预约演示
更新于:2026-03-08

University of Sassari
更新于:2026-03-08
概览
标签
肿瘤
神经系统疾病
皮肤和肌肉骨骼疾病
小分子化药
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 小分子化药 | 5 |
| 排名前五的靶点 | 数量 |
|---|---|
| ACHE(乙酰胆碱酯酶) | 1 |
| CAIX(碳酸酐酶IX) | 1 |
| Tubulin(微管蛋白) | 1 |
| P-gp(P-糖蛋白-1) | 1 |
关联
5
项与 University of Sassari 相关的药物靶点- |
作用机制- |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 P-gp抑制剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 AChE抑制剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
42
项与 University of Sassari 相关的临床试验NCT07006194
Nicotinamide Levels in Serum, Aqueous Humor, and Tear Film in Glaucoma and Correlations With Mitochondrial Damage-Associated Molecular Patterns (mtDAMPs) and Senescence-Associated Secretory Phenotype (SASP)
Background Glaucoma is a multifactorial, chronic, and progressive eye disease characterized by the irreversible loss of retinal ganglion cells (RGCs). It is the second leading cause of blindness globally, with approximately 76 million patients affected in 2020, a number expected to rise to 120 million in the coming decades, especially in Africa and Asia. Elevated intraocular pressure (IOP) is a significant risk factor for glaucoma, and its reduction remains the only scientifically proven approach to slowing visual function decline. However, many glaucoma patients continue to experience visual loss even with IOP values within the normal range, due to the disease's multifactorial nature. Besides IOP reduction, there is a need for direct neuroprotection to address other factors causing RGC damage.
Mitochondrial dysfunction has emerged as a critical early factor in RGC damage, making glaucoma, at least in part, resemble a mitochondrial disease. Mitochondria play a key role in cellular functions such as energy production, redox metabolism, and maintaining mitochondrial function. In glaucomatous conditions, mitochondrial damage leads to the release of mitochondrial damage-associated molecular patterns (mtDAMPs), triggering chronic inflammation and tissue damage. This inflammation is exacerbated by the Senescence-Associated Secretory Phenotype (SASP), a process where senescent cells secrete bioactive molecules that contribute to cellular dysfunction and glaucoma progression.
Among potential neuroprotective approaches, nicotinamide (NAM) and its precursor nicotinamide riboside (NR) are gaining attention. These compounds support mitochondrial function and NAD production, which is essential for cellular vitality. Research indicates that reductions in NAM levels correlate with glaucoma progression. For instance, studies have shown that NAD precursors may prevent or delay RGC degeneration, suggesting a promising adjunctive treatment for glaucoma patients.
Study Objectives The study aims to measure NAM levels in ocular and non-ocular biological fluids (serum, aqueous humor, and tear fluid) of patients at different stages of glaucoma. The study will correlate NAM concentrations with disease severity and mitochondrial function markers. Furthermore, NAD levels in peripheral blood mononuclear cells (PBMCs) will be assessed to investigate potential biomarkers for glaucoma progression. A secondary objective is to evaluate the impact of dietary supplementation with nicotinamide and nicotinamide riboside (iNAD®) on NAD levels in pharmacologically controlled glaucoma patients.
Methods This cross-sectional, case-control, multi-center study involves three universities: University of G. d'Annunzio Chieti-Pescara, University of Pisa, and University of Sassari. Biological fluid analyses will be conducted at the Animal Biology Laboratory affiliated with the University of G. d'Annunzio Chieti-Pescara.
Patients will be categorized into four groups:
1. Uncontrolled glaucoma patients scheduled for glaucoma surgery.
2. Controlled glaucoma patients scheduled for cataract surgery.
3. Healthy controls undergoing cataract surgery.
4. Pharmacologically controlled glaucoma patients supplemented with iNAD®. Biological fluids (plasma, PBMCs, tear fluid, and aqueous humor) will be collected for analysis, including measures of NAD, mtDAMPs, and SASP components. These measures aim to provide insights into the molecular mechanisms underlying glaucoma and potential biomarkers for disease progression.
