预约演示
更新于:2026-04-25

Shanxi Medical University
更新于:2026-04-25
概览
标签
肿瘤
其他疾病
神经系统疾病
小分子化药
化学药
合成多肽
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 小分子化药 | 13 |
| 化学药 | 6 |
| 合成多肽 | 3 |
| 间充质干细胞疗法 | 2 |
| 未知 | 1 |
关联
28
项与 山西医科大学 相关的药物作用机制 LCN2 inhibitors [+2] |
原研机构 |
最高研发阶段申请上市 |
首次获批国家/地区- |
首次获批日期- |
89
项与 山西医科大学 相关的临床试验NCT07539545
A Comparative Study of Minimal Accessory-Incision-Assisted Endoscopic Breast-Conserving Surgery With Minimal Auxiliary Incisions Versus Conventional Open Breast-Conserving Surgery: A National Multicenter, Open-Label, Randomized Controlled Trial (MECO-BCS)
This study is a multicenter, open-label, randomized controlled trial. The study aims to evaluate differences in operative efficiency (e.g., operative time), economic effect, surgical safety (e.g., surgical complication rates), postoperative aesthetics (e.g., BREAST-Q scores, Harris scores, SCAR-Q scores and Ueda scores), and oncological safety (e.g., margin status, no local recurrence survival) between patients undergoing M-E-BCS and patients undergoing C-O-BCS.
开始日期2026-04-01 |
申办/合作机构  四川大学华西医院(四川省国际医院) 四川大学华西医院(四川省国际医院) [+4] |
NCT07276919
Perioperative Tamsulosin for Treating Voiding Dysfunction Following Prostate Biopsy: A Multicenter, Randomized, Controlled, Open-Label Study
The goal of this clinical trial is to learn if the drug Tamsulosin Hydrochloride works to treat urination difficulties after a prostate biopsy in men.The main questions it aims to answer are:
* Does Tamsulosin Hydrochloride lower the rate of Acute Urinary Retention (inability to urinate) after a prostate biopsy?
* Does Tamsulosin Hydrochloride improve participants' urination symptoms, quality of life, urine flow rate, and post-void residual urine volume?
Researchers will compare the drug group to a control group (receiving no preventive medication) to see if Tamsulosin Hydrochloride works to prevent urination problems.
Participants will:
* Be randomly assigned to either take Tamsulosin Hydrochloride or receive no preventive medication.
* If in the drug group, take one capsule of Tamsulosin Hydrochloride every night, starting 3 days before the biopsy and continuing until 6 days after the biopsy (10 doses total).
* Undergo a standard prostate biopsy procedure (either through the rectum or perineum).
* Return to the clinic 7 days after the biopsy for a follow-up checkup, which will include questions about their symptoms (IPSS and QoL scores) and tests to measure urine flow and bladder emptying.
* Does Tamsulosin Hydrochloride lower the rate of Acute Urinary Retention (inability to urinate) after a prostate biopsy?
* Does Tamsulosin Hydrochloride improve participants' urination symptoms, quality of life, urine flow rate, and post-void residual urine volume?
Researchers will compare the drug group to a control group (receiving no preventive medication) to see if Tamsulosin Hydrochloride works to prevent urination problems.
Participants will:
* Be randomly assigned to either take Tamsulosin Hydrochloride or receive no preventive medication.
* If in the drug group, take one capsule of Tamsulosin Hydrochloride every night, starting 3 days before the biopsy and continuing until 6 days after the biopsy (10 doses total).
* Undergo a standard prostate biopsy procedure (either through the rectum or perineum).
* Return to the clinic 7 days after the biopsy for a follow-up checkup, which will include questions about their symptoms (IPSS and QoL scores) and tests to measure urine flow and bladder emptying.
开始日期2026-02-01 |
申办/合作机构 北京大学第一医院 [+1] |
ChiCTR2500111605
The Impact of Health Education Based on Information Framing Effect on Self-Management Behavior of Artemisia Pollen Allergic Rhinitis Patients
开始日期2025-11-06 |
申办/合作机构  山西医科大学 山西医科大学 [+1] |
100 项与 山西医科大学 相关的临床结果
登录后查看更多信息
0 项与 山西医科大学 相关的专利(医药)
登录后查看更多信息
6,409
项与 山西医科大学 相关的文献(医药)2026-12-31·Human Vaccines & Immunotherapeutics
HPV vaccine awareness and uptake among women attending cervical screening in health-resource-limited areas of China: A multicenter cross-sectional study
Article
作者: Hu, Shangyin ; Shi, Jingyi ; Wang, Junling ; Feng, Ruimei ; Duan, Rufei ; Jia, Xinhua ; Zhang, Tai ; Yang, Chunxia ; Zhang, Hongyun ; Qiao, Youlin ; Li, Zhifang ; Zhao, Yuqian ; Da, Xi’ao
Evidence shows HPV vaccination reduces infection, precancer and cervical cancer, yet coverage in health-resource-limited of China remains uncertain. We assessed awareness, uptake and correlates among women attending cervical screening, and examined associations with screening outcomes. We conducted a cross-sectional study in eight county sites in 2023-2024 among women aged 35-64 y. A standardized questionnaire captured sociodemographic factors, awareness and vaccination. Cervical samples were tested for hrHPV. Outcomes were awareness, vaccination, hrHPV, HPV16/18 and CIN2+. Associations were estimated using modified Poisson models with site fixed effects and HC3 robust errors. Adjusted prevalence ratios (aPRs) and covariate-standardized marginal estimates were reported. We included 93,027 unique participants. Awareness was 45.15% and vaccination 6.73%. hrHPV prevalence was 11.38% and CIN2+ detection was 0.70%. In 2024 versus 2023, awareness was lower (40.66% vs 51.34%) while vaccination was higher (7.61% vs 5.53%; aPR 1.25, 95% CI 1.18-1.32). Awareness and uptake declined with age; coverage was 23.23% at ages 35-39 and 0.29% at ages 60-64. Urban residence and higher education were associated with uptake (urban aPR 1.19, 95% CI 1.11-1.27; bachelor's or higher aPR 1.73, 95% CI 1.58-1.90). The age-by-year interaction was significant, with standardized gains concentrated at ages 35-49. Vaccination was associated with lower HPV16/18 infection (aPR 0.66, 95% CI 0.52-0.85) but not with overall hrHPV or CIN2+. HPV vaccine awareness and uptake were low among women aged 35-64 y in health-resource-limited areas, with strong age, educational and urban-rural gradients and marked site heterogeneity. Uptake increased in 2024, mainly at ages 35-49, and vaccination was associated with a lower prevalence of HPV16/18 infection.
