预约演示
更新于:2026-05-12

Regulus Therapeutics, Inc.
更新于:2026-05-12
概览
标签
消化系统疾病
其他疾病
神经系统疾病
ASO
小分子化药
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| ASO | 3 |
| 小分子化药 | 1 |
关联
4
项与 Regulus Therapeutics, Inc. 相关的药物靶点 |
作用机制 miR-17抑制剂 [+2] |
在研适应症 |
非在研适应症- |
最高研发阶段临床1期 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 miR-155拮抗剂 [+1] |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 miR-132抑制剂 [+2] |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
7
项与 Regulus Therapeutics, Inc. 相关的临床试验NCT05521191
A Phase 1b, Double-Blind, Placebo-Controlled, Multiple Ascending Dose and an Open-Label Fixed-Dose Study in Patients With Autosomal Dominant Polycystic Kidney Disease to Evaluate the Safety, Tolerability, Pharmacodynamics, and Pharmacokinetics of RGLS8429
Primary Objectives
* To assess the safety and tolerability of RGLS8429
* To assess the impact of RGLS8429 on ADPKD biomarkers
Secondary Objectives
* To assess the impact of RGLS8429 on height-adjusted total kidney volume (htTKV)
* To characterize the pharmacokinetic (PK) properties of RGLS8429
* To assess the impact of RGLS8429 on renal function
* To assess the safety and tolerability of RGLS8429
* To assess the impact of RGLS8429 on ADPKD biomarkers
Secondary Objectives
* To assess the impact of RGLS8429 on height-adjusted total kidney volume (htTKV)
* To characterize the pharmacokinetic (PK) properties of RGLS8429
* To assess the impact of RGLS8429 on renal function
开始日期2022-10-06 |
申办/合作机构 |
NCT05429073
A Phase 1, Double-Blind, Placebo-Controlled, Single Ascending Dose Study in Healthy Volunteers to Evaluate the Safety, Tolerability, and Pharmacokinetics of RGLS8429
Primary Objective
• To assess the safety and tolerability of single ascending doses of RGLS8429
Secondary Objectives
* To identify dose-limiting toxicity (DLT) and to determine the maximum tolerated dose (MTD) of a single SC dose of RGLS8429
* To characterize the pharmacokinetic (PK) properties of RGLS8429
• To assess the safety and tolerability of single ascending doses of RGLS8429
Secondary Objectives
* To identify dose-limiting toxicity (DLT) and to determine the maximum tolerated dose (MTD) of a single SC dose of RGLS8429
* To characterize the pharmacokinetic (PK) properties of RGLS8429
开始日期2022-06-10 |
申办/合作机构 |
NCT04536688
A Phase 1b, Multicenter, Open-Label, Adaptive Design Study to Evaluate the Safety, Tolerability, Pharmacokinetics, and Pharmacodynamics of RGLS4326 Administered Via SC Injection to Patients With Autosomal Dominant Polycystic Kidney Disease
Primary Objective
• To assess the dose response relationship between RGLS4326 and ADPKD biomarkers
Secondary Objectives
* To characterize the pharmacokinetic (PK) properties of RGLS4326 in plasma and urine
* To assess the safety and tolerability of RGLS4326
• To assess the dose response relationship between RGLS4326 and ADPKD biomarkers
Secondary Objectives
* To characterize the pharmacokinetic (PK) properties of RGLS4326 in plasma and urine
* To assess the safety and tolerability of RGLS4326
开始日期2020-10-13 |
申办/合作机构 |
100 项与 Regulus Therapeutics, Inc. 相关的临床结果
登录后查看更多信息
0 项与 Regulus Therapeutics, Inc. 相关的专利(医药)
登录后查看更多信息
17
项与 Regulus Therapeutics, Inc. 相关的文献(医药)2017-05-01·Molecular cancer therapeutics2区 · 医学
Lipid Nanoparticle–Mediated Delivery of Anti-miR-17 Family Oligonucleotide Suppresses Hepatocellular Carcinoma Growth
2区 · 医学
Article
作者: Sung, Eric ; Davis, Scott ; Chau, B. Nelson ; Li, Jian ; Kaimal, Vivek ; Magnus, Jill ; Estrella, Heather ; Prudente, Rene ; Lee, Robin ; Yang, Xia ; Sorourian, Mehran ; Walls, Marlena ; Huang, Xinqiang ; Zabludoff, Sonya ; Pavlicek, Adam ; Karmali, Priya ; Lee, Edmund C.
Abstract:
Hepatocellular carcinoma (HCC) is one of the most common human malignancies with poor prognosis and urgent unmet medical need. Aberrant expression of multiple members of the miR-17 family are frequently observed in HCC, and their overexpression promotes tumorigenic properties of HCC cells. However, whether pharmacologic inhibition of the miR-17 family inhibits HCC growth remains unknown. In this study, we validated that the miR-17 family was upregulated in a subset of HCC tumors and cell lines and its inhibition by a tough decoy inhibitor suppressed the growth of Hep3B and HepG2 cells, which overexpress the miR-17 family. Furthermore, inhibition of the miR-17 family led to a global derepression of direct targets of the family in all three HCC cell lines tested. Pathway analysis of the deregulated genes indicated that the genes associated with TGFβ signaling pathway were highly enriched in Hep3B and HepG2 cells. A miR-17 family target gene signature was established and used to identify RL01-17(5), a lipid nanoparticle encapsulating a potent anti-miR-17 family oligonucleotide. To address whether pharmacologic modulation of the miR-17 family can inhibit HCC growth, RL01-17(5) was systemically administrated to orthotopic Hep3B xenografts. Suppression of Hep3B tumor growth in vivo was observed and tumor growth inhibition correlated with induction of miR-17 family target genes. Together, this study provides proof-of-concept for targeting the miR-17 family in HCC therapy. Mol Cancer Ther; 16(5); 905–13. ©2017 AACR.
2017-01-01·Methods in molecular biology (Clifton, N.J.)
Assessing Anti-miR Pharmacology with miRNA Polysome Shift Assay
Article
作者: Androsavich, John R
Target engagement measurements are critical for evaluating developmental drug candidates and their pharmacological activity. microRNA (miRNA) Polysome Shift Assay enables measurement of anti-miR drug target engagement (i.e. extent of miRNA inhibition) without the need to pre-identify or pre-validate downstream miRNA-regulated genes. This makes it useful for assessing anti-miR activity in target tissues or cells where biology of the inhibited miRNAs may not be well understood. In addition, miRNA Polysome Shift Assay can be multiplexed to assess inhibition of multiple miRNAs by a single anti-miR, thus guiding drug optimization for enhancing or avoiding these activities as desired. This chapter outlines the miRNA Polysome Shift Assay technique, describes sample preparation and quality control, and how to calculate and interpret results.
2017-01-01·Methods in molecular biology (Clifton, N.J.)
Competitive Argonaute-Based RNA Immunoprecipitation for Investigation of Transcriptomic Response to Anti-miR
Article
作者: Androsavich, John R
Identification and validation of microRNA (miRNA) target genes is essential for gaining a better understanding of the many different functions miRNAs have in healthy and diseased cells. From a practical standpoint, validated target genes are also useful for monitoring pharmacological activity of developmental therapeutics that modulate miRNAs, such as anti-miRNA oligonucleotides (anti-miR). Here, we describe a method that uses changes in Argonaute 2-RNA immunoprecipitation in response to competition by anti-miR, titrated ex vivo, as physical evidence for target validation.
204
项与 Regulus Therapeutics, Inc. 相关的新闻(医药)2026-04-30
Talking over pharma pipelines with a major drugmaker’s development chief, he noted that if you looked at the biggest 50 drugs by revenue, the final one on the list brought in about $4 billion last year.
Number one, of course, was Keytruda, at $32 billion.
To understand what goes into pivotal trials at the R&D 15 — our in-depth review of the biggest pharma research spenders — you have to recognize that the industry’s bar for success has been steadily rising for years. To get a callout by the CEO in their end-of-year reviews, you need to be in blockbuster territory. Real success takes peak sales projections of $4 billion to $5 billion. Pipelines-in-a-product remain the holy grail of R&D, with the prospect of expanding indications and rising revenue.
In this club, you play with big stakes for even larger pots. As a result, you’re seeing a continued shift toward big population drugs, where remarkable efficacy and reliable safety are essential. That competition for winners that can match the success of the obesity drug franchises has driven a growing demand for deals in China, the US and Europe.