Statistical Analysis Sample size estimation was calculated using G*Power for a priori one-way ANOVA analysis. Assuming a mean NAM concentration of 0.14 μM (SD=0.12) for glaucoma patients and 0.19 μM (SD=0.13) for controls, with a power of 80% and alpha of 0.05, a minimum of 246 patients (82 per group) are required.
Mitochondrial dysfunction has emerged as a critical early factor in RGC damage, making glaucoma, at least in part, resemble a mitochondrial disease. Mitochondria play a key role in cellular functions such as energy production, redox metabolism, and maintaining mitochondrial function. In glaucomatous conditions, mitochondrial damage leads to the release of mitochondrial damage-associated molecular patterns (mtDAMPs), triggering chronic inflammation and tissue damage. This inflammation is exacerbated by the Senescence-Associated Secretory Phenotype (SASP), a process where senescent cells secrete bioactive molecules that contribute to cellular dysfunction and glaucoma progression.
Among potential neuroprotective approaches, nicotinamide (NAM) and its precursor nicotinamide riboside (NR) are gaining attention. These compounds support mitochondrial function and NAD production, which is essential for cellular vitality. Research indicates that reductions in NAM levels correlate with glaucoma progression. For instance, studies have shown that NAD precursors may prevent or delay RGC degeneration, suggesting a promising adjunctive treatment for glaucoma patients.
Study Objectives The study aims to measure NAM levels in ocular and non-ocular biological fluids (serum, aqueous humor, and tear fluid) of patients at different stages of glaucoma. The study will correlate NAM concentrations with disease severity and mitochondrial function markers. Furthermore, NAD levels in peripheral blood mononuclear cells (PBMCs) will be assessed to investigate potential biomarkers for glaucoma progression. A secondary objective is to evaluate the impact of dietary supplementation with nicotinamide and nicotinamide riboside (iNAD®) on NAD levels in pharmacologically controlled glaucoma patients.
Methods This cross-sectional, case-control, multi-center study involves three universities: University of G. d'Annunzio Chieti-Pescara, University of Pisa, and University of Sassari. Biological fluid analyses will be conducted at the Animal Biology Laboratory affiliated with the University of G. d'Annunzio Chieti-Pescara.
Patients will be categorized into four groups:
1. Uncontrolled glaucoma patients scheduled for glaucoma surgery.
2. Controlled glaucoma patients scheduled for cataract surgery.
3. Healthy controls undergoing cataract surgery.
4. Pharmacologically controlled glaucoma patients supplemented with iNAD®. Biological fluids (plasma, PBMCs, tear fluid, and aqueous humor) will be collected for analysis, including measures of NAD, mtDAMPs, and SASP components. These measures aim to provide insights into the molecular mechanisms underlying glaucoma and potential biomarkers for disease progression.
Statistical Analysis Sample size estimation was calculated using G*Power for a priori one-way ANOVA analysis. Assuming a mean NAM concentration of 0.14 μM (SD=0.12) for glaucoma patients and 0.19 μM (SD=0.13) for controls, with a power of 80% and alpha of 0.05, a minimum of 246 patients (82 per group) are required.
开始日期2025-10-01 |
申办/合作机构 |
NCT06652776
The Italian COPD Registry. the DescribinG BUrden of COPD and Occurrence of MortaLity in a Cohort of Italian Patients
Chronic obstructive pulmonary disease (COPD) is a treatable but debilitating medical condition associated with persistent symptoms and chronic airflow obstruction. Despite the availability of multiple therapeutic options, COPD is the third leading cause of death worldwide and has a substantial socioeconomic impact.
The present real life study is aimed at describing the clinical and functional characteristics, treatment patterns, impact of exacerbations and comorbidities and their association with mortality in a large cohort of Italian patients with COPD.
The present real life study is aimed at describing the clinical and functional characteristics, treatment patterns, impact of exacerbations and comorbidities and their association with mortality in a large cohort of Italian patients with COPD.