2026-07-01·EXPERIMENTAL NEUROLOGY
Sulforaphane improved cognitive behavior in APP/PS1 mice via promoting structural and functional synaptic plasticity in the hippocampus
Article
作者: Wang, Ying ; Shang, Zeyu ; Li, Xinxin ; Wang, Zhaojun ; Hu, Yanjie ; Gao, Qichao ; Cai, Hongyan ; Wu, Shengnan ; Wu, Meina
Cognitive dysfunction is the main clinical feature of Alzheimer's disease (AD), and oxidative stress is considered a critical contributor to AD pathogenesis. Sulforaphane (SFN), an aliphatic isothiocyanate predominantly derived from cruciferous vegetables, has been reported to exert antioxidant and neuroprotective effects; however, its impact on synaptic plasticity and the underlying electrophysiological mechanisms in AD remain unclear. In this study, ten-month-old male APPswe/PS1dE9 double transgenic (APP/PS1) mice and wild-type (WT) littermates were randomized to four groups (WT + saline, WT + SFN, APP/PS1 + saline, and APP/PS1 + SFN). SFN was administered at 10 mg kg-1via intraperitoneal injection for 30 d in vivo and 1 μM concentration in vitro. Then, we investigated whether SFN improves cognition by restoring hippocampal synaptic plasticity in APP/PS1 mice. It was suggested that SFN administration significantly attenuated Aβ₁₋₄₂-induced oxidative damage in vitro and improved spatial learning and reference memory deficits in APP/PS1 mice. Bioinformatics analysis suggested that SFN modulated synapse-related pathways associated with AD. Consistent with these findings, electrophysiological recordings demonstrated that SFN alleviated hippocampal long-term potentiation (LTP) impairment. Moreover, SFN increased the expression of synaptic proteins PSD-95 and synaptophysin and enhanced dendritic complexity and dendritic spine density in the hippocampal CA1 region, as assessed by immunoblotting and Golgi staining. Together, these results indicated that SFN ameliorated cognitive deficits in AD mice by alleviating LTP inhibition and restoring synaptic structural integrity, supporting a role for synaptic plasticity in the neuroprotective effects of SFN.
2026-07-01·COLLOIDS AND SURFACES B-BIOINTERFACES
Room-temperature synthesis of fluorine-anion engineered cerium oxyfluoride nanozymes via valence-regulation strategy for prompt uric acid biosensing
Article
作者: Li, Min ; Zhang, Wenjun ; Wang, Yuheng ; Ruan, Xuehua ; Zhang, Yumin ; He, Gaohong ; Huang, Yixiang ; Xu, Zixuan
The catalytic activity of Ce-based nanozymes is significantly positive-correlated with the Ce3 + ratio. In this work, Ce3+ content in cerium oxyfluoride nanozymes was efficiently and directly regulated with fluorine-induced charge compensation between fluorine and oxygen, and stabilized by the electronegativity-driven inductive effect, through the facile and room-temperature synthesis for the enzyme /H2O2-free and highly sensitive UA detection. Remarkably, cerium oxyfluoride exhibited Vmax of 196 × 10-8 M·s-1 and Km of 0.347 mM, indicating the superior oxidase-like activity. Thereby, ascribed to the antioxidant property of uric acid (UA), UA was effectively and promptly detected, i.e., a wide linear range of 0.3-1.2 mM, and ultra-low detection limit of 4.5 μM, as well as the significant anti-interference against the common urinary components. Prospectively, the electron-density reforming strategy through the strong electronegative fluorine introduction, extremely regulated the valence of cerium in cerium oxyfluoride to enhance the catalytic activity, via the synthetic route of mildness and facileness, fulfilling the effective real-time detection of UA.
101
项与 山西医科大学 相关的新闻(医药)2026-04-22
·百度百家
2026年医学考研季正呈现出一种奇特的“分裂图景”:一边是临床医学专硕的报名通道被考生挤得“水泄不通”,深圳大学临床医学专硕报考比从2025年的1:3.2飙升至1:4.8,武汉大学临床医学专硕统考名额缩减至119人但报考人数逆势增长23%,复录比达1:1.2;另一边则是学术学位(学硕)方向“门庭冷落”,南开大学临床医学学硕复试线从2025年的295分降至2026年的280分,山西医科大学基础医学学硕报录比从2025年的9:1跌至2026年的4:1,调剂名额占比超40%。
当绝大多数医学生基于“务实”考量涌向专硕这条看似安稳的轨道时,一个被忽视的提问正浮出水面:学硕这条路径真的意味着“前途黯淡”吗?或许,现实正在给出反差性答案——在特定赛道,学硕正悄然转化为“稀缺资产”。
解剖“热”与“冷”——现象背后的驱动逻辑
专硕之“热”:为何成为主流选择?
政策引力构成最直接的“终极诱惑”。临床医学专硕的“四证合一”模式——硕士学历证、学位证、执业医师资格证和住培合格证——将住院医师规范化培训(住培)与硕士教育深度融合,大幅缩短了临床准入周期。学生需完成33个月的科室轮转,直接参与临床诊疗,这种“边学边用”的模式使毕业生已具备独立处理常见病的能力。
市场明牌为专硕提供了清晰的就业路径。专硕毕业生因“四证合一”优势,在地市级三甲医院招聘中更受青睐。例如,资料显示徐州市中心医院2025年招聘中,75%的临床岗位明确要求“四证合一”背景。这种与医院规范化培训及编制岗位的直接挂钩,为考生提供了可见的确定性。
即时收益满足当下的现实压力。2025年数据显示,专硕首年住院医师月薪约1.2-1.8万元,相比学硕初期收入(约0.8-1.2万元)有明显优势。对希望尽快经济独立、减轻家庭负担的考生而言,专硕提供了更早接触临床、获得经济回报的路径。
学硕之“冷”:传统困境与普遍认知
培养周期长构成心理屏障。科研型学硕聚焦基础研究与技术创新,三年时光多在实验室度过,需完成细胞培养、动物模型构建、基因测序等科研训练,毕业要求至少1-2篇SCI论文。如果毕业后要从事临床工作,需要另外进行3年住院医师规范化培训,职业起步相对缓慢。
科研压力大形成过滤机制。学硕需要投入大量时间进行实验设计、数据分析和论文撰写,产出不确定性高,心理负荷重。与传统认知中医院临床岗位对学硕需求弱的印象叠加,使得学硕的就业前景看似模糊、狭窄。
就业模糊性加深了考生顾虑。在普遍认知中,学硕毕业生在临床就业市场上面临困境,出路似乎被限定在高校、科研院所或政策研究机构。这种刻板印象与专硕清晰的职业通道形成鲜明对比,进一步加剧了学硕的遇冷趋势。
重新定义“冷”与“热”——学硕被低估的价值赛道
赛道一:生物医药研发的“硬核需求”
产业升级正在重塑人才需求结构。中国生物医药行业正从“仿制跟随”向“自主创新”全面跃迁,国内企业在PD-1、ADC、小核酸等前沿技术方向加速布局。2025–2026年创新药研发管线、跨国授权合作及全球交易活跃度均显著提升,这对具备系统科研能力的人才形成持续强劲的需求拉动。
岗位匹配显示出学硕的独特优势。生物医药行业高端技术岗位核心需求旺盛,涉及创新药早期靶点发现、抗体药物工程、基因/细胞治疗、生物信息学与AI辅助药物设计等前沿方向。学硕生在这些领域的基础研究、临床前研发、转化医学等岗位具备独特优势——他们既懂医学又具备深度科研素养,这类复合型人才在市场中供不应求。
薪酬天花板可能更高。资料显示,创新药研发科学家、生物信息与AI辅助研发工程师等岗位,对具备科研背景、实验设计能力与跨团队协作经验者需求迫切。博士毕业后到手年收入可达35万元以上,且涨薪速度快,紧跟前沿技术者更易抓住行业风口。
赛道二:“医学+X”交叉学科的“未来风口”
国家战略牵引形成明确导向。2026年,国务院政府工作报告进一步提出“一体推进教育科技人才发展、建设国家交叉学科中心”。现阶段国家急需的高层次人才多集中于交叉学科领域,优化高校学科布局、推动学科交叉融合成为适应未来产业发展的必由之路。
交叉学科项目获得实质性支持。中南大学湘雅医学院2026年度研究生交叉创新培育项目设立“医工交叉”“医理交叉”“医文交叉”“医信交叉”四个申报方向,拟立项20项,重点项目资助3万元/项,一般项目资助1.5万元/项。这种资源倾斜为交叉学科研究提供了物质保障。
前沿领域应用展示广阔前景。智能医学工程专业——AI+医疗的超级复合人才——正在改写医生的定义。医疗AI企业推想科技的人事总监透露:“既懂CT影像解读又能优化算法的毕业生,我们愿意支付35万年薪起。”该专业就业宽度广阔,既可进入高端设备商担任产品工程师,也能在顶级三甲医院智能医疗中心担任技术医师,还能加入互联网医疗平台做算法专家。
赛道三:高端科研与教育机构的“储备力量”
高水平大学和科研平台对学术研究人才保持持续需求。北京协和医学院数据显示,学硕进入三甲医院科研岗的比例是专硕的2.3倍。尽管传统临床岗位对学硕需求有限,但在高校附属医院、科研院所的科研岗位招聘中学硕占据明显优势。
国家科研体系建设为学硕提供了成长空间。2026年2月27日中共中央政治局会议强调要“加快高水平科技自立自强”,这一政策导向为长期投入基础科研的人才创造了宏观环境。生物医学数据科学、精准医学等新兴专业正站上风口,华西医院近期以60万年薪招聘的临床数据分析总监,要求必须兼具医学统计知识和基因组学背景。
趋势与信号——政策与市场正在说什么?