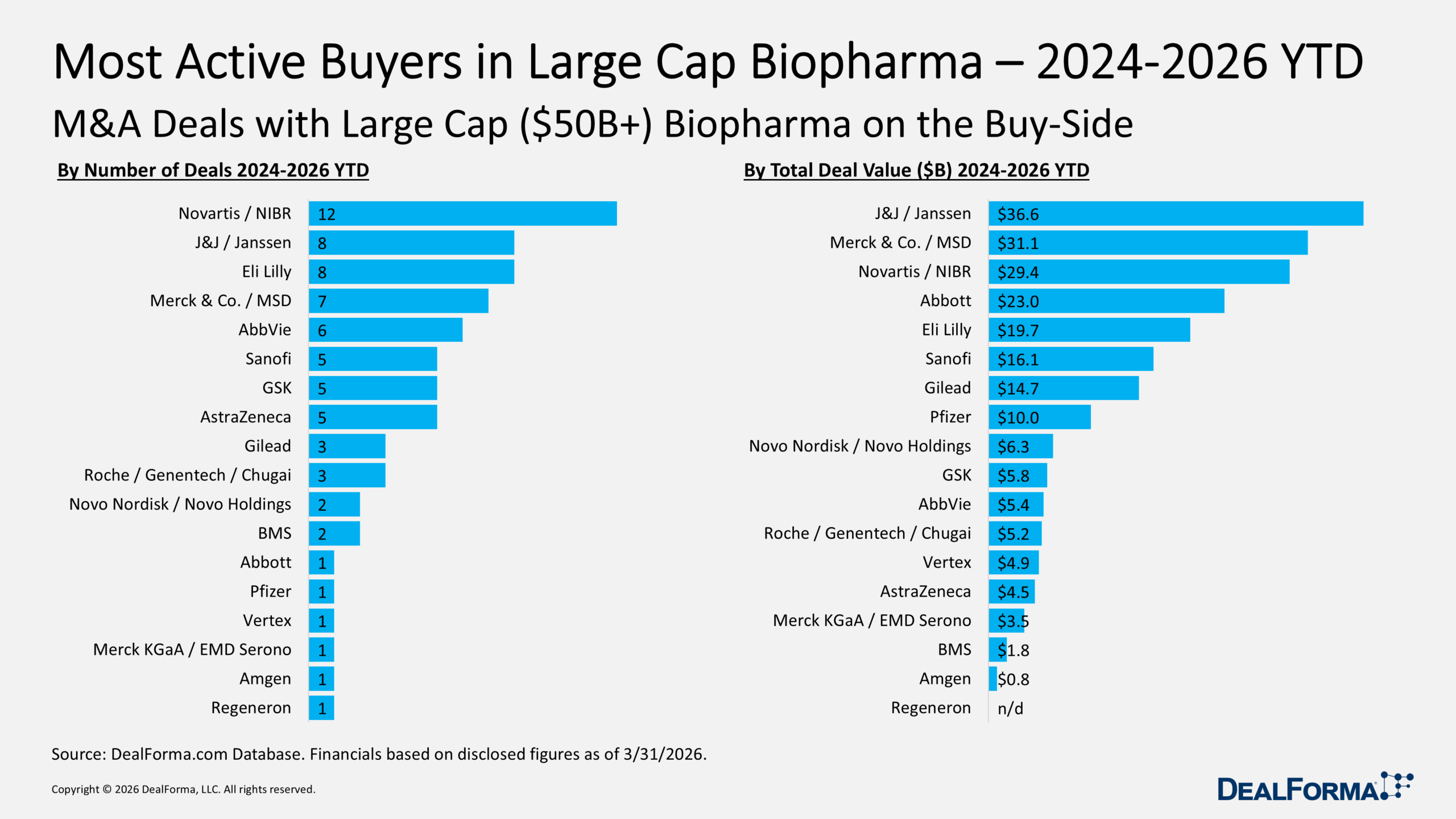
Together, J&J, Merck and Novartis committed close to $100 billion to M&A deals from the start of 2024 through the end of this year’s first quarter. And the tab is growing as a group of biopharmas look to replace their aging franchise therapies that are losing patent protection.
The upside to that dealmaking will be better therapies to control some debilitating and very lethal diseases. The downside is that there’s less and less interest in marginal products. Right now, small markets are the kiss of death.
1. Merck:
When being No. 1 means you have to try even harder
The scoop:
Over the past 12 years, Merck has become the Keytruda company — with a few other blockbusters. And in the next few years it will try to become the Big Pharma that kept investors happy after moving on from its record-setting franchise.
A lot of people in the business don’t remember that back in 2014, when Keytruda was approved, Merck was barely a player in oncology. Now it’s fleshed out a plan to transition to an injectable version of Keytruda that can keep a chunk of that megablockbuster revenue flowing, while looking to M&A to fill the rest of the gap when the original version loses exclusivity. And analysts have been fretting about Merck’s wobbly sales pace for its Gardasil vaccine, which adds only more pressure.
Just weeks ago it worked out a hefty
$6.7 billion deal
for Terns Pharmaceuticals to see if it could steal a march on Novartis’s Scemblix and earn billions more in chronic myeloid leukemia — which will have a lot to do with its ability to win an initial OK and then migrate to frontline therapy.
It’s all part of a $15 billion M&A commitment that Merck has made to oncology since the start of 2019, and there may well be billions more added to that tally by the end of this year.
Even at a significantly reduced budget of close to $16 billion, Merck remains the top spender in drug R&D among the big 15. And while oncology has been dominant, CEO Rob Davis and research chief Dean Li have been casting a wide net.
Merck has been focused on drugs with a clear clinical timeline to market, which helps explain why it came out on top of a bidding war to gain
Cidara Therapeutics’ antiviral drug
for a bit more than $9 billion last year. And it plunked
down $10 billion
for Verona Pharma and its newly approved COPD drug.
Merck can never expect to have another Keytruda and is building around a steady development of several major market drugs.
On that list are the oral PCSK9 enlicitide decanoate, which posted positive results for
slashing bad cholesterol
in Phase 3 a few months ago. Merck has been racing AstraZeneca on that front, looking to score big after the original injectables failed to catch on. The FDA is reportedly fast-tracking the review along with sacituzumab tirumotecan (sac-TMT), a TROP2 ADC from a multibillion-dollar deal with Kelun. That’s performed well in Phase 3 when paired with Keytruda — so well that the investor Blackstone will
provide $700 million
to back Merck’s 15 Phase 3s in 2026. Another Phase 3 combo program is underway for calderasib (MK-1084) with an eye on developing a new KRAS G12C drug.
Then there’s the IBD drug tulisokibart (MK-7240), which has been moving into a slate of Phase 3 trials.
In the meantime, Merck has been working on a patent strategy that could extend its market protection for Keytruda well into 2029 while positioning its new injectable version of Keytruda to maintain a large chunk of the market after biosimilar manufacturers try to position their new IV knockoffs.
2. Roche: Setting its sights on new franchises and spending big on R&D
The scoop:
For a player like Roche, with two R&D organizations straddling the globe, late-stage development requires plenty of forward thinking about the franchises its losing, and the franchises it needs to add to keep the Swiss francs adding up.
On the way out: Its top-selling drug Ocrevus, which earned $9 billion and change last year and faces a patent cliff starting in 2029.
On its way in: High hopes are being polished up among the analysts with positive
Phase 3 data
for fenebrutinib, an oral BTK inhibitor Roche badly needs to look better than its top blockbuster.
The drug achieved non-inferiority to Ocrevus in the first big Phase 3 readout for primary progressive multiple sclerosis, with a 12% reduction in the risk of disease progression. Researchers also boasted about a particularly positive readout for upper limb function. A Phase 3 for relapsing MS is looming, and will make up the key data being presented for an approval on both types of MS.
A lot is riding on that one. If they can make a case for marginally better efficacy in an easier-to-take pill (and overcome worry about higher numbers of elevated liver enzymes), they can make a case for a blockbuster replacement. If not, the generic rivals will bust up the franchise, leaving a large gap to fill elsewhere.
That could have come from giredestrant —
until a Phase 3 failure
in estrogen receptor-positive, HER2-negative breast cancer. The setback left analysts to ponder whether Roche’s dreams of a multibillion-dollar treatment had been destroyed. Roche has put in an application with the FDA, expecting that the positive late-stage data in advanced breast cancer will be enough. But the trial flop could limit its market potential.
Roche has a lengthy Phase 3 path to travel on trontinemab, the latest amyloid buster to rouse excitement in Alzheimer’s. Evidence shows
it can remove amyloid
, a chief suspect in disease progression, along with reduced brain bleeds that afflict the two marketed rivals from Eli Lilly and Eisai/Biogen. But it will take quite some time to see how the drug works on disease progression — particularly when they cross over to prospective patients who have yet to suffer from memory loss. If it can get over the finish line, there’s significant potential.
Roche is also attempting a late entry in another key category that has gripped many in the R&D 15: obesity. And it’s following the same route Eli Lilly took with Zepbound, pursuing a
GLP-1/GIP combo approach with CT-388
that produced weight loss numbers that were quite similar to the drug that now dominates new prescriptions. There’s excitement about data showing the drug could avoid plateauing on weight loss. But Roche has a long way to go in Phase 3 before it can prove it.
It grabbed CT-388 with its $2.7 billion deal for Carmot Therapeutics, which also came with the drug CT-996. Roche has high hopes for a triple approach with petrelintide, an amylin analog from Zealand Pharma.
Traditionally, once a drug seizes a best-in-class rep, it’s incredibly difficult to leapfrog it in the market. But the obesity market may be so substantial that those rules may not hold.
Back in early 2023, Roivant CEO Matt Gline was beaming when he talked about the Phase 2b data that they were seeing for RVT-3101, freshly picked up in a deal with Pfizer. Roche R&D execs were also pretty impressed, and nabbed it for $7.1 billion. Roche put it right on a Phase 3 fast track for ulcerative colitis and Crohn’s disease.
The anti-TL1A antibody approach has become a hot target at several big players. Merck made it a top priority with its $10.8 billion deal to buy Prometheus Biosciences and tulisokibart (MK-7240). Sanofi and partner Teva have their own advanced program for duvakitug.
CEO Thomas Schinecker includes pegozafermin, a MASH drug obtained in the 89bio buyout, and zilebesiran, a hypertension therapy partnered with Alnylam, among other top prospects in Phase 3. Zilebesiran was steered into Phase 3 despite a mid-stage flop, but the partners believe they found the right dose to take to regulators, once they have the data to back it up.
Dealmaking got Roche to this point with its late-stage pipeline, and dealmaking will continue to be a big part of its planning. Schinecker also hasn’t ignored the pacts being signed in China, so don’t be surprised if you hear of something new on that score in the coming months.
3. AstraZeneca: With more than 100 Phase III studies underway, analysts expect some big readouts in 2026
The scoop:
When I started working on this piece in late March, AstraZeneca’s R&D group scored two significant wins on the same day. Its
in vivo
CAR-T, acquired in a $1 billion deal for EsoBiotech, provided a batch of
promising data on a tiny group
of multiple myeloma patients. And its COPD drug tozorakimab,
an IL-33-targeting antibody
, came up with positive results in a pair of Phase 3 studies — after it flunked Phase 2.
Significantly, tozorakimab was developed internally by the R&D team and has a peak sales estimate of $3 billion to $5 billion.
