预约演示
更新于:2026-05-21

Beijing University of Traditional Chinese Medicine
更新于:2026-05-21
概览
标签
神经系统疾病
其他疾病
心血管疾病
小分子化药
中药
多糖药物
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 小分子化药 | 18 |
| 中药 | 5 |
| 多糖药物 | 3 |
| 诱导性多能干细胞 | 1 |
| CAR-T | 1 |
关联
28
项与 北京中医药大学 相关的药物靶点- |
作用机制 免疫调节剂 |
原研机构 |
非在研适应症- |
最高研发阶段批准上市 |
首次获批国家/地区 中国 |
首次获批日期2005-11-18 |
302
项与 北京中医药大学 相关的临床试验ITMCTR2026000306
Infrared thermography and EEG mechanisms of the Xu technique in the Liuzijue Qigong for its liver-soothing effects
开始日期2026-02-28 |
申办/合作机构 |
ITMCTR2026000218
The Effect of Multimedia Traditional Chinese Medicine Health Education Based on Response-Adaptive Randomization on Functional Dyspepsia and Study on Key Techniques
开始日期2026-02-20 |
申办/合作机构 |
CTR20260240
多中心、随机、双盲、安慰剂平行对照评价苏合颗粒治疗慢性萎缩性胃炎(脾虚瘀滞证)的有效性及安全性的Ⅱ期临床试验
主要目的:
评价苏合颗粒治疗慢性萎缩性胃炎的有效性。
次要目的:
评价苏合颗粒治疗慢性萎缩性胃炎的安全性。
探索苏合颗粒治疗慢性萎缩性胃炎的合理剂量。
探索苏合颗粒治疗慢性萎缩性胃炎的PD特征。
开始日期2026-02-06 |
申办/合作机构  北京中医药大学 北京中医药大学 [+1] |
100 项与 北京中医药大学 相关的临床结果
登录后查看更多信息
0 项与 北京中医药大学 相关的专利(医药)
登录后查看更多信息
8,390
项与 北京中医药大学 相关的文献(医药)2026-12-31·Human Vaccines & Immunotherapeutics
Economic evaluation of 22 HPV vaccination strategies to minimize the risk of penile, anal, and oropharyngeal cancers in Chinese males: A Markov model analysis
Article
作者: Zhang, Xin ; Zhou, Liangru ; Qin, Ruixi ; Chai, Peipei ; Wang, Di ; Xu, Gan ; Luo, Weihua ; Liu, Bingjie
Males can benefit significantly from human papillomavirus (HPV) vaccination. This study developed a decision-Markov model to evaluate the lifetime effectiveness and cost-effectiveness of 22 HPV vaccination strategies for preventing penile, anal, and oropharyngeal cancers. From a healthcare system perspective, vaccination strategies targeting males aged 14 and 40 were compared against no vaccination. Key parameters, including disease progression probabilities, cancer incidence, health state utilities, and direct medical costs, were sourced from the published literature and national databases. We calculated the number of cancer cases and deaths under different strategies and compared their cost-effectiveness. Results demonstrated that male vaccination is highly cost-effective. Vaccinating 14-y-old males averted between 73,724 and 416,654 cancer cases and between 1,102 and 6,229 deaths, depending on the vaccine valency and coverage rate. Similarly, strategies targeting 40-y-old males demonstrated substantial benefits. Overall, 20 of the 22 strategies were deemed highly cost-effective against a willingness-to-pay threshold of three times the national gross domestic product per capita. Sensitivity analysis confirmed the results' robustness. This study provides economic evidence to inform national HPV vaccination policies, demonstrating that including males, particularly adolescents, in vaccination programs represents a cost-effective investment for reducing the burden of HPV-associated cancers in China.
2026-12-31·PHARMACEUTICAL BIOLOGY
Xijiaqi Formula attenuates myocardial infarction-induced chronic heart failure by inhibiting TGF-β1/Smads-mediated fibroblast activation
Article
作者: Shang, Hong-Cai ; Lin, Sheng ; Zhou, Lin-Nan ; Li, Wan-Ting ; Xia, Huan ; Xia, Gui-Yang ; Wei, Xiao-Hong ; Zhang, Qian ; Chen, Jie ; Wu, Xue-Fen ; Wu, Yu-Zhuo
CONTEXT:
Chronic heart failure (CHF) is the common concern following myocardial infarction. Xijiaqi Formula (XJQ) effectively treats CHF clinically while the underlying mechanism remains unclarified.
