16 million funding round of a San Diego pharmaceutical startup Thesan Pharmaceuticals.
The series A financing will be used to develop new approaches in treating inflammatory skin disorders.
“For many years, drug development in the dermatology field has been dominated by old drugs just reformulated into more cosmetically appealing foams, gels or lotions,” said Gordon Foulkes, Thesan’s executive chairman and CEO, in a news release. “Thesan’s mission is to develop new chemical entities with novel mechanisms of action in order to substantially improve treatment outcomes for patients with a variety of dermatological diseases.”
Thesan, based in San Diego, has licensed the technology from the University of California, Irvine and the products have been developed in the laboratory of professor Daniele Piomelli, Louise Turner Arnold Chair of Neurosciences at UCI and the director of Drug Discovery and Development at the Istituto Italiano di Tecnologia (IIT) in Italy.
“A novel mechanism of action with dramatic efficacy, coupled with such solid chemistry is a great starting point for the formation of a new company,” said Giovanni Ferrara, venture partner at Novartis Venture Funds, in Thesan’s news release. “Combined with a world-class team, it made for a compelling investment.”
100 项与 Istituto Italiano di Tecnologia 相关的药物交易
登录后查看更多信息
100 项与 Istituto Italiano di Tecnologia 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年06月09日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
1
28
临床前
其他
1
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
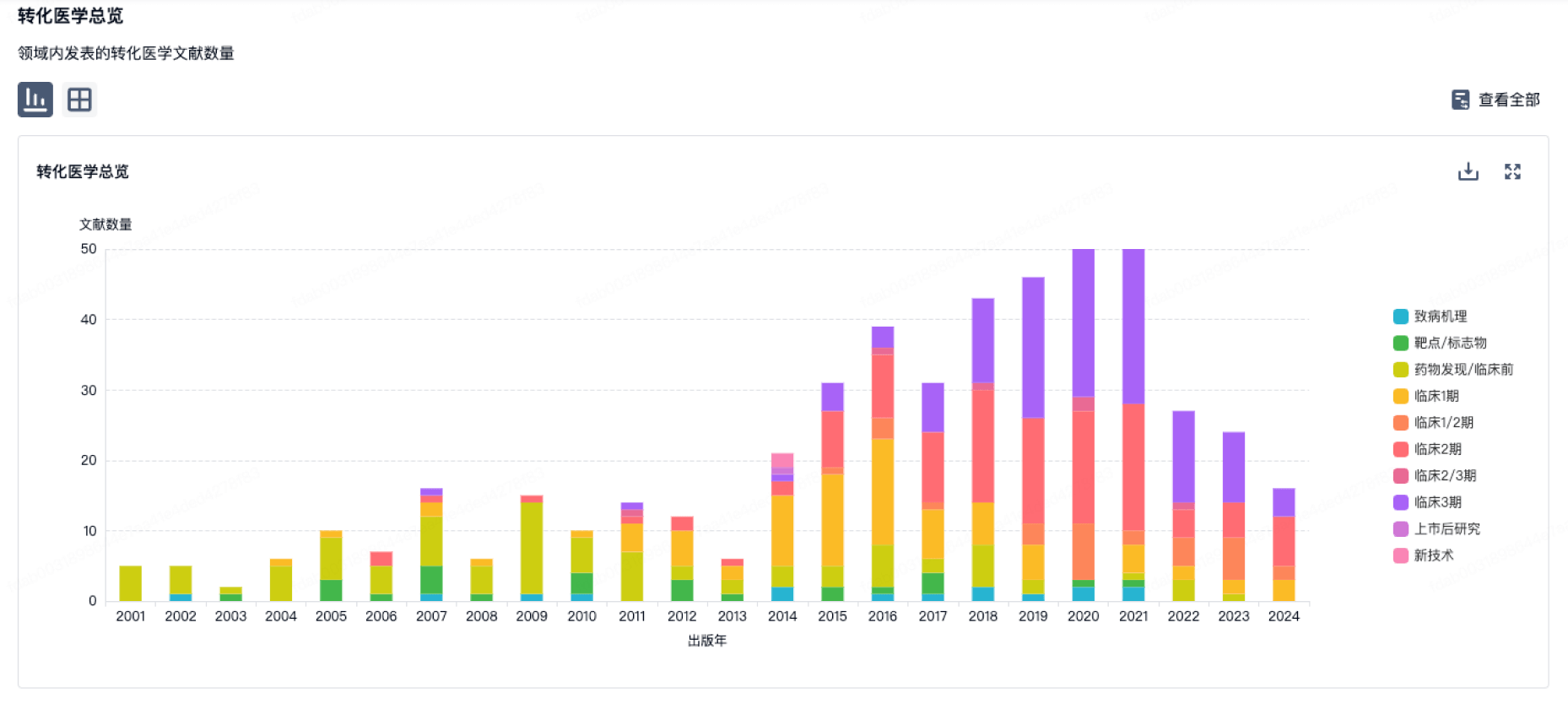
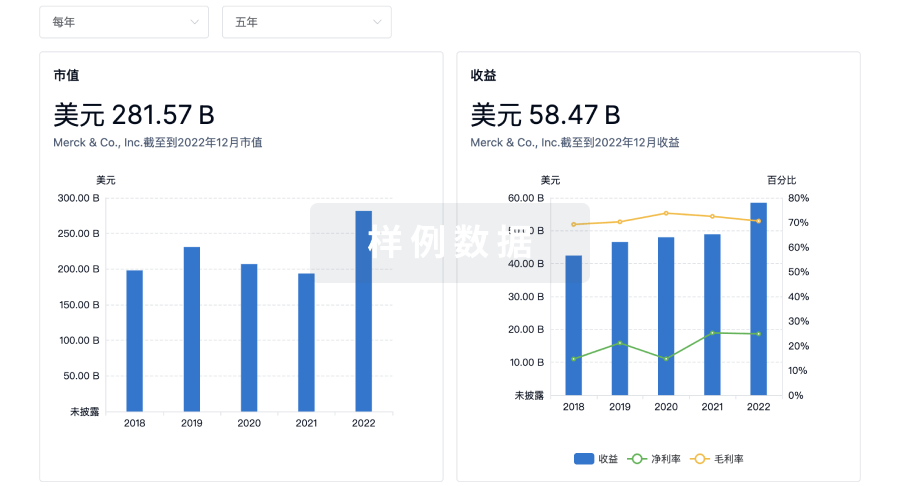
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
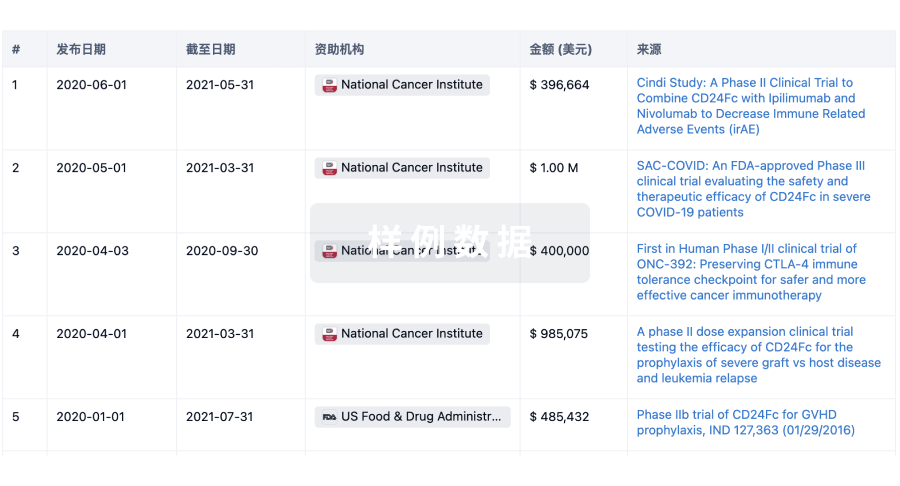
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
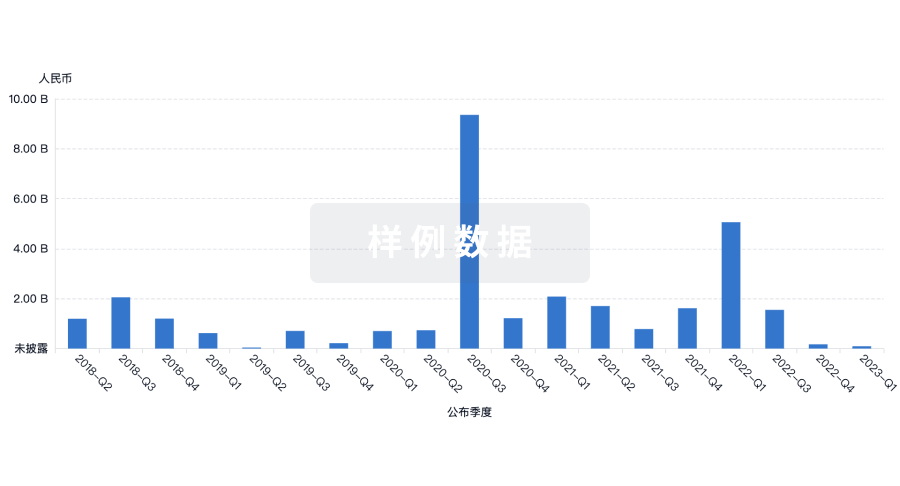
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
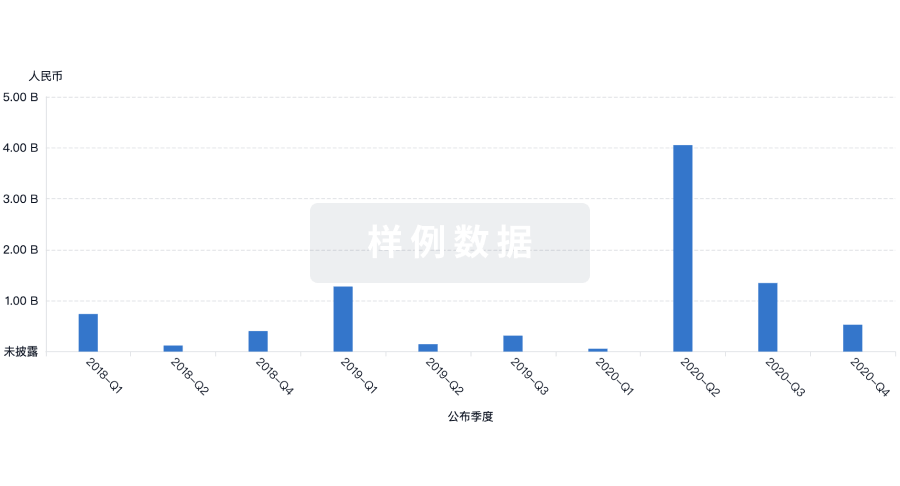
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