开始日期2024-09-30 |
申办/合作机构  University of Milan University of Milan [+9] |
NCT06286605
Horizontal Ridge Augmentation Using One-stage Guided Bone Regeneration (GBR) for Bone Defect Class IV of Cawed and Howell With Collagene Membrane (OssMem) and a Mix of Bovine Bone Substitute (A-Oss) and Autogenous Bone Versus A-Oss and LCR-A, a Synthetic Bone : a Randomized Controlled Trial
The aim of this randomized controlled trial is to evaluate the tridimensional bone stability after horizontal one-stage GBR using collagene membrane (OssMem) with a mix of Bovine Bone Substitute (A-Oss) and autogenous bone (test group) versus A-Oss and LCR-A, a synthetic bone (control group).
开始日期2024-03-01 |
申办/合作机构 |
100 项与 University of Sassari 相关的临床结果
登录后查看更多信息
0 项与 University of Sassari 相关的专利(医药)
登录后查看更多信息
7,670
项与 University of Sassari 相关的文献(医药)2026-04-01·AMBIO
Wolves on the phone: Public calls reveal a rise in urban concerns as wolves recolonize human-dominated areas
Article
作者: Brogi, Rudy ; Cerri, Jacopo ; Apollonio, Marco ; Marshall-Pescini, Sarah ; Lazzaroni, Martina ; Mattioli, Luca ; Neirotti, Giovanna
Abstract:
European wolf populations are expanding into human-dominated landscapes, triggering novel interactions with citizens and public concerns that may disrupt the traditional urban–rural divide in wolf attitudes and reshape conservation paradigms. We modelled the spatiotemporal distribution and valence of the wolf reports received through a dedicated phone service in Tuscany, Italy (2021–2024). Reports were significantly more common in: (i) late winter, aligning with the peak dispersal period and increased wolf movements. (ii) recently recolonized areas, suggesting a wolf-novelty effect, and (iii) urban areas, where negative valence was also more likely. Public concerns about wolves are increasingly emerging in urban areas, potentially disrupting the traditionally more supportive urban stance on wolf presence. In one of the European regions where wolf recovery began earlier and progressed further, our findings signal a broader shift in public attitudes that may weaken support for wolf conservation, potentially anticipating similar developments in areas of more recent recovery.Graphical Abstract
2026-04-01·Data in Brief
A human fecal metaproteomic dataset from celiac disease patients on gluten-free diet with or without poly-autoimmunity
Article
作者: Dore, Maria Pina ; Pira, Giovanna ; Errigo, Alessandra ; Uzzau, Sergio ; Pes, Giovanni Mario ; Abbondio, Marcello ; Manetti, Roberto ; Sau, Rosangela ; Tanca, Alessandro ; Bibbò, Stefano
This dataset provides the fecal metaproteome profiles of 28 celiac disease patients on a gluten-free diet, distinguished by the presence or absence of co-occurring autoimmune conditions. The resource includes raw liquid chromatography-tandem mass spectrometry (LC-MS/MS) files, database search results, protein/peptide identification outputs, and taxonomic/functional annotation outputs, along with comprehensive anthropometric, clinical, and dietary metadata for each patient. The identified proteins originate from microbial, human, and plant sources, consistent with the multi-database search strategy used. This collection is designed for reuse in meta-analyses and integrative studies exploring functional changes in the gut microbiome related to auto-immune status and dietary variables. The complete dataset is available via the ProteomeXchange Consortium with the identifier PXD069517.