For CEO Pascal Soriot, the announcements offered a payback for AstraZeneca’s growing bet on R&D, which jumped to $14.2 billion in 2025 from $13.6 billion in 2024. And listening to Soriot discuss the strategy to grow revenue to $80 billion in 2030, there was no mistaking the company’s intention of staying laser-focused on drugs with multibillion-dollar potential.
Soriot got into the top five based on a slate of new cancer drugs that reversed years of failure in the clinic. But with Novo Nordisk offering the latest lesson on how a reliance on a couple of big drugs can blow up in your face, he’s built a development machine that entered this year with more than 100 Phase 3 studies underway.
“Think about that,” he told his audience of investors and analysts. “One hundred Phase 3 trials. It’s an enormous momentum going through the pipeline. This year, we should have 20 Phase 3 readouts. Those readouts, fingers crossed of course, if they are positive, they will collectively drive another more than $10 billion of peak revenue.”
In vivo
was called out in that session. Along with the
oral PCSK9 drug AZD0780
, an
oral GLP-1 called elecoglipron
that scored in Phase 2, eight homegrown ADCs, radioligands, next-generation I/O bispecifics, “in particular, rilvegostomig” (despite all the problems we’ve seen with TIGIT), the
BCMA/CD19 targeted AZD0120
, gene therapies and a new T cell-engager platform, you could say that AstraZeneca has a lot on its plate.
Soriot is a relentless builder. That’s led to one of the most ambitious plays in China, where he has devoted a considerable amount of his own time sharpening both commercialization and R&D activities in a country that has made biopharma a top economic priority.
4. Eli Lilly: The most stubborn drug developer in biopharma goes big on R&D as obesity delivers
The scoop:
Now that Eli Lilly has sealed its rep in obesity, trouncing a sometimes hapless Novo Nordisk, it’s no wonder that Eli Lilly juiced its R&D budget in 2025, betting on a new round of obesity drugs while broadening its research horizons.
The FDA wasted no time in approving Lilly’s oral obesity pill orforglipron, perhaps a reward for all the US investments the pharma giant has laid out before the Trump administration. It’s a few months behind Novo, but Lilly has already shown that its marketing team can compete and catch up. And then there’s retatrutide, its
next-gen triple-G agonist
, where Lilly has been balancing solid results on weight loss against side effects.
Staying out front of the global obesity market won’t be easy, as other giants like Merck and Pfizer make a play. But Lilly earned its kudos and has a lot of momentum in its favor.
Lilly had similarly high hopes for its Alzheimer’s drug, but weak efficacy data has been holding down sales. Can it
do better with remternetug
, its N3pG beta amyloid treatment? The late-stage candidate hopes to do what the current marketed drugs do in clearing amyloid, but with a self-injectable that will be far easier for patients to use than the current infusion. That could go a long way to building a market of patients who are at risk of developing Alzheimer’s years in the future.
There’s a lot here to suggest that Lilly faces another uphill battle in Alzheimer’s, but it’s been plugging away for decades and shows no signs of conceding defeat or calling a truce. To the contrary.
Last fall CEO David Ricks wooed Carole Ho away from her chief medical officer post at Denali Therapeutics — which is celebrating its first new drug approval — and made her president of Lilly Neuroscience. Ho led the
$6.3 billion deal
to buy Centessa Pharmaceuticals and its sleep disorder drug cleminorexton (ORX750), which is in Phase 2 studies for narcolepsy and hypersomnia.
That’s a chunk of change for a relatively small market, which has analysts thinking that Eli Lilly may be targeting a much bigger opportunity in fatigue and focus — which would put it much more squarely in a familiar realm of population drugs.
Alongside Ho, Adrienne Brown was put in charge of immunology while R&D chief Daniel Skovronsky was given a range of impressive titles. So look for more deals in the hot immunology arena as well.
Lilly is not known for big buys. It’s been more comfortable going after biotechs like Scorpion Therapeutics, where it beefed up its oncology franchise with the
PI3Kα inhibitor program STX-478
, targeting a large segment of the breast cancer market. Now in Phase 3, it now faces a major rival in Novartis, which recently picked up its own contender for a less-toxic treatment in a $2 billion deal for Synnovation Therapeutics.
Lilly upped the ante with its purchase of Kelonia Therapeutics, putting up $3.25 billion in cash on a $7 billion M&A deal that will make Lilly a player in
in vivo
CAR-T. That’s become one of the hottest areas in biotech, with companies looking to leapfrog allogeneic versions of CAR-T that have moved slowly in the clinic. The original autologous CAR-Ts delivered jaw-dropping data, but in the US have proven too complex for most community cancer centers, where the bulk of treatment occurs.
Lilly has built a powerhouse rep, and it’s also splashing some big bucks to go after a new supercomputer with Nvidia. It remains one of the top pharmas to watch in R&D after going a long way to shed its historical profile as the slowest player in the big leagues.
5. Johnson & Johnson: The healthcare giant breaks out of the pack with a big buyout
The scoop:
CEO Joaquin Duato likes to talk up the big numbers of marketed and experimental drugs under J&J’s broad umbrella. But when it comes to near-term growth drivers, the discussion always boils down to a few big players.
On the marketed side, that means products like Carvykti — the star CAR-T out of China — and Tecvayli, which sparked recent headlines with data for treatment-refractory patients with multiple myeloma. The Tecvayli/Darzalex combo was approved as a second-line treatment for multiple myeloma after FDA Commissioner Marty Makary blessed it with a special rapid review. Its IL-23 Tremfya has broken above $5 billion in sales, leaving the ever optimistic Duato predicting a crest above $10 billion. The psoriasis drug icotrokinra Icotyde just got approved, the payoff of a longtime partnership with Protagonist and another example of how J&J’s business development team has scored key deals for the pharma giant.
J&J has reportedly been in talks to
buy its partner Protagonist
, but so far, no deal. And with Protagonist’s stock up 140% over the last year, that would be pricey.
And now that Caplyta has an FDA nod to be marketed for major depressive disorder — the big bet that came with the Intra-Cellular buyout for $14.6 billion — J&J can be more confident about generating more than $5 billion a year for that franchise.
That has all gone a long way to putting the loss-of-exclusivity for Stelara in the rearview mirror, as Duato delighted in telling analysts during its Q4 results call.
It’s not all been clear sailing. The Phase 3 for another depression therapy,
aticaprant, was axed
after it flopped (as did a key rival). And its other big depression program for seltorexant stumbled badly last fall when it failed to beat out quetiapine extended release in a head-to-head study. J&J had vowed to stand by both after the Caplyta buyout. And there’s been plenty of trouble for milvexian, partnered with Bristol Myers Squibb, which I cover in more depth below.
One of its next big bets was on Halda, which J&J
made a deal to acquire for $3.05 billion
at the end of 2025. Using tech derived inside the biotech — which was founded by Yale’s Craig Crews — researchers targeted BRD4 “in the presence of androgen receptors” common in prostate cancer, as explained by
Endpoints’ science writer Ryan Cross
. In an early study, the biotech found it had a big impact on PSA, a common biomarker for prostate cancer. And the same approach on RIPTACs — regulated induced proximity targeting chimeras — could work in a variety of cancers, potentially overcoming resistance to currently marketed drugs with more targeted therapies that protect most healthy cells.
Another closely watched drug in J&J’s pipeline is JNJ-4804, which targets both IL-23 and TNF. Now in Phase 2, the drug is going after ulcerative colitis, psoriatic arthritis and Crohn’s disease. And Royalty Pharma
recently bet $500 million
on its future in autoimmunity, co-funding its research work on the therapy for the next two years.
That’s quite a vote of confidence.
Whatever happens with Protagonist, J&J has been effective in nailing down blockbuster deals over the years. And those deals have included some very bold plays, which include reaching into China for a game-changer when that was still considered suspicious.
Given its track record on M&A, you can expect more ahead in 2026.
6. Novartis: Key patent losses keeps R&D scrambling to build late-stage pipeline
The scoop:
Novartis has been engaged in a delicate balancing act with its late-stage portfolio and rising generic competition. In its 2025 wrap-up, CEO Vas Narasimhan championed sales growth for mainstays like Kisqali, Kesimpta and Pluvicto. All of which are expected to help fill up most of the $4 billion hole this year from the loss of exclusivity on Entresto, Promacta and Tasigna. And with Cosentyx facing knock-offs by the end of the decade, there’s lots more pipeline work being done.
Some analysts have been keen to trumpet the blockbuster potential for ianalumab, which has been
racking up positive data
for Sjögren’s disease and primary immune thrombocytopenia (ITP), a blood disorder. It’s targeted at B-cell driven autoimmune disorders. Novartis bagged that drug in its buyout of MorphoSys in 2024, which was primarily about pelabresib — an oncology program that
quickly disappointed
Novartis with evidence of emergent malignancies.
It’s going to take more than one added blockbuster to satisfy Wall Street. So it was at least slightly concerning that Narasimhan had to push back readouts for its late-stage pipeline by a bit.
Now zigakibart is
expected to deliver
Phase 3 data for kidney disease in early 2027, while pelacarsen should make its Phase 3 data debut for lipoprotein(a) now in the second half of this year.
Zigakibart is part of a trio of drugs in the pharma giant’s IgA nephropathy (IgAN) portfolio — including Vanrafia and Fabhalta. A couple of months ago Vanrafia flunked a Phase 3 for kidney function decline. That was close enough for Novartis, though, as it plans to push ahead on an OK for a full IgAN approval.
Travere had a near miss on the same endpoint, before its approval, so don’t count Novartis out yet.
Novartis likely had Cosentyx in mind when it agreed to pay $12 billion for Avidity last fall. That’s significantly higher than the single-digit billion-dollar deals Narasimhan is noted for — which includes Tourmaline Bio, Regulus Therapeutics and recently Excellergy.