OBJECTIVE:
This study was designed to investigate the effect and mechanism of XJQ on CHF in rats following MI.
MATERIALS AND METHODS:
UHPLC-Q-TOF-MS/MS was utilized to analyze the constituents of XJQ. Post-MI CHF was induced in rats by permanent ligation of the left anterior descending coronary artery. Cardiac function was assessed by echocardiogram and hemodynamic. Myocardial morphology, fibrosis, and ultrastructure were evaluated through HE staining, Masson staining, and transmission electron microscopy, respectively. Myocardial transcriptomics were employed to identify genes that may be altered in response to XJQ treatment. Consistently, XJQ inhibited TGF-β1-induced myofibroblast activation in vitro.
RESULTS:
XJQ significantly improved cardiac function and structure, mitigating fibrosis, edema, and mitochondrial damage, while reducing key biomarkers (BNP, NT-proBNP, cTnT, Ang II). Transcriptomic analysis indicated that the differentially expressed genes influenced by XJQ were predominantly associated with extracellular matrix (ECM) remodeling. Notably, XJQ inhibited the upregulation of ECM proteins, including Adamts2, Fbln1, Itgbl1, and Ltbp3 mRNAs, as well as the proteins TGF-β1, P-Smad2/3, and MMP2. Additionally, in vitro experiments revealed that XJQ significantly suppressed the activation of myofibroblasts induced by TGF-β1.
CONCLUSIONS:
XJQ significantly attenuated the progression from MI to CHF by inhibiting fibroblast activation mediated by TGF-β1/Smads signaling. The molecular mechanisms underlying this effect appear to be intricately linked to its regulation of ECM remodeling proteins, specifically Adamts2, Itgbl1, Fbln1, and Ltbp3.
2026-12-31·Gut Microbes
Gut microbiota metabolic reprogramming drives the development of metabolic diseases in the host
Review
作者: Wei, Xue ; Wang, Yanrong ; Guan, Yuanyuan ; Huang, Beibei ; Li, Lingru ; Zheng, Yanfei ; Sun, Wenlong
Metabolic diseases pose a major global health challenge, the pathogenesis of which centers on "metabolic reprogramming"; that is, the adaptive or pathological rewiring of metabolic pathways. Emerging evidence indicates that gut microbiota dysbiosis triggers its metabolic reprogramming prior to host disease onset and plays a pivotal role in the development of metabolic disorders. However, unlike host metabolic reprogramming, which has been well characterized, the pathogenic mechanisms resulting from gut microbiota metabolic reprogramming remain poorly understood, creating a critical knowledge gap regarding its role in systemic metabolic diseases. To address this gap, this review introduces the concept of gut microbiota metabolic reprogramming and establishes its foundational role in systemic metabolic disease. We propose that gut microbiota metabolic reprogramming constitutes an early pathogenic event, preceding and potentially driving subsequent metabolic alterations in the host. Within this framework, we systematically reveal that an imbalance in the gut microbiota leads to its significant metabolic reprogramming, including lipid, glucose, amino acid, and uric acid metabolism, which in turn regulates host-wide metabolic and immune homeostasis and contributes to the development of metabolic diseases. By integrating these mechanisms into a coherent model, our work provides a novel paradigm for understanding metabolic regulation. This model refines the fundamental pathophysiology of metabolic disorders and highlights new possibilities for targeting the microbiome for the prevention and treatment of metabolic disorders.