政策风向:科技创新与学科交叉的双重支撑
教育部在2026年的政策导向中明确要求临床医学专硕占比提升至70%,学硕招生规模缩减30%,但重点向“卡脖子”领域(如基因编辑、AI医疗)倾斜。这种“有压有保”的策略表面上看压缩了学硕规模,实则优化了结构——将资源集中于前沿交叉学科。
国务院学位委员会、教育部印发的《研究生教育学科专业目录(2022年)》将交叉学科作为一个门类正式“入驻”,下设智能科学与技术、纳米科学与工程等一级学科。2026年国务院政府工作报告提出“建设国家交叉学科中心”,这一信号表明国家对交叉学科发展的重视程度。
市场信号:新兴产业崛起与人才需求结构变化
人工智能医疗公司正高薪争夺学硕人才。资料显示,自然语言处理方向的AI医疗岗位年薪可达40万元,具备医学与数据科学交叉能力的人才更具市场稀缺性。数字医疗技术的发展推动了“微生物工厂”生产复杂药物,大幅降低生产成本的同时,也创造了新的就业岗位。
药企研发岗位为学硕提供了另一条路径。生物医药行业人才需求集中在研发及临床岗位,且对AI+生物交叉复合型人才需求提升明显。高端研发岗位通常要求硕士以上学历,博士在基因编辑等领域更具优势,而学硕作为博士预备阶段,在这一链条中占据重要位置。
战略选择与行动指南——你适合走学硕之路吗?
自我评估:哪些学生更适合考虑学硕?
内心驱动是首要考量因素。学硕适合那些对探索疾病本质、机理研究有强烈好奇心与热情的学生,而不仅仅是对临床操作技能感兴趣。如果享受发现问题、设计实验、分析数据的过程,而非追求即时临床反馈,学硕可能是更合适的选择。
能力特质决定适应程度。学硕需要较强的逻辑思维、耐心和抗压能力,能够忍受实验室的寂寞和科研的不确定性。相比之下,专硕更适合动手能力强、喜欢与人打交道、能够承受高强度值班(日均接诊30-50人次)和夜班压力的学生。
职业愿景指向不同方向。如果志不在局限于单一临床操作,而是向往产业研发、前沿科技创新、高校科研或成为跨领域专家,学硕提供了更广阔的可能性。专硕则更适合坚定走临床医生道路、希望尽快经济独立、目标普通三甲医院就业的考生。
关键决策点:如何规划学硕之路?
导师选择成为成败关键。寻找研究方向前沿、产业资源丰富或学术声誉卓著的导师至关重要。导师风格(“放养”vs“手把手”)需要与自身特点匹配——自主性强的学生可能适合“放养”型导师,而需要更多指导的学生则应选择“手把手”型导师。
研究方向需要平衡科学价值与应用潜力。选择既有科学价值又有应用潜力的课题,关注AI辅助新药研发、智能诊断设备开发、高端生物医用材料等交叉学科方向。中南大学湘雅医学院的交叉创新培育项目提供了医工、医理、医文、医信四个方向的参考。
产业衔接应提前布局。在读期间通过实习、项目合作、学术会议主动接触产业界,构建可转移的技能组合(如数据分析、专利撰写、项目管理)。浙江大学智能医学工程专业的课程表上,Python编程与解剖学并列,机器学习与病理学同排,这种课程设计本身就体现了产业衔接的思路。
在喧嚣中聆听内心的声音
2026年医学考研的“冷热抉择”本质上是个体在短期现实利益与长期发展潜力之间、在已知赛道与未知蓝海之间的一次战略权衡。专硕代表了一条清晰、稳健的职业道路——3年获得“四证合一”,直接进入临床岗位;学硕则意味着拥抱不确定性,以更长时间的投入博取在创新领域更广阔的可能性。
医学教育正面临深度重构。教育部要求2026年临床医学专硕占比提升至70%,学硕招生规模缩减30%,但重点向“卡脖子”领域倾斜。这并非简单的数量增减,而是结构优化——将有限资源集中于最有价值的交叉学科方向。
没有绝对的正确选择,只有基于深刻自我认知与趋势判断的合适之选。当绝大多数人涌向专硕时,学硕这条路径可能正从“冷门”转变为“稀缺”;当AI医疗公司愿意为“既懂医学又懂算法”的人才支付35万年薪时,传统的“冷热”定义正在被重新书写。
在这场医学考研的“冷热抉择”中,你是愿意跟随人群挤上专硕的“独木桥”,还是敢于探索学硕这条看似冷清却可能通往未来的“新航道”?
2026-04-22
·有驾
2026年医学考研季正呈现出一种奇特的“分裂图景”:一边是临床医学专硕的报名通道被考生挤得“水泄不通”,深圳大学临床医学专硕报考比从2025年的1:3.2飙升至1:4.8,武汉大学临床医学专硕统考名额缩减至119人但报考人数逆势增长23%,复录比达1:1.2;另一边则是学术学位(学硕)方向“门庭冷落”,南开大学临床医学学硕复试线从2025年的295分降至2026年的280分,山西医科大学基础医学学硕报录比从2025年的9:1跌至2026年的4:1,调剂名额占比超40%。
当绝大多数医学生基于“务实”考量涌向专硕这条看似安稳的轨道时,一个被忽视的提问正浮出水面:学硕这条路径真的意味着“前途黯淡”吗?或许,现实正在给出反差性答案——在特定赛道,学硕正悄然转化为“稀缺资产”。
解剖“热”与“冷”——现象背后的驱动逻辑
专硕之“热”:为何成为主流选择?