There are two key late-stage drugs that inspired the neuromuscular deal, along with multibillion-dollar expectations for both: delpacibart etedesiran (del-desiran) for myotonic dystrophy type 1 and delpacibart braxlosiran (del-brax) for facioscapulohumeral muscular dystrophy (FSHD).
The deal also delivered delpacibart zotadirsen (del-zota) in Duchenne muscular dystrophy, and with it peak sales projections that tend to drag well below the blockbuster mark. That should be first up for an accelerated approval.
Like everyone else in the R&D 15, R&D disappointment is no stranger to Novartis. There were blockbuster expectations for the PI3K drug Piqray in breast cancer at one point, but Roche has crimped any future in that direction with the approval of Itovebi, a better drug that’s now used in frontline therapy.
Novartis, though, doesn’t take disappointment lying down and handed over $2 billion upfront to get a new drug that could do Piqray and Itovebi one better. The drug is dubbed SNV4818, and Novartis threw in a billion dollars in milestones to purchase the PI3Kα a few weeks ago.
The big idea here is that
SNV4818 is designed
to spare healthy wild-type PI3Kα and zero in on the mutated version in cancer cells. That may well eliminate much of the toxicity that has scuttled its combination approaches. And it gives Novartis a new inside track on a market that covers 40% of HR+/HER2- breast cancer, plus a shot at other solid tumors as well.
The pipeline isn’t solely devoted to blockbusters. Novartis developed Coartem for malaria decades ago and sold it cheap in Africa, priced for the poor. Now that new drug-resistant strains are coming along, Novartis has a new combo approach — GanLum — that
looks just as effective
.
There are major players that wouldn’t spend a dime on a program like that. So some added credit is due here.
Don’t be surprised if Novartis picks up its pace on China deals. Argo Biopharma (a siRNA deal focused on cardio) and SciNeuro (in Alzheimer’s) have been brought into the fold recently. And R&D was a big focus with its announced
$480 million plan
to beef up research and manufacturing in China.
7. Pfizer:
Repeated setbacks taint expectations, setting the stage for more dealmaking
The scoop:
Pfizer has a long tradition of disappointing investors in late-stage development, and 2025 was no exception.
Its sickle cell drug inclacumab
went down in the summer
, following the withdrawal of Oxbryta. That led to some quick math about the billions wasted on the Global Blood Therapeutics buyout — though the GBT drug osivelotor remains in the pipeline. Danuglipron for obesity
was a bust
. Not surprisingly, a raft of experimental programs at Seagen were slashed, but that’s standard after a big buyout. Pfizer’s once magnificent hopes for gene therapy came to an end — along with much of the rest of the AAV field — early last year with the discontinuation of sales of the $3.5 million hemophilia B treatment Beqvez. And weeks ago trouble recruiting for the Phase 3 study of its
Lyme disease vaccine
left a cloud over that program. It failed the trial, but will still go to the FDA.
The Lyme disease setback was quickly followed by an even more troublesome issue with its newly acquired obesity drug that came with the $10 billion Metsera acquisition. It didn’t fail. But it also
didn’t impress many analysts
with its rather bland weight-loss numbers. Pfizer is going for an easier monthly dose, but it’s also arrayed against two well-established leaders in the field, both of which are doubling down on their own second-gen therapies. And they aren’t alone.
The bar for carving out a blockbuster entry in obesity will be very high. And based on their Q4 review of the pipeline you can expect a relentless focus there.
Every time Pfizer experiences a flop, it goes right back at it, inking new deals and pushing R&D to come up with a few new winners. And there’s always plenty to talk about in Phase 3 to excite the discussion anew.
Pfizer trumpeted an early cut of
Phase 2 progression free data
for atirmociclib, which it hopes can succeed its blockbuster CDK 4/6 Ibrance in breast cancer. Ibrance earned more than $4 billion last year despite less competitive overall survival data compared to its rivals. Atirmociclib, though, still has a few years of late-stage trials before it can reach the finish line on first-line metastatic breast cancer.
Cancer clearly remains a major focus in Phase 3. The company has accelerated its work on mevrometostat for castration-resistant prostate cancer, looking to build on the positive data researchers gathered in a Phase 1 combo with Xtandi and released at an ASCO session early last year. Metastatic treatment-resistant patients
saw a 49% decline
in the risk of disease progression or death. Pfizer has big hopes for this drug with a rapid-fire plan to win a quick FDA OK, but some analysts have been less than impressed with the data they’ve seen on other EZH2 inhibitors, which they believe might be typical of the class.
Now that Pfizer has had a chance to absorb the Seagen pipeline and come up with its own strategy, several of its oncology plays are centered around the assets it acquired.
There’s the anti-HER2 ADC disitamab vedotin, multi-tumor prospect sigvotatug vedotin, which should offer up Phase 3 data soon, and PDL1V, another Seagen ADC that targets PD-L1 expressing cells with a microtubule-disrupting agent payload. Pfizer is pushing PDL1V — which could theoretically work in a variety of tumors — into pivotal trials, but it’s been flying largely under the radar. Henlius, though, has flagged its own rival in the pipeline, touting its prospects of coming out ahead.
Finally, prifetrastat (PF-07248144) is one of several KAT6 inhibitors angling for the spotlight.
Padcev remains Pfizer’s top prospect for expansion studies, with a slate of new Phase 3 data on bladder cancer. Their Padcev/Keytruda combo beat out chemo in pre-surgery muscle invasive bladder cancer cases, promising a larger market for their drug out of Seagen. It’s unlikely, though, that Pfizer can recapture the glory it saw during the pandemic, when its mRNA vaccine out of BioNTech scored huge — only to fade away faster than once thought possible.
8. Bristol Myers Squibb: A high wire act in Phase 3 leaves analysts fretting over their future
The scoop:
Bristol Myers has had its sights set on a successor franchise to Revlimid and Pomalyst for years. And 2026 is shaping up as a key turning point for a pair of programs that it hopes will fill at least part of the big gap left by generic competition.
Iberdomide and mezigdomide are a pair of molecular glues in late-stage development. Mezigdomide looked promising in a Phase 2 study back in 2023 and was hailed earlier this year for
a win in Phase 3
. We have yet to see the numbers, and some analysts want to see how the head-to-head with Pomalyst looks.
Iberdomide, meanwhile, is
up for a decision by the FDA
this summer in multiple myeloma, and could break past into blockbuster territory once on the market.
They’re both part of a class of drugs dubbed cereblon E3 ligase modulators (or CELMoD), the tip of the spear in protein degradation.
A third CELMoD, golcadomide, is in Phase 3 for large B-cell lymphoma.
Meanwhile, dark clouds have been gathering around another late-stage effort for milvexian, an oral factor XI inhibitor that was seen as a potentially safer successor to the blockbuster blood thinner Eliquis. Two big studies have been scrapped on
disappointing data
and the field is still reckoning with the failure of Bayer’s oral program. Regeneron and Novartis are pursuing injectables, which could eclipse the orals on efficacy.
Admilparant, an LPA1 antagonist which showed promise in Phase 2 for IPF, is up on the frontlines of the pipeline, along with pumitamig (PD-L1 x VEGF-A bispecific antibody with BioNTech) and the radiopharmaceutical RYZ101, which it picked up in its $4.1 billion buyout of RayzeBio. Like most of the rest of the top 15, Bristol Myers has been busily scooping up biotechs to fill out late-stage development work. Cobenfy, approved for schizophrenia in 2024, came out of Karuna and is now triggering worries for its late-stage program for Alzheimer’s-related psychosis following some “irregularities” that delayed an interim analysis.
Bristol Myers doesn’t have a whole lot of wiggle room when it comes to the late-stage pipeline. It’s being hammered by the loss of exclusivity on legacy blockbusters, with Opdivo — its current reigning champ — and Eliquis looking at a patent cliff event over the next two years. Revlimid is in significant decline. Even with some new approvals contributing to the bottom line, revenue slipped slightly in ’25 and will slide even more this year. So its recently approved franchise therapies and Phase 3 drugs are all meant to be major contributors. Which is why this is one big pharma that frequently signals to the biopharma research community that it has no appetite for small-market drugs.
Its late-stage pipeline today was shaped by a reorganization two years ago that dropped a slate of early and mid-stage drugs from the pipeline that were seen as runners-up in the blockbuster race. That included a CTLA-4 drug that was seen as a cut below Yervoy. To compete at the level Bristol Myers needs, it all has to be seen as a contender for first-in-class or best-in-class.
9. GSK: The new CEO brings some added enthusiasm to play in bolstering late-stage expectations
The scoop:
Historically, GSK has avoided anything flashy. It buys biotechs, but tends to go for deals on the lower end of the bolt-on spectrum. Its pipeline work has drawn mixed reviews over the years, sometimes spurring the activist set. The new, new thing in biopharma R&D often fails to pan out, as they know from experience: One of the reasons why the company saw a spike in R&D spending last year was due to its write-off on belrestotug, its TIGIT program partnered up while Hal Barron was running research. The original deal with iTeos cost $625 million in upfront cash.
GSK execs aren’t diving into obesity, either — or at least they’re avoiding the commercial slugfest developing around GLP-1. It’s too frothy.
So after its ADC Mo-Rez (mocertatug rezetecan) delivered strong Phase 1b data recently, it caught our attention. GSK heralded the results in platinum-resistant ovarian or endometrial cancer as “extremely exciting.”
They were so excited, GSK said it would
pivot directly into an ambitious Phase 3 program
with five trials set to begin soon. The ADC was picked up in a modestly priced licensing deal with China’s Hansoh Pharma, targeting the B7-H4 antigen. And if the data hold up, it will help make GSK a force to be reckoned with in oncology, a goal that ex-CEO Emma Walmsley set out when she took over a decade ago, picking up the pieces left from its big asset swap with Novartis in 2015.
It’s not the only ADC GSK is excited about. The other one — also from Hansoh — is dubbed Ris-Rez and targets small cell lung cancer. And GSK talked it up in its Q4 review earlier this year.