2026-03-01·JOURNAL OF DAIRY SCIENCE
Single-step genomic evaluation for production and type traits in the Italian Mediterranean buffalo
Article
作者: Biffani, S ; Campanile, G ; Gombia, Y ; Rossi, D ; Cesarani, A ; Zullo, L ; Gubitosi, L ; Gómez-Carpio, M ; Neglia, L ; Cimmino, R ; Di Vuolo, G
Single-step GBLUP (ssGBLUP) is becoming the most used method to predict breeding values in livestock, offering several advantages in terms of computational efficiency and simplifying the genetic evaluation process by integrating genomic, pedigree, and phenotypic information in a single step. Genomic information is now available for the Italian Mediterranean buffalo (IMB), and its inclusion in the genetic evaluation system could increase both evaluation accuracy and genetic progress of the breeding objectives. The aim of this study was to test the feasibility of ssGBLUP and to present the first results of the implementation of a genomic evaluation for IMB. Phenotypic information on production traits (milk yield adjusted to 270 d, fat and protein yield and content, and cheese yield) and morphology traits (feet and legs scores and udder teat scores) were used in this study. Production records included 792,200 lactations from 293,633 buffalo cows born from 1984 to 2021. Morphological traits were from 99,609 buffalo cows from 2004 to 2023. Regarding the genotypes, a total of 3,647 genotyped animals were used. Data were analyzed fitting 2 multitrait animal models, a 6-trait model for production data, and a 2-trait model for morphology data. Breeding values (BV) were estimated with BLUP and ssGBLUP models, both considering unknown parent groups. The methods were compared in terms of correlation between BV and genetic trends. Results were also validated with the linear regression (LR) method. Three different scenarios were used according to the cut-off year used to create the partial datasets, namely T2013, T2016, and T2018. The genomic and nongenomic BV were strongly correlated, and genetic trends for each trait were similar. The average increase in accuracy moving from BLUP to ssGBLUP across traits ranged from +3% to +12%. The LR method statistics confirmed the effectiveness of the ssGBLUP method. The average validation correlations across production traits and scenarios for BLUP and ssGBLUP by female and bull groups were 0.54 and 0.47, and 0.63 and 0.52, respectively. Accuracies were also higher with ssGBLUP (0.62/0.55) compared with BLUP (0.53/0.51). The best dispersion values (i.e., closer to 1) were observed for ssGBLUP (T2013, T2016). The ssGBLUP method provided better results across genotyped and nongenotyped animals, particularly in terms of a relative increase in accuracy associated with the inclusion of phenotypes. These results showed that implementing ssGBLUP in the breeding program can generate more accurate predictions for production and morphological traits in dairy IMB.
1
项与 University of Sassari 相关的新闻(医药)2024-04-03
A routine blood test that measures a patient's inflammation levels could improve the early diagnosis and management of a wide range of debilitating autoimmune diseases. The Systemic Inflammation Index (SII) uses information from routine laboratory data to measure inflammation in the body and examining this index in a new way could provide vital answers.
A routine blood test that measures a patient's inflammation levels could improve the early diagnosis and management of a wide range of debilitating autoimmune diseases.
The Systemic Inflammation Index (SII) uses information from routine laboratory data to measure inflammation in the body and examining this index in a new way could provide vital answers says Strategic Professor of Clinical Pharmacology Arduino Mangoni from Flinders University's College of Medicine and Public Health.
"The index, that measures inflammatory cells in the blood, could play a key role in early diagnosis, patient management strategies and health initiatives to help with autoimmune diseases," says Professor Mangoni.
Autoimmune diseases, which include a range of more than 80 different illnesses from inflammatory bowel disease (Crohn's disease and ulcerative colitis), rheumatoid arthritis to type 1 diabetes and multiple sclerosis, occur when the immune system attacks the body.
The diseases affect around 5 per cent of people in Australia and New Zealand and often have symptoms that are extremely distressing and debilitating and, if undetected, can lead to serious organ and body tissue damage.
In a new study by Professor Mangoni and Italian Professor Angelo Zinellu from the University of Sassari, the researchers carried out a systematic review and meta-analysis of multiple research articles on the potential use of the index in diagnosing the presence and severity of autoimmune diseases.
"A key element for the successful management of these diseases is being able to identify them at an early stage and then provide targeted treatment," says Professor Mangoni.
"Currently available biomarkers of inflammation, measured in the blood, have limited diagnostic accuracy in several types of immunological diseases leading to harmful delays in the diagnosis and treatment of these conditions.