政策引力构成最直接的“终极诱惑”。临床医学专硕的“四证合一”模式——硕士学历证、学位证、执业医师资格证和住培合格证——将住院医师规范化培训(住培)与硕士教育深度融合,大幅缩短了临床准入周期。学生需完成33个月的科室轮转,直接参与临床诊疗,这种“边学边用”的模式使毕业生已具备独立处理常见病的能力。
市场明牌为专硕提供了清晰的就业路径。专硕毕业生因“四证合一”优势,在地市级三甲医院招聘中更受青睐。例如,资料显示徐州市中心医院2025年招聘中,75%的临床岗位明确要求“四证合一”背景。这种与医院规范化培训及编制岗位的直接挂钩,为考生提供了可见的确定性。
即时收益满足当下的现实压力。2025年数据显示,专硕首年住院医师月薪约1.2-1.8万元,相比学硕初期收入(约0.8-1.2万元)有明显优势。对希望尽快经济独立、减轻家庭负担的考生而言,专硕提供了更早接触临床、获得经济回报的路径。
学硕之“冷”:传统困境与普遍认知
培养周期长构成心理屏障。科研型学硕聚焦基础研究与技术创新,三年时光多在实验室度过,需完成细胞培养、动物模型构建、基因测序等科研训练,毕业要求至少1-2篇SCI论文。如果毕业后要从事临床工作,需要另外进行3年住院医师规范化培训,职业起步相对缓慢。
科研压力大形成过滤机制。学硕需要投入大量时间进行实验设计、数据分析和论文撰写,产出不确定性高,心理负荷重。与传统认知中医院临床岗位对学硕需求弱的印象叠加,使得学硕的就业前景看似模糊、狭窄。
就业模糊性加深了考生顾虑。在普遍认知中,学硕毕业生在临床就业市场上面临困境,出路似乎被限定在高校、科研院所或政策研究机构。这种刻板印象与专硕清晰的职业通道形成鲜明对比,进一步加剧了学硕的遇冷趋势。
重新定义“冷”与“热”——学硕被低估的价值赛道
赛道一:生物医药研发的“硬核需求”
产业升级正在重塑人才需求结构。中国生物医药行业正从“仿制跟随”向“自主创新”全面跃迁,国内企业在PD-1、ADC、小核酸等前沿技术方向加速布局。2025–2026年创新药研发管线、跨国授权合作及全球交易活跃度均显著提升,这对具备系统科研能力的人才形成持续强劲的需求拉动。
岗位匹配显示出学硕的独特优势。生物医药行业高端技术岗位核心需求旺盛,涉及创新药早期靶点发现、抗体药物工程、基因/细胞治疗、生物信息学与AI辅助药物设计等前沿方向。学硕生在这些领域的基础研究、临床前研发、转化医学等岗位具备独特优势——他们既懂医学又具备深度科研素养,这类复合型人才在市场中供不应求。
薪酬天花板可能更高。资料显示,创新药研发科学家、生物信息与AI辅助研发工程师等岗位,对具备科研背景、实验设计能力与跨团队协作经验者需求迫切。博士毕业后到手年收入可达35万元以上,且涨薪速度快,紧跟前沿技术者更易抓住行业风口。
赛道二:“医学+X”交叉学科的“未来风口”
国家战略牵引形成明确导向。2026年,国务院政府工作报告进一步提出“一体推进教育科技人才发展、建设国家交叉学科中心”。现阶段国家急需的高层次人才多集中于交叉学科领域,优化高校学科布局、推动学科交叉融合成为适应未来产业发展的必由之路。
交叉学科项目获得实质性支持。中南大学湘雅医学院2026年度研究生交叉创新培育项目设立“医工交叉”“医理交叉”“医文交叉”“医信交叉”四个申报方向,拟立项20项,重点项目资助3万元/项,一般项目资助1.5万元/项。这种资源倾斜为交叉学科研究提供了物质保障。
前沿领域应用展示广阔前景。智能医学工程专业——AI+医疗的超级复合人才——正在改写医生的定义。医疗AI企业推想科技的人事总监透露:“既懂CT影像解读又能优化算法的毕业生,我们愿意支付35万年薪起。”该专业就业宽度广阔,既可进入高端设备商担任产品工程师,也能在顶级三甲医院智能医疗中心担任技术医师,还能加入互联网医疗平台做算法专家。
赛道三:高端科研与教育机构的“储备力量”
高水平大学和科研平台对学术研究人才保持持续需求。北京协和医学院数据显示,学硕进入三甲医院科研岗的比例是专硕的2.3倍。尽管传统临床岗位对学硕需求有限,但在高校附属医院、科研院所的科研岗位招聘中学硕占据明显优势。
国家科研体系建设为学硕提供了成长空间。2026年2月27日中共中央政治局会议强调要“加快高水平科技自立自强”,这一政策导向为长期投入基础科研的人才创造了宏观环境。生物医学数据科学、精准医学等新兴专业正站上风口,华西医院近期以60万年薪招聘的临床数据分析总监,要求必须兼具医学统计知识和基因组学背景。
趋势与信号——政策与市场正在说什么?
政策风向:科技创新与学科交叉的双重支撑
教育部在2026年的政策导向中明确要求临床医学专硕占比提升至70%,学硕招生规模缩减30%,但重点向“卡脖子”领域(如基因编辑、AI医疗)倾斜。这种“有压有保”的策略表面上看压缩了学硕规模,实则优化了结构——将资源集中于前沿交叉学科。
国务院学位委员会、教育部印发的《研究生教育学科专业目录(2022年)》将交叉学科作为一个门类正式“入驻”,下设智能科学与技术、纳米科学与工程等一级学科。2026年国务院政府工作报告提出“建设国家交叉学科中心”,这一信号表明国家对交叉学科发展的重视程度。
市场信号:新兴产业崛起与人才需求结构变化
人工智能医疗公司正高薪争夺学硕人才。资料显示,自然语言处理方向的AI医疗岗位年薪可达40万元,具备医学与数据科学交叉能力的人才更具市场稀缺性。数字医疗技术的发展推动了“微生物工厂”生产复杂药物,大幅降低生产成本的同时,也创造了新的就业岗位。
药企研发岗位为学硕提供了另一条路径。生物医药行业人才需求集中在研发及临床岗位,且对AI+生物交叉复合型人才需求提升明显。高端研发岗位通常要求硕士以上学历,博士在基因编辑等领域更具优势,而学硕作为博士预备阶段,在这一链条中占据重要位置。
战略选择与行动指南——你适合走学硕之路吗?
自我评估:哪些学生更适合考虑学硕?
内心驱动是首要考量因素。学硕适合那些对探索疾病本质、机理研究有强烈好奇心与热情的学生,而不仅仅是对临床操作技能感兴趣。如果享受发现问题、设计实验、分析数据的过程,而非追求即时临床反馈,学硕可能是更合适的选择。
能力特质决定适应程度。学硕需要较强的逻辑思维、耐心和抗压能力,能够忍受实验室的寂寞和科研的不确定性。相比之下,专硕更适合动手能力强、喜欢与人打交道、能够承受高强度值班(日均接诊30-50人次)和夜班压力的学生。
职业愿景指向不同方向。如果志不在局限于单一临床操作,而是向往产业研发、前沿科技创新、高校科研或成为跨领域专家,学硕提供了更广阔的可能性。专硕则更适合坚定走临床医生道路、希望尽快经济独立、目标普通三甲医院就业的考生。
关键决策点:如何规划学硕之路?