GSK expects Mo-Rez can earn more than £2 billion a year, making it a key drug in its quest for £40 billion a year in revenue by 2031, up from 2025’s £32.7 billion. Ris-Rez, meanwhile, has picked up a breakthrough drug designation at the FDA and earned equally ambitious peak sales projections.
You could call all that part of GSK’s tactical acceleration to achieve some longtime strategic goals.
Rounding out its newfound prospects for oncology, GSK is following up its
IDRx buyout
— just $1 billion upfront — from a little more than a year ago with pivotal plans for velzatinib/IDRX-42. If successful, new CEO Luke Miels will be headed back to an arena where he found success at AstraZeneca.
And he clearly relishes that prospect.
Miels, a marketing specialist, offered a similar projection for bepirovirsen, its
hep B “functional cure,”
projecting equivalent peak sales as its top cancer drug. Some of the analysts, though, aren’t ready to go that high, but breaking the billion-dollar mark seems plausible.
Then there’s the pivotal campaign for camlipixant, a P2X3 antagonist for chronic cough that’s also earned some blockbuster love, along with fresh prospects for Jemperli.
In some respects, a brighter horizon for GSK reflects Miels’ cheerleading style for R&D. Where Walmsley would be somewhat more guarded, he’s encouraged high expectations. But it’s also based on hard data, where he’s been reaping the rewards of Walmsley’s dogged ambitions for GSK.
GSK has had its successes, particularly with Shingrix, which has helped the company manage the market volatility for RSV. And the global pharma continues to talk up vaccines and respiratory, two traditional mainstays. It’s also been a longtime runner-up to Gilead Sciences in the HIV market, whereto majority-owned ViiV continues to carve out significant sales.
10. AbbVie: Researchers set out to prove they’re more than just Skyrizi and Rinvoq
The scoop:
Skyrizi and Rinvoq kept AbbVie in the big leagues as Humira gradually faded, and AbbVie execs have shown real zeal about adding to those franchises with new indications. Moving beyond its two cash cows, there’s a slate of late-stage bets — along with an aspirational play for
in vivo
CAR-T.
AbbVie had high hopes for their neuroscience division when it paid $8.7 billion to get Cerevel Therapeutics back in 2024. The readouts aren’t over, but it had to write off a good chunk of that when
emraclidine foundered
in a slate of schizophrenia studies. It did better with another drug picked up in the buyout when tavapadon hit in Parkinson’s. But execs can’t be happy with a group of modest peak sales projections.
Maybe AbbVie can do better with bretisilocin, a 5-HT2A receptor agonist and 5-HT releaser which had a positive slice of Phase 2a data for major depressive disorder when AbbVie bought it. It
didn’t pay a lot for it
, failing to disclose the upfront and leaving it at $1.2 billion all in for the psychedelic mimic. But it’s a near-term shot in a high-risk segment of the business that’s been seeing growth with Vraylar and Ubrelvy.
ABBV-932 rounds out its top neuroscience prospects.
Then there’s oncology.
Etentamig (ABBV-383) was dosed in Phase 3 for the first time in 2024, highlighting AbbVie’s shot at a BCMAxCD3 bispecific T-cell engager for multiple myeloma. By silencing the Fc tail, researchers hope to extend the half-life of the drug so they can stretch dosing to every four weeks. The downside is it’s late to the game, with Tecvayli from J&J out on the market and following up with a recent accelerated approval for a combo with Darzalex.
Pivekimab sunirine (PVEK), its CD123 ADC, was filed with the FDA last fall. AbbVie submitted early Phase 1/2 data for blastic plasmacytoid dendritic cell neoplasm (BPDCN), a rare and aggressive cancer, hoping to gain a quick entry on a drug that earned a breakthrough drug designation from the FDA.
Temab-A (ABBV-400), which targets cancers with high c-Met expression, got started in Phase 3 with a shot at colorectal cancer, where it’s being studied as a monotherapy and in combo with bevacizumab. AbbVie is looking for data from a slate of mid- to late-stage studies to gauge its potential for solid tumors.
ABBV-706, an SEZ6 targeted ADC, is in trials for solid tumors.
You don’t hear a lot about BoNT/E, the fast-acting botulinum neurotoxin that works for shorter spans. AbbVie picked it up in mid-stage development way back in 2018, bringing a potential rival to Botox in-house. That treatment was filed with the FDA a year ago and could bolster its big business in aesthetics.
AbbVie will go the distance when it feels there’s sufficient upside, and that’s what it committed to do when the company bought Capstan Therapeutics, one of the leading early-stage biotechs working in the sizzling
in vivo
CAR-T space. There’a a very long row to hoe on that one, but a win here could put AbbVie out front with a next-gen cancer drug. Also, the $2.1 billion deal didn’t cost the moon, giving it a relatively low-cost method for generating some added enthusiasm for the pipeline.
11. Sanofi: Another new CEO looks for a better path in Phase 3
The scoop:
Big pharma players with a solid record of success in Phase 3 don’t fire their CEOs, so Paul Hudson’s exit a couple of months ago set the stage for a recital of all his setbacks in R&D.
Hudson bet big on immunology at Sanofi, where board members have repeatedly demonstrated their readiness to make a change when they feel a course correction is needed. So when the strategy failed to set the company squarely on a new path that could carry it past Dupixent, the enormous cash cow (out of Regeneron) that pays the freight at Sanofi, the rest was Greek theater.
New CEO Belén Garijo will need to make some tough decisions, and that typically starts with the pipeline.
About two years ago, and Hudson was cheering mid-stage data for its anti-inflammatory OX40L drug amlitelimab, once billed as a product that could excel over Dupixent. These days execs spend their time defending mixed data from the Phase 3 program that followed. Analysts highlighted lower efficacy than Dupixent for atopic dermatitis and Sanofi has been dropping indications instead of building the pipeline-in-a-product it badly needs.
It’s not a bust, but it is far from megablockbuster heaven.
The same “meh” reaction followed last summer’s approval of rilzabrutinib (Wayrilz) for immune thrombocytopenia (ITP). That followed a major setback on atopic dermatitis. It edged ahead after the key drug in the Principia Biopharma buyout, tolebrutinib, hit a series of setbacks in clinical trials. And the oral TNF inhibitor balinatunfib has gone down on data twice in mid-stage studies, failing an effort to prove it could work as a monotherapy.
That only leaves the anti-CD40L frexalimab from one of Hudson’s old lists of experimental drugs capable of generating more than $5 billion a year in revenue. That’s in Phase 3 for multiple sclerosis, after a failure for Sjögren’s disease. Its bispecific lunsekimig, meanwhile, hit in mid-stage studies of asthma and an inflammatory sinus condition, but flopped in atopic dermatitis.
Sanofi stayed at the blockbuster feast because of its partnership with Regeneron on Dupixent. But it’s only managed to disappoint analysts with its COPD data for next-gen drug itepekimab. That leaves R&D working away to keep building Dupixent as it gets close to a reckoning on its patents in the early 2030s.
Sanofi also isn’t catching much of a break with analysts on its latest CEO pick. Garijo never established Merck KGaA’s reputation in R&D and there are plenty of doubters that she can do it at Sanofi. Nevertheless, she’s holding the reins now.
12. Novo Nordisk: After fumbling the lead in obesity, R&D hunts for a new way forward
The scoop:
Novo Nordisk has been channeling more cash into its R&D division in recent years, emboldened by its mega-blockbuster semaglutide and vowing to beat back competition from fierce rival Eli Lilly.
Some of that money went into a critical head-to-head study of its GLP-1/amylin analog against Lilly’s tirzepatide, and Novo lost — big time. With that went its market cap, which has shriveled back to levels from five years ago.
Novo had a big head start on Eli Lilly, and its steady backward drift — as a tide of new GLP-1s courses through the industry — has escaped no one. The early arrival of its pill has done nothing to change perceptions.
Novo Nordisk isn’t done with CagriSema. With weight loss averaging more than 20%, why should it be? But a lot of attention has now shifted to zenagamtide (amycretin), a GLP-1/amylin dual agonist that was slated to begin Phase 3 this year with both oral and injected versions.
Adding insult to injury, Novo also had to concede late last year that its Phase 3 attempt to see if semaglutide could help Alzheimer’s patients came up as a loser. Practically no one expected it to succeed, and Novo clearly flagged the lottery-ticket strategy, but the stock took a hit anyway.
But don’t expect Novo to take a back seat on obesity just because of a few key setbacks. That was apparent with its failed bid for Metsera, which fell — inevitably — to Pfizer. But it also has UBT251, a triple agonist that it hopes can put it out front again.
Long a leader in the diabetes field, Novo is also using its deep scientific knowledge in the field with its late-stage program for ziltivekimab to see if it can make an impact in cardiovascular disease with a disease-modifying approach. That’s another huge field, with enormous potential — if Novo can catch a break.
13. Amgen: The new R&D chief is determined to score big with protein degradation
The scoop:
The star of Amgen’s late-stage pipeline remains MariTide, its obesity drug that’s meant to rival the two frontrunners. There’s a twist, built around GIPR antagonism, as opposed to the agonism that has succeeded so far.
Over the years, Amgen CEO Bob Bradway has managed to foster star coverage for its leading experimental drugs. Lumakras, its breakthrough offering on KRAS, was an example of that. But there are too many questions about the obesity market now to get a free pass among the analysts. For Amgen to succeed with obesity, it’d have to clear a very high bar on efficacy (that hasn’t looked good), hold the line on safety and offer a better dosing regimen to compete against the leaders.
That’s not so easy, particularly as more and more rivals elbow their way into the obesity pipeline looking for any kind of edge. And Amgen still has a ways to go before it can hunt up an approval.
In reviewing 2025 for investors, R&D chief Jay Bradner offered its Phase 3 drug desodoliveb for Sjögren’s disease. And there’s daxdilimab.
But there were also a couple of duds.
Bradner axed an anti-OX40 rocatinlimab collaboration with Kyowa Kirin — a precursor to the Japanese company’s decision to scrap the whole thing — as well as the program for bemarituzumab, which produced positive data for gastric cancer.