"This issue has prompted the search for new, more accurate biomarkers of immunological diseases to enhance diagnosis and overall management.
"Among these candidate biomarkers, those that are derived from routine blood tests measuring the number of specific cell types, such as neutrophils, lymphocytes, and monocytes, have been increasingly studied in immunological diseases.
"One of these haematological indices, the SII, has been shown to be particularly accurate in the diagnosis of other conditions characterised by excess inflammation and dysregulated immunity, such as coronavirus disease 2019 (COVID-19).
"Our study of all the evidence so far confirms that it is very likely that the SII is superior to currently available biomarkers and could be routinely used in clinical practice to optimally diagnose and manage patients with immunological diseases," he says.
100 项与 University of Sassari 相关的药物交易
登录后查看更多信息
100 项与 University of Sassari 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年03月25日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
临床前
5
1
其他
登录后查看更多信息
当前项目
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
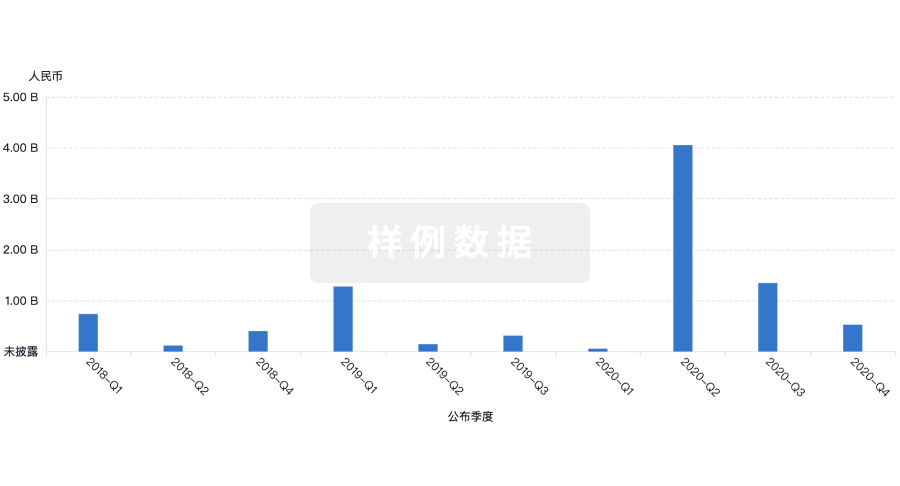
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
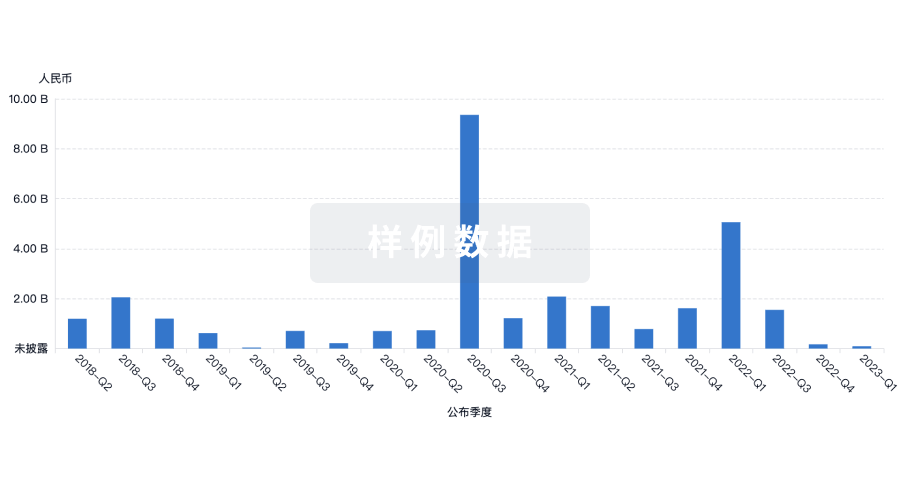
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