导师选择成为成败关键。寻找研究方向前沿、产业资源丰富或学术声誉卓著的导师至关重要。导师风格(“放养”vs“手把手”)需要与自身特点匹配——自主性强的学生可能适合“放养”型导师,而需要更多指导的学生则应选择“手把手”型导师。
研究方向需要平衡科学价值与应用潜力。选择既有科学价值又有应用潜力的课题,关注AI辅助新药研发、智能诊断设备开发、高端生物医用材料等交叉学科方向。中南大学湘雅医学院的交叉创新培育项目提供了医工、医理、医文、医信四个方向的参考。
产业衔接应提前布局。在读期间通过实习、项目合作、学术会议主动接触产业界,构建可转移的技能组合(如数据分析、专利撰写、项目管理)。浙江大学智能医学工程专业的课程表上,Python编程与解剖学并列,机器学习与病理学同排,这种课程设计本身就体现了产业衔接的思路。
在喧嚣中聆听内心的声音
2026年医学考研的“冷热抉择”本质上是个体在短期现实利益与长期发展潜力之间、在已知赛道与未知蓝海之间的一次战略权衡。专硕代表了一条清晰、稳健的职业道路——3年获得“四证合一”,直接进入临床岗位;学硕则意味着拥抱不确定性,以更长时间的投入博取在创新领域更广阔的可能性。
医学教育正面临深度重构。教育部要求2026年临床医学专硕占比提升至70%,学硕招生规模缩减30%,但重点向“卡脖子”领域倾斜。这并非简单的数量增减,而是结构优化——将有限资源集中于最有价值的交叉学科方向。
没有绝对的正确选择,只有基于深刻自我认知与趋势判断的合适之选。当绝大多数人涌向专硕时,学硕这条路径可能正从“冷门”转变为“稀缺”;当AI医疗公司愿意为“既懂医学又懂算法”的人才支付35万年薪时,传统的“冷热”定义正在被重新书写。
在这场医学考研的“冷热抉择”中,你是愿意跟随人群挤上专硕的“独木桥”,还是敢于探索学硕这条看似冷清却可能通往未来的“新航道”?
2026-04-21
在再生医学与细胞治疗的前沿领域,真正的突破往往来自长期深耕基础研究、并敢于直面临床空白的探索者。
重症肢体缺血(CLI)简单来说,它是外周动脉疾病最危险的阶段,下肢动脉严重狭窄或闭塞,休息时肢体依然严重缺血,已经出现疼痛、溃疡甚至坏死,属于血管急症,不及时处理很可能截肢。
被宣判“截肢”的千万人,无路可走。失去一条肢体,意味着行动能力彻底丧失、生活质量归零,长期康复、假肢适配与家庭照护,带来沉重的经济与精神压力。
并且截肢后引发的感染、血栓、心肺并发症,还会进一步推高死亡风险。一组冰冷的数据,道出了这一领域治疗空白的残酷现状:中国每年新增CLI患者约22万,全球每年新发高达130万;确诊后6个月截肢率达40%,1年死亡率超过20%;20%–40%的患者,最终会被贴上“无治疗选择”的标签。
目前临床主流治疗手段均存在明显局限,传统手术的介入治疗难以处理严重钙化、远端细小血管;药物仅能缓解疼痛、改善症状,无法修复已坏死的组织与供血系统;传统间充质干细胞疗法以调节局部微环境为主,无法直接形成有供血功能的新生血管网络。
并非所有患者都具备介入或手术条件。对大量终末期患者而言,现有的治疗体系几乎没有真正有效的选择。
20年深耕血管再生,
不愿成果只停在论文
黄永德教授的科研生涯,几乎全部围绕血管与再生展开。从香港中文大学博士毕业后,他先后赴斯坦福大学、康奈尔大学从事博士后及助理教授研究,2016年回到港中文任教,深耕血管生物学与干细胞定向分化近20年。
长期的基础研究让他形成了清晰的判断,想要真正改变重症肢体缺血(CLI)患者的结局,仅靠改善微循环、缓解症状远远不够,必须实现功能性血管的原位再生。
而iPSC技术的成熟,为这一目标提供了可行路径——它能够稳定、无限地提供细胞来源,并且可以高效分化为高纯度内皮细胞,这正是构成血管内壁、承担血流运输的核心功能细胞,也是实现血管再生不可或缺的“基础原料”。
当实验室的关键突破,具备了挽救患者的潜力,黄永德教授决定走出象牙塔。2023年底,他和几位好友成立NutrigeneAI脉源创科生物科技有限责任公司,希望通过将手中的成果产业化,为重症肢体缺血患者探索一个新的治疗选择。
图:NutrigeneAI脉源创科创始人 黄永德 Prof.Jack Wong
在他看来,科研的最终价值不只体现在论文与实验数据中,更在于能否真正解决临床难题、挽救患者生命:“不想让研究只停留在论文和实验里,想给那些被宣判无药可医的人,一个真实的希望。”
从“河道淤堵”到“水系重生”
相较于传统治疗思路,黄永德教授选择了一条更贴近疾病本质的技术路线。针对CLI患者血管闭塞、钙化严重、远端血流无法重建的临床痛点,团队将研究重心放在iPSC定向分化内皮细胞这一核心方向上,通过干细胞技术获得构成血管的关键细胞,再通过功能优化,使其能够在缺血环境下稳定形成新的血管结构。
对于这一核心技术的解读,黄永德教授教授在访谈中分享了一个生动的比喻。四肢就像一片需要灌溉的农田,血管就是贯穿其中的河流,负责输送血液、养分,维持组织的生机与活力。而重症肢体缺血,本质就是这条生命河道因严重钙化、堵塞彻底断流,农田彻底干涸、庄稼坏死,最终只能面临截肢的结局。
传统疗法始终困在“修旧河”的逻辑里,难以突破:常规药物就像给干涸的农田洒水、撒肥,只能短暂缓解症状,根本无法恢复河道的持续供水;手术介入则像是强行开挖、疏通淤堵的河道,不仅创伤大、风险高,面对已经完全闭塞、钙化严重的细小血管,往往无从下手,更无法从根源上解决问题。
脉源创科跳出了“修旧河”的思维,选择了一条“造新河”的全新路径。团队以iPSC为源头,定向分化出高纯度的血管内皮细胞——这是构成血管内壁的核心“建筑材料”,是真正能搭建血管的“砖瓦”。
更关键的是,团队研发出独有的增效启动专利技术,用内源性细胞因子给这些细胞做了一次“特种训练”,不做基因编辑、纯天然诱导,让它们的增殖、迁移、成管能力实现跨越式提升。
这些经过“强化训练”的增效型内皮细胞,就像一支自带施工图的智能工程兵:它们会精准奔赴缺血的病灶区域,手拉手搭建起全新的、有功能的血管“新河”,同时清理局部炎症的“淤泥”,动员身体自身的修复能力,加固周边的肌肉组织。
最终,在原本彻底断流的肢体里,重建出一套完整的新生血管水系,让血液重新顺畅流淌,从根源上逆转缺血坏死的进程。为血流恢复、组织修复与保肢治疗提供核心支撑。
同时,团队依托自研的Nutrigene AI系统,逐步建立从细胞分化预测、影像分析到全流程质控的智能化管控,可稳定制备纯度超98%的内皮细胞,支持3D自动化培养与规模化扩增,单批次产能可达百亿级,为后续标准化、工业化量产和全球商业化落地,打下了坚实的基础。