Where Amgen execs see their most likely near-term growth coming from is franchise expansion. But it has also had to stare down the FDA, which wanted their rare disease drug Tavneos to be pulled after the agency spotlighted a reassessment of the benefit/risk profile the drug presented.
Given the less-than-stellar prospects of its late-stage pipeline, it’s no wonder that Bradner turned to Dark Blue, buying out the UK biotech for $840 million earlier in the year. The move puts Bradner back in the driver’s seat of protein degradation, an early and long-lasting love of his.
14. Regeneron: Cash cows carry it through a challenging fight in R&D
The scoop:
Regeneron execs Len Schleifer and George Yancopoulos have been working hard to get investors excited about the near-term prospects for their late-stage pipeline, beginning with their LAG-3 fianlimab. But it can be an uphill climb with the analysts who haven’t been that impressed with late arrivals in questionable markets.
Where Yancopoulos boasts about a drug with “best-in-class” potential, some analysts wonder how well they can do with a follow-up to Bristol Myers Squibb’s pioneering LAG-3. Bristol Myers has had plenty of trouble with the drug since it was approved. It isn’t effective enough to work as a monotherapy, so it has to be rolled out as a combo. Follow-up studies have failed, and rivals like Immutep have seen their candidates go down in flames, blighting a field that once looked like it would be a successor to CTLA-4 and PD-L1.
Libtayo (the PD-1 cemiplimab) itself is a follow-up to Keytruda and Opdivo. It hit the $1.4 billion revenue mark last year, leaving it well behind the two market leaders as Regeneron’s research group pushes for expanded approvals. Building that market with combo drugs like fianlimab is important for Regeneron, which is run by two of the most persistent self-made billionaires in biotech. Another oncology prospect in mid-stage development is marlotamig (REGN7075), an EGFRxCD28 bispecific.
Cancer overall, though, has proven to be a complex challenge for Regeneron. Odronextamab — its CD20xCD3 candidate — has run into repeated issues at the FDA as rivals crowd around. And when linvoseltamab was approved last summer (as Lynozyfic) for multiple myeloma, more delays had forced it further behind rivals at Pfizer and J&J.
In addition, Regeneron has had trouble following up on its own market leader Dupixent. Last year its next-gen replacement partnered with Sanofi, itepekimab, fizzled in late-stage testing. But during Regeneron’s Q4 call Schleifer shifted the spotlight to “long-acting IL-13, IL-4, and IL-4/13 bispecifics as well as… a new soupy doopy molecule that is a new version of Dupixent that was naturally selected that might have even more improved properties.”
In the meantime, Sanofi’s been working on legal strategies to extend the patents on Dupixent to 2040. Where R&D fails, patent thickets can often fill the gap.
Regeneron has a much broader focus these days than when it achieved early successes with Dupixent and Eylea, including a foray into obesity and diabetes. Last summer it forked over a modest $80 million in cash to China’s Hansoh Pharma for olatorepatide, a GLP-1/GIP drug following elimination Lilly’s dominant play with tirzepatide. Even Regeneron says the drug looks similar to Eli Lilly’s, but it sees it as a pathway to developing a better drug that can maintain the muscle lost to GLP-1s. China’s late-stage trial passed muster in Phase 3 a few weeks ago, with Regeneron laying the groundwork for pivotal development ex-China.
Trevogrumab (REGN1033) may help with the muscle loss issues, where researchers have been posting evidence of significantly reduced loss of lean mass. And there’s a triplet with garetosmab for obesity, though safety issues have clouded expectations.
Once again, though, Regeneron finds itself competing in a hot, crowded field dominated by market leaders. Coming out on top will prove a tremendous challenge.
Regeneron has had success with its C5 siRNA drug cemdisiran, reporting positive Phase 3 data for generalized myasthenia gravis as it steered toward an FDA filing. Jefferies, for one, has estimated sales of a billion dollars. But it’s been flying under the radar until recently at Regeneron, which rarely happens at a biotech led by such bullish players.
15. Gilead: Time to bury the dead and move ahead
The scoop:
Gilead’s MO on the R&D side has remained largely unchanged during Daniel O’Day’s seven-year tenure as CEO. It buys up intriguing new drugs from cutting-edge biotechs and watches as the experiments gradually fail in the clinic. But each year it’s saved by a growing, dominant HIV business.
Biktarvy, its 3-in-1 daily tablet for HIV, provides about half of Gilead’s revenue. The R&D group has a deep understanding of HIV and the maintenance therapies needed to keep it in check, developing regimens that are easier on patients while keeping the big bucks rolling in. Yes, there have been failures in HIV — notably in GS-1720 and and GS-4182 recently — but its dominance in HIV shows no signs of deterioration.
You don’t have to look long at the deals Gilead has done to see the missteps in cancer. Immunomedics is a prime example. The recent Q4 and 2025 review contains no reference to TIGIT, once a hot field in immunotherapy that “seems doomed,” in the words of analyst Tim Anderson. And does anyone remember magrolimab, the anti-CD47 “don’t eat me” antibody?
O’Day, though, isn’t throwing in the towel. On the contrary. Gilead recently decided to go all in on anito-cel, buying out its partner Arcellx in a deal worth $7.8 billion. It believes the CAR-T can outperform Carvykti, the reigning champ in CAR-T, and help revive its commercial work in a field that has been flagging with its aging Kite originals.