让科研成果,真正走到患者身边
只靠科学家做不出药,只懂产业抓不住源头创新,只谈资本没有长期耐心。脉源创科的核心竞争力,来自科研、产业、资本的黄金互补。
吴广基是与黄永德教授相识已久的挚友。作为公司的财务总监(CFO),吴广基先生拥有在国泰君安、鼎珮资本等头部金融机构资深从业经历,长期专注生物医药领域投融资与企业管理。并当选2026~2027年度香港东华三院总理。作为黄永德教授多年的朋友,他既懂行业逻辑,也懂长期价值,负责为公司搭建稳健的资本结构、治理体系与长期发展节奏,让研发不用为短期资金焦虑,专注啃最难的临床硬骨头。
王标博士获颁受 2025 年国家级启明人才选拔 (以前的千人计划),拥有海外顶尖学术背景与跨国药企、国内龙头创新药企的完整产业经验,曾在百时美施贵宝、强生、先声药业等平台负责研发、注册与产业化工作。他最懂“实验室成果如何变成药品”,负责把技术做成可量产、可申报、可落地的产品,打通临床转化最关键的“最后一公里”。
图:NutrigeneAI脉源创科CEO 王标 Dr.Bill Wong
昔日好友因“解决重症缺血患者无药可医”的共同目标走到一起,没有陌生磨合,一联手就是成熟的黄金三角。
我们的专利已经在多个国家申请,黄永德教授更荣获香港创新企业家奖及前海港澳台大赛铜奖与“最受投资者青睐奖”,充分验证了技术的原创性和产业化潜力。
快一点落地,让更多人用得起
面对重症肢体缺血患者刻不容缓的治疗需求,脉源创科制定了“速度优先、兼顾广度、分步落地”的产品与临床策略,在保证研发严谨性的同时,尽可能缩短转化周期,让创新疗法更早抵达患者。
团队采用两代产品循序渐进的研发路径,兼顾短期临床落地与长期商业化价值。第一代产品是HLA配型即用型细胞。依托已建立合作的临床级iPSC细胞系快速启动研发,无需从头构建细胞资源,大幅压缩前期准备周期。两条核心细胞株即可覆盖南中国约15%人口,能够以最快速度推进临床前验证与试验启动,让首批无治疗选择的患者优先受益。
第二代产品则是基因编辑通用型细胞,通过免疫原性优化与基因编辑技术,进一步降低细胞排斥风险,最终实现全人群通用、现货式供应。尽管面临更高的临床与监管要求,团队仍将其作为长期核心方向,打造真正可规模化、普惠化的下一代细胞药物。
为保障产品顺利进入临床,团队从细胞培养、制备到质量控制全流程合规化,确保细胞产品稳定、安全、可重复,满足人体试验与未来申报的严苛要求。
按照规划,公司正全力推进GMP标准化生产体系建设,并积极筹备启动研究者发起的早期临床研究(IIT),优先与国内顶尖三甲医院血管外科、介入科合作,基于中国患者人群验证疗法的安全性与有效性,为后续IND申报与多中心临床打下基础。
同时,团队充分借助海南博鳌乐城国际医疗旅游先行区的政策优势,依托特许医疗、特许临床研究、特许国际交流等支持政策,提前开展前沿疗法的临床应用与真实世界数据积累。相关数据可用于支持国内注册申报,有效缩短整体研发与上市周期,让患者在产品正式获批前就有机会获益。
在不断推进研发的过程中,团队始终没有忘记以可及性为初心,让患者用得上、用得起。
iPSC技术的核心优势之一,是能够实现标准化、自动化、规模化生产。脉源创科采用自动化3D培养体系,实现单批次高产能产出,从源头降低生产成本,摆脱个体化细胞治疗“高价难普惠”的行业困境。
团队始终将患者可负担性作为核心目标,致力于将单疗程治疗费用控制在普通家庭可承担的普惠区间,让保肢治疗不再因价格成为奢望。
在全球化布局上,公司同步推进美国、日本等国际市场的临床与合作准备,探索海外授权模式,以国际市场收益反哺国内研发与患者可及,最终形成“国内可及、全球价值”的可持续发展模式。
做血管再生的转化先锋
在黄永德教授的规划里,脉源创科的使命远不止开发一款细胞药物、解决一个适应症,更在于走出一条可复制、可推广的中国原创科研转化路径。
团队希望以重症肢体缺血这一全球空白为起点,树立iPSC血管再生领域的转化标杆,为国内干细胞创新药行业提供参考,推动更多源头创新成果走向市场、走向世界。
面向未来,脉源创科将持续深耕血管再生赛道,在CLI领域扎实推进临床与产业化的同时,逐步将技术平台拓展至缺血性心脏病、脑血管缺血、慢性创面修复、外周血管疾病等更广泛的未满足临床需求领域,构建以血管再生为核心、多适应症覆盖的全球化技术与产品平台。
“我们的终极目标,是让无药可医的血管疾病成为历史。”他表示,对每一位身处绝境的患者而言,保住一条腿,就是保住行走的权利、生活的尊严、家庭的完整与未来的可能。
对团队而言,这场始于实验室的血管再生探索,最终要奔赴的,是让更多人重获健康、重拾对生活热爱的长远使命。
专家点评:
尚骏投资总经理陈奕群
从投资视角看,脉源创科具备以下关键特质:
第一是 CLI患者基数大(中国年新增22万,全球130万),且20%-40%患者无有效治疗选择,截肢率高、患者保肢意愿强,支付支撑明确。
第二是 iPSC定向分化内皮细胞+增效启动专利技术,从"修旧河"转向"造新河",加上他们有AI系统支持自动化规模化生产,单批次百亿级产能,解决成本问题。
第三是从团队来看,黄永德教授(近20年血管再生研究)+ 吴广基(资本背景)+ 王标博士(产业化经验),黄金三角配置,互补性强、信任基础好。
这是一家值得深入尽调的早期细胞治疗企业。如果IIT临床数据良好、IND推进顺利,具备成长为血管再生细分赛道龙头的潜力。
从实验室走向市场
记录科研成果的产业化征程
再伟大的科学发现,只有走出实验室,才能真正改变世界。从基础研究到临床应用,从专利申请到商业转化,从校园实验室到产业化公司,每一步都是科研价值实现的关键跨越。
在全球科技竞争的背景下,科研机构正在成为创新策源地。无论是顶尖高校的技术转移办公室,还是国家实验室的成果转化中心,又或是医院的临床转化平台,它们都在孵化着改变未来的种子。这些机构的每一笔专利交易、每一次技术授权、每一个衍生企业诞生,都标志着科学智慧向生产力的转化。
橙果动态专栏致力于追踪记录这一转化过程的每个重要节点。我们以科研机构为视角,追踪国内外顶尖院校、研究所、医院等机构的成果转化动态,建立"机构-成果-转化"的完整图谱。通过持续追踪,我们希望呈现:哪些技术正在寻求产业化?哪些科研团队正在走向创业?哪些机构在持续产出高价值成果?哪些机构的转化机制值得借鉴?
让我们一起见证科学如何改变世界,记录每一次从0到1的突破。
——橙果动态专栏
▲创新者100人
丨创新者100人丨
▲人物专栏
丨人物专栏丨
▲橙果动态(成果转化)
▼高校
丨北京大学丨清华大学丨复旦大学丨上海交通大学丨
丨浙江大学丨中山大学丨同济大学丨华中科技大学丨
丨四川大学丨中南大学丨暨南大学丨首都医科大学丨
丨湖南大学丨南开大学丨天津大学丨西南医科大学丨
丨南京大学丨兰州大学丨重庆大学丨安徽医科大学丨
丨南方科技大学丨重庆交通大学丨北京航空航天大学丨
丨哈尔滨理工大学丨西安交通大学丨重庆医科大学丨
丨加州大学旧金山分校丨吉林大学丨温州医科大学丨
丨加州大学伯克利分校丨宁夏医科大学丨新乡医学院丨
丨东北林业大学丨天津工业大学丨山西医科大学丨
丨东北大学丨济南大学丨河南科技大学丨
丨沈阳药科大学丨山东中医药大学丨
丨成都中医药大学丨山东大学丨
▼医院
丨上海九院丨华山医院丨中山医院丨北京协和医院丨
丨华西医院丨湘雅医院丨齐鲁医院丨武汉协和医院丨
丨阜外医院丨北医一院丨北医三院丨南京鼓楼医院丨
丨南方医科大学南方医院丨中山大学附属第三医院丨
丨华中科技同济医院丨福建医科大学第一附属医院丨
丨中国科学技术大学附属第一医院丨北京世纪坛医院丨
丨西安交大附一院丨四川省人民医院丨上海长征医院丨
丨湖南省肿瘤医院丨北京天坛医院丨四川省肿瘤医院丨
丨南通大学附属医院丨安徽医科大学第一附属医院丨
丨武汉大学中南医院丨福建医科大学孟超肝胆医院丨
丨厦门大学附属心血管病医院丨常州市第一人民医院丨
丨江苏省肿瘤医院丨江苏省人民医院丨北京潞河医院丨
丨南京医科大学第二附属医院丨北京儿童医院丨
丨深圳儿童医院丨中山大学孙逸仙纪念医院丨
丨中山大学肿瘤防治中心丨东莞市人民医院丨
丨河南省肿瘤医院丨南京市口腔医院丨
丨中国医学科学院肿瘤医院深圳医院丨
丨河南科技大学第一附属医院丨
丨上海儿童医学中心丨
▼中国科学院
丨中国科学院金属研究所丨中国科学院深圳先进院丨
丨中国科学院苏州纳米技术与纳米仿生研究所丨
▼其他科研单位
丨中国医学科学院生物医学工程研究所丨