This is no long-term wager. The company is looking for a green light from the FDA just before Christmas.
Then there was the $3.15 billion upfront pact to buy out an ADC from Tubulis.
Gilead spent $1.68 billion in cash to bag Ouro, gaining the T cell engager gamgertamig, a BCMAxCD3 candidate now in Phase 1/2.
In biotech, sometimes you have to bury the dead (programs) and move ahead. O’Day’s clearly decided that the time has come for Gilead. Its durable success in HIV gives him the means to do just that.
临床2期临床3期临床结果上市批准并购
2026-04-30
·大毒师
以下项目共同展现了 ADPKD 治疗领域从单一通路激素控制向多元化、机制驱动型疗法的明显转变,其中 HRS 9057 代表了新兴的细胞内小分子药物类别,RNA 疗法着眼于基因调控,而生物制剂和蛋白质矫正剂则拓展了精准治疗的范围。
ADPKD囊肿发病机制及相关治疗。图示为ADPKD发病机制中涉及的主要分子和细胞机制,包括:PC1/PC2功能丧失和钙稳态紊乱;影响纤毛形成的鞭毛内转运(IFT)复合物突变;JAK/STAT、NF-κB、Wnt/β-catenin和mTOR信号通路改变;Ras/MAPK和PI3K/Akt异常激活;cAMP信号通路失调;PC1错误折叠导致的内质网应激;以及线粒体功能障碍伴糖酵解和乳酸生成改变。图示的药物干预措施包括托伐普坦(V2受体阻滞剂)、二甲双胍(AMPK激活剂)、雷帕霉素/依维莫司(mTOR抑制剂)以及PDE和钙调磷酸酶通路调节剂。图中底部展示了新兴的基因疗法(CRISPR/Cas9、AAV介导的递送、反义寡核苷酸)。实线红线表示活跃的信号通路,虚线红线表示被破坏或抑制的通路。[Kolísková P (2024) Primary cilia-associated signalling in squamous cell carcinoma of head and neck region. Front. Oncol. 14:1413255. doi: 10.3389/fonc.2024.1413255]
全球 ADPKD 治疗方案开发机制集群图
🌍JMKX003142(济民可信,中国)
JMKX003142 是一种首创的小分子抑制剂,靶向细胞内囊肿增殖通路。临床前研究表明,该化合物能剂量依赖性地抑制肾上皮细胞增殖,且无钩状毒性。目前,该化合物正在中国开展一项针对快速进展型常染色体显性多囊肾病 (ADPKD) 患者的 II 期随机、双盲、安慰剂对照临床试验,同时正在进行 I 期 QTc 间期安全性研究。
🌍HRS 9057(成都盛迪药业/江苏恒瑞,中国)
HRS 9057是由江苏恒瑞旗下子公司成都盛迪药业研发的一种中国原创的I类创新小分子药物。该药旨在作为一种疾病修饰疗法用于治疗常染色体显性多囊肾病(ADPKD),其作用机制为在细胞内抑制肾上皮细胞增殖和囊肿形成。与血管加压素拮抗剂不同,HRS 9057并非主要通过激素通路发挥作用,而是靶向驱动囊肿形成的细胞内在信号网络。该化合物于2023年底获得国家药品监督管理局(NMPA)批准,进入ADPKD的临床试验,目前在中国处于早期I/II期临床开发阶段,主要研究重点是安全性、耐受性和探索性疗效。
🌍Tamibarotene (RN 014) –日本Rege Nephro 公司
塔米巴罗汀由日本 Rege Nephro 公司研发,是一种基于诱导多能干细胞 (iPSC) 衍生疾病模型而开发的视黄酸受体 (RAR) 激动剂,用于治疗常染色体显性多囊肾病 (ADPKD)。其作用机制在于调节抑制囊肿形成和上皮细胞去分化的转录程序。该药物为口服给药,是一种非激素、非 cAMP 直接作用的治疗方法。塔米巴罗汀已在日本完成 II 期临床试验的受试者招募,该试验设有安慰剂对照,是目前处于临床开发阶段的、由日本主导的最先进的 ADPKD 创新项目之一。
🌍Farabursen (RGLS8429) – Regulus / 诺华
Farabursen 是一种靶向 miR-17 的反义寡核苷酸,最初由 Regulus Therapeutics 开发,后被诺华收购并继续推进研发。该药物旨在通过解除 miR-17 介导的抑制作用来恢复 PKD1 和 PKD2 的表达,从而从遗传信号核心治疗常染色体显性多囊肾病 (ADPKD)。Farabursen 在 1b 期临床研究中展现出令人鼓舞的生物标志物和肾脏体积改善效果,目前正在包括美国以外地区在内的多个国家开展III 期临床开发。它是目前最成熟的基于 RNA 的 ADPKD 疗法之一。
🌍VX 407 – Vertex Pharmaceuticals
VX 407 是一种蛋白质校正小分子,旨在改善特定 PKD1 突变亚群中缺陷多囊蛋白 1 (PC1) 的折叠和转运。由Vertex公司开发的VX 407项目采用精准医疗策略,针对那些基因突变导致PC1蛋白错误折叠但可能可恢复的患者。VX 407目前正在全球范围内开展II期临床试验,具有高度的突变选择性,这意味着它未来将主要用于基因分层的ADPKD患者群体,而非普遍使用。
🌍艾伯维/卡利科抗妊娠相关血浆蛋白A抗体(ABBV CLS 628)
ABBV CLS 628 是一种靶向妊娠相关血浆蛋白A(PAPP A)的单克隆抗体,由艾伯维和卡利科生命科学公司联合开发。PAPP A 调节胰岛素样生长因子信号通路,从而促进囊肿扩张。该生物制剂代表了一种非小分子药物治疗常染色体显性多囊肾病(ADPKD)的方法,目前正在国际临床试验中心进行II期临床开发。它反映了人们对肾小管特异性机制之外的生长因子调节机制日益增长的兴趣。
🌍SGLT2抑制剂(达格列净、恩格列净——学术界和产业界项目)
SGLT2抑制剂(如达格列净和恩格列净)最初获批用于治疗糖尿病和慢性肾病(CKD),目前主要在欧洲和日本进行ADPKD的临床试验。它们的作用机制是间接的,涉及代谢、血液动力学和抗炎作用,而非直接抑制囊肿。目前正在进行多项 II 期和 III 期学术试验,通常设有安慰剂或标准治疗对照组,其中包括将 SGLT2 抑制剂与托伐普坦联合使用的研究。
🌍利昔伐坦 (Lixivaptan) – Centessa Pharmaceuticals
利昔伐坦是一种新一代血管加压素 V2 受体拮抗剂,旨在保留托伐普坦的促囊肿作用,同时降低肝毒性风险。目前,利昔伐坦正处于全球(包括美国以外地区)III 期临床试验的后期阶段,其作用机制并非改变,而是一种优化。
[Clerici S, Boletta A. Polycystic kidney disease Trends in Molecular Medicine, 2026; 32, 313-314]
🌍全球 ADPKD 治疗研发管线
药物公司地区阶段核心机制新颖性×成熟度关键状态说明主要网址链接托伐普坦(OPC 41061)大冢制药美国、欧盟和日本批准V2受体拮抗剂6/10目前唯一获批的疾病修饰疗法(DMT),但其疗效受限于利尿和肝毒性https://pkdcure.org/research/pipeline利昔伐坦
Centessa
美国和欧盟
III 期
新一代 V2 受体拮抗剂(对肝脏更安全)
7/10
正处于已验证通路的后期优化阶段
https://www.abnewswire.com/pressreleases/autosomal-dominant-polycystic-kidney-disease-pipeline-drugs-report-2025-innovative-treatments-drug-candidates-clinical-trials-and-market-trends_767131.htmlABBV‑CLS‑628AbbVie / Calico美国和欧盟II 期抗 PAPP A 单克隆抗体8/10FDA 快速通道;生长因子生物学https://clinicaltrials.gov/study/NCT06902558Farabursen (RGLS8429)
Regulus → Novartis
美国和欧盟
II 期
anti‑miR‑17 (RNA therapy)
9/10
恢复PKD1/2表达;强有力的生物标志物数据
https://pkdcure.org/research/pipeline/VX‑407 (AGLOW)Vertex美国和欧盟II 期PC1蛋白校正剂8/10针对PKD1变异体的精准治疗https://clinicalstudies.pkdcure.org/clinical-studies/HRS-9057
恒瑞/盛迪
中国
I/II期
细胞内囊肿生长途径抑制剂
7/10
首创非激素小分子
https://bydrug.pharmcube.com/news/detail/bff0f912050a13fecce02c651728024e他米巴罗汀 (RN 014)Rege Nephro日本II期视黄酸受体 (RAR) 激动剂7/10诱导多能干细胞发现;转录调控https://clinicaltrials.gov/study/NCT06289998恩格列净(ADPKD)
汉诺威医学院
欧盟、美国
III 期
SGLT2 抑制剂(代谢)
6/10
慢性肾病药物再利用
https://clinicaltrials.eu/trial/study-of-empagliflozin-for-patients-with-autosomal-dominant-polycystic-kidney-disease/达格列净——STOP PKD科隆大学欧盟II 期SGLT2 抑制剂5/10学术性概念验证http://pkdinternational.org/studiesDAPA Tolvaptan 组合
庆应大学
日本
II 期
SGLT2 + V2RA
6 / 10
组合策略
https://rctportal.mhlw.go.jp/en/detail?trial_id=UMIN000046275兰瑞肽(DIPAK)欧盟学术研究欧盟III期生长抑素类似物5/10肾脏获益有限https://www.ajkd.org/article/S0272-6386%2824%2901032-1/fulltext奥曲肽LAR
学术
欧盟
II/III期
生长抑素类似物
5/10
长效靶向抑制剂
二甲双胍美国国立卫生研究院资助的学术研究美国、欧盟II期AMPK激活4/10代谢再利用方法https://pmc.ncbi.nlm.nih.gov/articles/PMC6010317/依维莫司(Everolimus)
诺华
欧盟
II期
mTOR抑制剂
3/10
长期疗效不佳
博舒替尼(Bosutinib)辉瑞美国、欧盟II期Src激酶抑制剂3/10毒性限制替塞伐替尼(Tesevatinib,KD019)
Kadmon
美国、欧盟
II期
Src/EGFR抑制剂
3/10
已停止研发
雷公藤内酯醇学术中国II 期抗增殖药物4/10区域特异性https://pmc.ncbi.nlm.nih.gov/articles/PMC5123007/AZD1613(PIONEER PKD)
阿斯利康
美国、欧盟
I 期
新型小分子
4/10
早期安全性/药代动力学
https://clinicalstudies.pkdcure.org/clinical-studies/GLP-1 RA美国学术界美国II 期代谢/抗炎4/10探索性用途再利用RGLS4326
Regulus
美国
I 期
第一代抗 miRNA 药物
4 / 10
Farabursen 的前体
https://pkdcure.org/research/pipeline/JMKX003142济民可信中国II 期细胞内囊肿增殖通路抑制剂7/10中国研发的 BIC 潜力;非激素类小分子
2026-04-18
·药时代
合作机会来了!
药时代BD团队现有优质项目:用于治疗MASH和IPF,口服、每日一次,具有三重作用的 11β-HSD1抑制剂。MASH适应症已完成IIa期临床,具有非常积极的顶线结果和良好的安全性,在三重指标(脂肪变性、炎症、纤维化)上有均衡改善。IPF适应症处于IND ready阶段,抗纤维化的效果已经在两个模型中观察到。备向PF-ILD、RIPF以及更广泛的纤维化肺病扩展的潜力。
了解详情,请点击这里:
用于治疗MASH和IPF,具有三重作用的口服FIC 11β-HSD1抑制剂 | 药时代BD项目
请感兴趣的公司立即联系:药时代BD团队
BD@drugtimes.cn
备注项目代号:DT-20260325-122
自特朗普第二任期开启以来,美国政坛风向骤变,生物医药行业首当其冲。美国国立卫生研究院(NIH)经费遭遇大幅削减;部分原本在与监管机构沟通后有望获批的FDA上市申请,最终却等来了完全回应函(CRL);而层出不穷的关税政策,更持续向药企施加压力。面对这场“政策风暴”,企业不得不重新审视研发预算,调整应对之策。
数据也印证了这一趋势:2025年研发投入排名前十的企业中,多数已着手削减开支。BMS、强生、辉瑞、默沙东、艾伯维和罗氏的研发投入均较2024年有所下降,部分公司降幅高达29%。少数企业虽仍保持增长,但增幅已远不及往年。
数据来源:fiercebiotech
01
默沙东
默沙东表示,商务开发活动收缩是2025年研发预算下降12%的主因,但凭借充足投入,仍稳居榜首。
2024年,默沙东共完成四笔单笔超5亿美元的交易,大幅推高了当年研发预算;而2025年,公司最大一笔商务开发支出仅为向LaNova Medicines支付3亿美元,用于完成PD-1/VEGF双抗MK-2010的技术转移。
公司核心研发机构:默沙东实验室全年研发支出达到108亿美元,同比增长约7亿美元,延续了自2015年以来持续增长的投入趋势。
2025年,默沙东研发管线收获了多项Ⅲ期临床数据,包括口服PCSK9抑制剂enlicitide decanoate以及HIV疗法组合doravirine-islatravir。2026年,公司计划公布更多关键数据,涵盖用于溃疡性结肠炎的tulisokibart、用于糖尿病性黄斑水肿的MK-3000,以及GLP-1相关候选药物efinopegdutide等。
在今年1月的摩根大通医疗健康大会上,默沙东CEO Robert Davis表示,截至2030年代中期,公司将迎来价值700亿美元的年度商业化机遇。
02
罗氏
肿瘤领域仍是罗氏研发投入最高的板块,公司明确将KRAS抑制剂divarasib列为重点推进项目之一,计划尽快提交其用于非小细胞肺癌的上市申请。与此同时,口服选择性雌激素受体降解剂(SERD)giredestrant在2025年取得关键Ⅲ期临床研究进展,已就HR阳性、HER2阴性乳腺癌的二线治疗适应症提交监管申请,并同步推进用于早期乳腺癌辅助治疗的注册研究。
此外,罗氏在2025年加速布局肥胖症领域,宣布了跻身全球肥胖症药企前三的宏伟目标,计划2030年前推动GLP-1/GIP双受体激动剂CT-388上市。
免疫学管线也在去年获得追加资金,推动肠道疾病候选药阿非奇单抗(afimkibart)开展多项Ⅲ期临床,计划2027年提交上市申请。
尽管部分领域投入增加,但罗氏表示,相关增幅已被基因治疗子公司Spark Therapeutics重组、肿瘤数据平台Flatiron Health资源整合带来的成本削减完全抵消。
03
强生
研发投入收缩,或与强生大举收购外部创新资产有关。2025年初,强生以146亿美元收购中枢神经系统药企Intra-Cellular Therapies,成为全年最大并购交易;11月再以30亿美元收购Halda Therapeutics,获得其调节诱导接近靶向嵌合体研发平台及前列腺癌候选项目,为肿瘤领域的创新布局提供新的技术路径。
在临床研发方面,2025年公司多项创新疗法取得重要进展。与 Protagonist 合作的斑块状银屑病候选药icotrokinra Ⅲ期疗效优于BMS的 Sotyktu;新一代双靶点CAR-T细胞疗法初步数据显示,10 例经多线治疗患者客观缓解率达 100%。
2026年初,强生与特朗普政府签署“最惠国”药价协议,并承诺在美国新建生产基地。
04
阿斯利康
2025年,阿斯利康研发投入持续增长,约占公司总营收的四分之一。公司多款潜在重磅药物取得临床成功,包括一款乳腺癌口服药、一款预计销售额达数十亿美元的自身免疫病药物,以及一项价值8亿美元的罕见病布局。
阿斯利康持续加码研发管线:承诺未来五年在华投资150亿美元;与JCR制药达成8.25亿美元罕见病合作;与石药集团签署协议,并于2025年6月合作后,今年初再敲定一项185亿美元的肥胖症重磅合作。
CEO Pascal Soriot在年末声明中着眼未来:“我们有超过100项Ⅲ期临床正在推进,其中大量变革性技术的临床试验持续扩容,有望彻底改变患者结局,并支撑公司2030年后的长期增长。”
05
礼来
2025年,礼来成为首家市值突破万亿美元的制药企业,重磅药替尔泊肽销售额登顶“药王”宝座。凭借替尔泊肽的热销与充裕现金流,公司研发投入同比大增21.4%。
礼来正通过数字化与人工智能技术强化药物发现能力:推出制药行业规模最大的超级计算机LillyPod,深化与英伟达的合作;建立Lilly Gateway Labs孵化网络,计划在韩国建立新的创新中心;并将合作延伸至Insilico Medicine、晶泰科技子公司Ailux,借助人工智能开发新药,以进一步推动全球创新药研发生态建设。
同时礼来布局基因治疗领域:以约7400万美元收购Adverum,获得一款黄斑变性基因治疗候选药;获得 MeiraGTx 一款候选药授权,该药曾帮助 11 名重症视网膜疾病患儿恢复视力。
礼来最紧迫的研发任务,是延续替尔泊肽在肥胖与糖尿病领域的成功。2026年4月1日,礼来宣布FDA批准其口服GLP-1受体激动剂orforglipron上市。该药位列Evaluate预测的2026年最受期待上市药物第二位,仅次于诺和诺德的CagriSema。
06
诺华
诺华年报显示,研发投入增长部分源于对近期收购资产的追加投资。继2024年以29亿美元收购MorphoSys后,诺华再斥资120亿美元收购晚期肌营养不良症生物科技公司Avidity Biosciences。诺华CEO Vas Narasimhan表示,此举旨在推动诺华成为神经肌肉疾病领域的领军企业。
2025年诺华并购动作密集:14亿美元收购心血管药企Tourmaline;17亿美元收购肾病专科药企Regulus Therapeutics;2月以9.25亿美元收购此前分拆的Anthos Therapeutics;引进Sironax的血脑屏障穿透技术。
在研发合作方面,诺华与BioArctic合作研发神经退行性疾病药物,从Arrowhead引进帕金森病候选药,与ProFound Therapeutics达成心血管领域的发现合作,并与中国企业Argo Biopharmaceutical签署siRNA协议,承诺最高50亿美元的投入。
今年,诺华有望收获过往并购的成果:未来数月将公布与Ionis合作开发的降低Lp(a)心血管风险候选疗法的关键数据;同时准备今年初向FDA提交ianalumab的上市申请,该药在原发性免疫血小板减少症的治疗中取得了重大突破。
07
辉瑞
辉瑞在2025年继续推进其创新研发战略,重点布局代谢疾病治疗领域。其中最引人关注的是完成对Metsera的收购,以100 亿美元将新一代肥胖症药企纳入麾下。此后,辉瑞再以最高19亿美元从复星医药子公司获得一款 GLP-1 候选药授权。
在管线管理方面,辉瑞在2025年持续对研发项目进行结构性优化,将资源集中于更具潜力的创新疗法。同时,公司在基因治疗领域也进行了战略调整,在优化既有项目组合的同时,从Beam Therapeutics引入新的基因编辑候选项目,以持续拓展其遗传疾病治疗管线。
在大型药企CEO中,辉瑞 CEO Albert Bourla曾被视为与特朗普关系最为密切的高管,但近期态度转变,公开抨击美国卫生部长小肯尼迪的反疫苗立场,并称Vinay Prasad为“问题人物”,指责其无视专业职员意见。
08
BMS
2025年,BMS收紧研发开支,研发预算削减11%,通过提升研发效率与降低资产减值支出实现成本节约。
公司临床前新分子实体研发投入为13亿美元,低于此前三年每年超过15亿美元的水平;但Ⅰ/Ⅱ/Ⅲ期临床投入增长,推动临床研发费用升至近46亿美元。临床与临床前总投入减少1.26亿美元,剩余12亿美元的降幅源于减值支出与并购相关结算。CEO Christopher Boerner叫停了alnuctamab等项目,目标是在20个月内削减15亿美元成本。
在研发管线方面,BMS正推进多项关键候选药物进入重要临床阶段。例如,用于多发性骨髓瘤的iberdomide在临床研究中取得进展,并通过研究方案调整推动相关试验继续推进;另一款多发性骨髓瘤候选药物mezigdomide在Ⅲ期临床研究中取得积极结果。同时,公司还在推进与强生合作开发的抗凝药物milvexian以及用于特发性肺纤维化治疗的admilparant等项目。通过持续优化后期临床项目,BMS正为未来创新疗法的推出奠定基础。
09
艾伯维
艾伯维在2025年继续保持较高水平的研发投入,并通过并购与合作持续扩展创新药物管线。公司全年研发预算约为91亿美元,资源重点投向神经精神疾病、细胞疗法以及代谢疾病等多个前沿治疗领域。
艾伯维CEO Rob Michael表示,并购仍是公司突破自免药物Skyrizi与Rinvoq、实现增长的核心战略。
2025年,艾伯维开展多项并购:以12亿美元获得Gilgamesh Pharmaceuticals的抑郁症候选药bretisilocin;以21亿美元收购Capstan Therapeutics,切入体内CAR-T这一热门赛道;同时布局T细胞衔接器、分子胶降解剂、新一代肥胖症药物等领域。与丹麦药企Gubra合作的胰淀素类似物近期公布数据,单剂量给药12周减重幅度接近10%。
在侧重外部创新的同时,艾伯维也优化了研发组织结构:11月终止与Calico Labs的长期合作,约百名相关员工被裁。
10
赛诺菲
管线接连失利加剧了管理层压力,CEO Paul Hudson曾表示2025年是“坎坷之路”。抗OX40配体抗体amlitelimab在哮喘Ⅱ期临床中失败;口服TNF抑制剂balinatunfib在银屑病中期研究中未达终点;与再生元合作的IL-33抗体itepekimab在慢阻肺的两项Ⅲ期研究中有一项失利。
这与公司年初目标相去甚远。Paul Hudson曾表示,2024年公司向“聚焦型科研驱动药企”转型取得重大进展,出售消费者健康业务Opella正是为了“优先聚焦研发”。然而,随着重磅药物Dupixent在2031年左右专利到期,营收缺口亟待填补,赛诺菲董事会失去耐心,于2026年2月解雇Paul Hudson,聘任德国默克的CEO Belén Garijo接任。董事会明确要求Garijo:提升研发效率、完善治理、强化创新能力。
参考资料:
1.https://www.fiercebiotech.com/special-reports/top-10-pharma-rd-budgets-2025
2.其他公开资料
封面图来源:即梦AI
小肯尼迪:把“灰产洗白”!
2026-04-16
上市仅两周即被FDA要求补数据!礼来口服GLP-1面临考验
2026-04-15
拒绝320亿美元收购?那就对了!
2026-04-14
沙利文发布《2026年全球过敏原特异性免疫治疗行业蓝皮书》(内附全文获取方式)
2026-04-13
临床试验获重要进展!北京天坛医院完成干细胞移植,改写帕金森治疗格局
2026-04-11
版权声明/免责声明
本文为原创文章。
本文仅作信息交流之目的,不提供任何商用、医用、投资用建议。
文中图片、视频、字体、音乐等素材或为药时代购买的授权正版作品,或来自微信公共图片库,或取自公司官网/网络,部分素材根据CC0协议使用,版权归拥有者,药时代尽力注明来源。
如有任何问题,请与我们联系。
衷心感谢!
药时代官方网站:www.drugtimes.cn
联系方式:
电话:13651980212
微信:27674131
邮箱:contact@drugtimes.cn
点击查看更多精彩内容!
临床3期临床申请临床2期
100 项与 Regulus Therapeutics, Inc. 相关的药物交易
登录后查看更多信息
100 项与 Regulus Therapeutics, Inc. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年06月06日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
临床前
3
1
临床1期
其他
11
登录后查看更多信息
当前项目
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或

转化医学
使用我们的转化医学数据加速您的研究。
登录
或

营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或

科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或

投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或

融资
发掘融资趋势以验证和推进您的投资机会。
登录
或

生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