如果您所在机构有成果转化、专利交易或技术授权案例,或您关注并追踪国内外科研机构的转化实践,欢迎向我们投稿(投稿请添加下方微信)。我们将以机构和成果为视角,系统化记录每一次从实验室到市场的转化突破,建立"机构-成果-转化"的完整图谱,让更多行业同仁了解科研成果产业化的路径与经验。
100 项与 山西医科大学 相关的药物交易
登录后查看更多信息
100 项与 山西医科大学 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年06月10日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
6
21
临床前
临床1期
1
5
其他
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
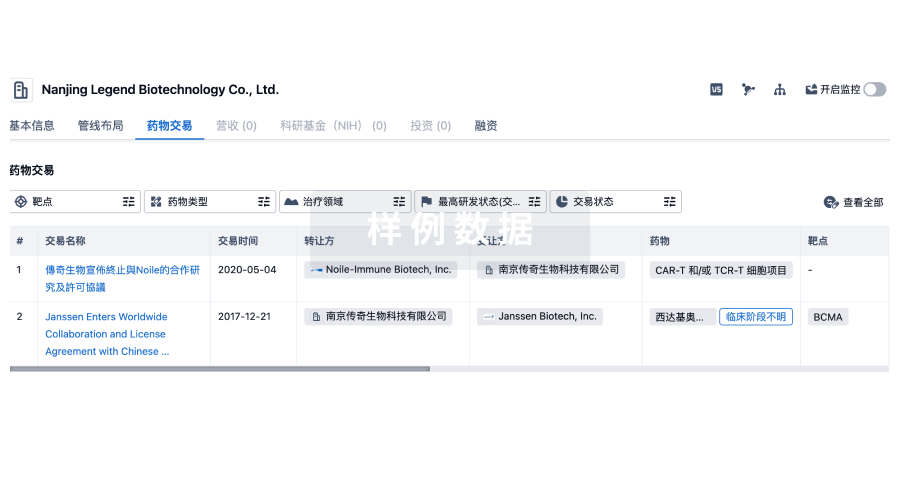
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
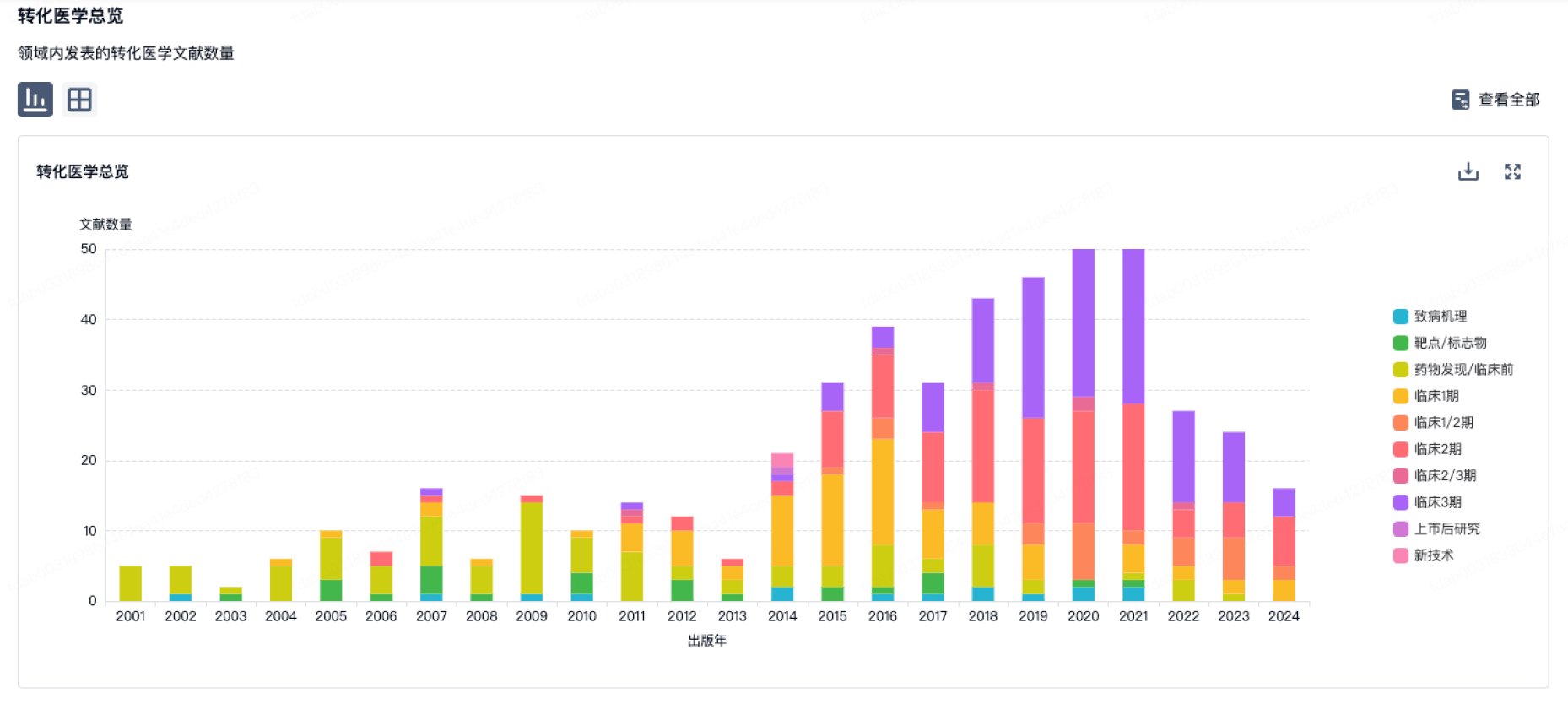
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
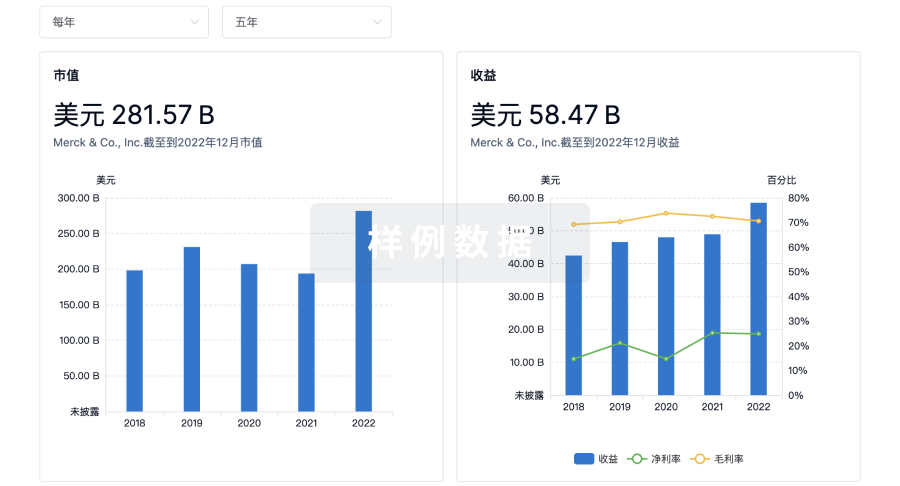
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
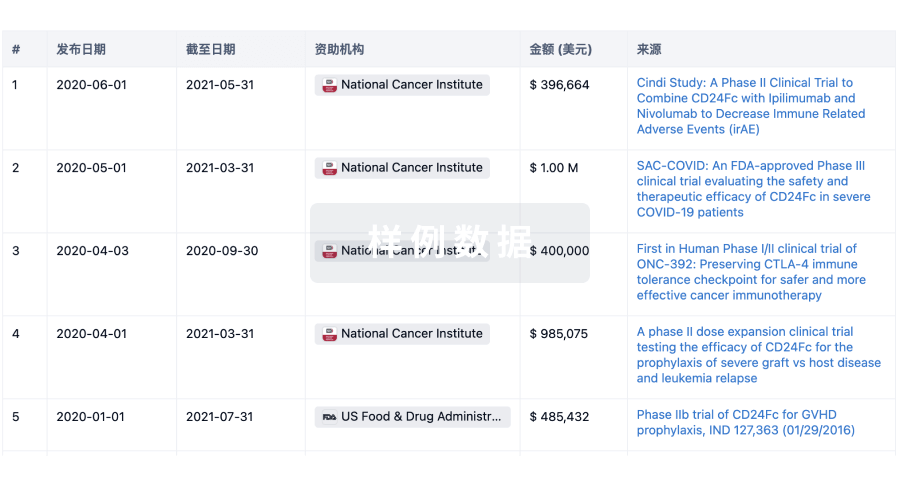
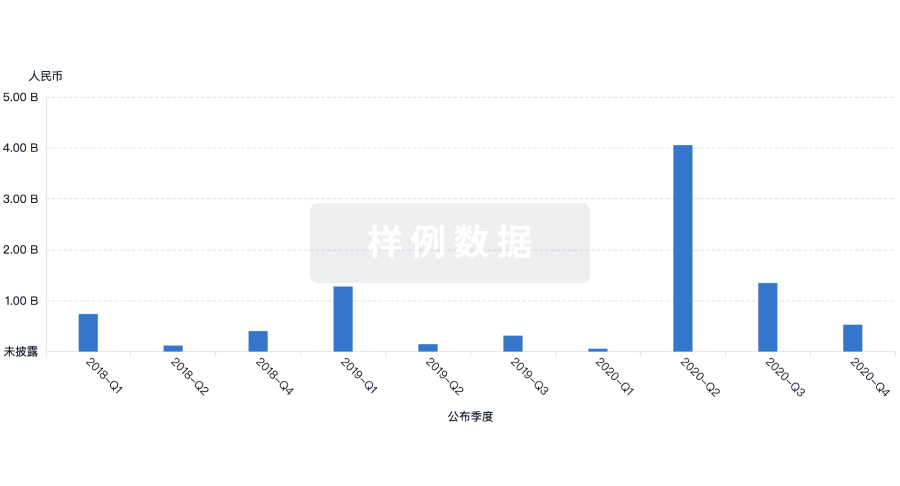
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
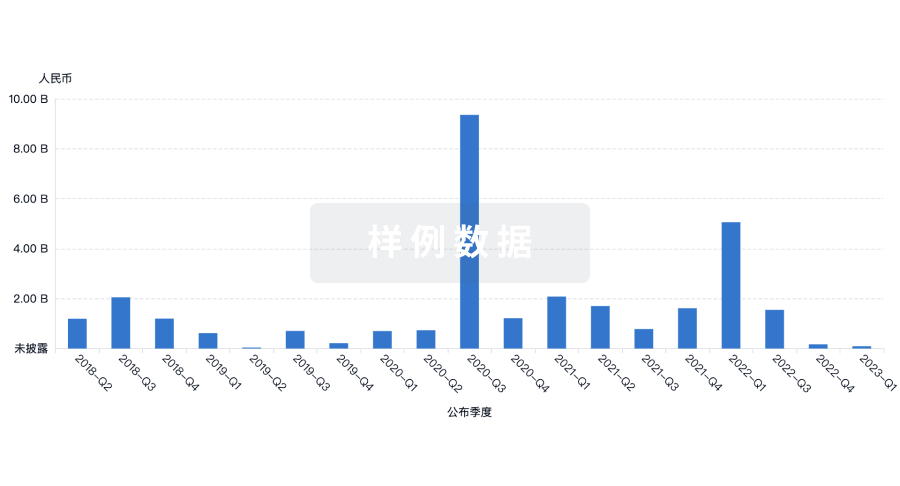
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用