524
项与 北京中医药大学 相关的新闻(医药)2026-05-20
·今日头条
> 中医药,传承千年的东方智慧,正通过人工智能等现代科技突破国际认知壁垒,在196个国家和地区获得传播与认可。
从智利将针灸纳入国家公共卫生体系,到新加坡研发AI舌诊系统,科技赋能不仅提升了中医药的临床效能与产业规模,更重塑其全球形象——但这一进程仍面临高质量数据稀缺等基础挑战。
## 国际突破:政策认可与标准主导
世界卫生组织数据显示,全球**113个成员国认可针灸等中医药诊疗方式**,29国制定相关法规,20国将其纳入医保体系。智利驻华大使巴勃罗・阿里亚兰表示,针灸已成为当地应用最广泛的补充医学之一,并正式纳入公共卫生体系,成为中智文化交流纽带。
在国际标准层面,中国主导制定全球现行**181项中医药ISO标准中的75%以上**,中药处方药在海外注册总数突破1000个。
三十年间,中药工业总产值从1996年的约**234亿元增长至2025年的8000余亿元**,规模扩大34倍;中医药SCI论文量从几十篇增至**2.6万篇**,占全球总数的43.5%。
## AI全场景赋能:从诊疗到制造的链式革命
人工智能已深度融入中医药全产业链,在四大应用场景取得实质进展:
- **临床诊疗经验传承**:通过AI将名老中医经验数智化,实现高效传承推广。例如,青岛西海岸新区上线中医药智能辅助诊断系统,覆盖35家基层医疗机构,上线一个月内**AI推荐诊疗14562次**,辅助诊断14952次。

- **中药新药研发提速**:AI助力新药成药性评估与靶点识别。中国中医科学院广安门医院团队采用人工智能算法,精准识别出子宫内膜异位症复发关键靶点VEGFA与LGALS3,定位香叶醇与丹酚酸B的靶向作用,成功转化中药创新药“活血消异颗粒”,并进入Ⅱ期临床试验。
- **循证研究证据生产**:AI提升临床疗效证据生产和转化效率,支持科学决策与监管。天津中医药大学团队构建“全景解析-靶向干预-调平纠偏”中药代谢生物学研究新范式,为诠释中药效应靶标提供新路径。
- **智能制造绿色转型**:AI推动传统生产向高品质、高效能、绿色化升级。上药杏灵采用“**PAT+AI+MSPC**”联合质控方案,将检测时间从2小时缩短至**3分钟**,准确率超98.5%。
江中药业通过连续制造技术与AI融合,2024年万元产值综合能耗和碳排放强度较2020年分别下降**37%和51%**。
## 案例深描:基层应用与前沿探索
在基层医疗,AI正降低服务门槛。上海中医药大学早在2010年研制首台中医四诊客观化集成诊断仪,如今上海市第七人民医院揭牌“中医药人工智能中心”,致力于打造“懂中医、会辨证”的垂类大模型。在国际前沿,新加坡政府网罗AI人才,推动中医与科技结合:
> 新加坡卫生部属下的全国医疗科技机构在开发“AI舌诊”,希望将其发展为远程治疗和日常健康管理的手段;同时首创推拿机器人,一名医师可同时操控多台设备,治疗中生成数据报告供后续参考。

此外,新加坡南洋理工大学与北京中医药大学东方医院成立联合实验室,用AI为糖尿病前期肥胖等慢性病提供预防治疗方案。
## 文化出海:品牌增值与教育赋能
科技助力中医药文化国际传播。湖北蕲春“蕲艾”品牌价值突破**150.36亿元**,全链产值超190亿元,企业通过智能化转型,在新加坡设立国际运营与研发中心,推动艾灸文化出海。
大明古艾公司采用智能滴灌技术,使蕲艾挥发油含量达**1.5%**,远超行业0.8%标准,并推出AI智能艾灸舱等产品。
在教育领域,北京中医药大学牵头研发的“**薪火中国药**”中医药教育大模型,参数规模达700亿,成为国内首款完成国家备案的中医药学科教育大模型,覆盖助学、助教、文化传播等场景,亮相世界数字教育大会。
## 挑战与未来:数据瓶颈与持续创新
尽管进展显著,AI赋能中医药仍面临基础性挑战。天津中医药大学副校长张俊华指出,可供AI模型训练的高质量数据集缺乏是一个关键突破点。随着国家科技专项在“十五五”期间的体系化布局,相关研究任务预计将取得重要成果。
中医药现代化三十年,从产值飙升到论文领跑,科技已为其注入新质生产力;未来,持续深化多学科融合与创新,将助力中医药在全球健康舞台上扮演更核心角色。
临床2期上市批准临床1期生物类似药
2026-05-20
创新,是鲁南制药基因里的密码。而沂蒙精神,则是鲁南制药发展的精神根基。张贵民,这位土生土长的临沂人,从小受沂蒙精神的滋养。出身科研的他,从科研一线人员到车间主任,再到鲁南制药集团党委书记、董事长、总经理,创新贯穿于他的整个职业生涯,而沂蒙精神则支撑他带领企业渡过一次次难关,实现一次次跨越。鲁南制药集团,正在“千亿鲁南 健康世界”的道路上不断迈进。搭建科研平台 筑牢创新根基在“经方与现代中药融合创新全国重点实验室”内,上千平方米的工作空间里,多位科研人员有条不紊地忙碌着。鲁南制药集团最近投放市场的抗肿瘤药物益立达利妥昔单抗注射液正是从这里研制成功的,科研人员夜以继日的努力,历经成千上万次的实验,为千万名患者带来了福音。以“经方与现代中药融合创新全国重点实验室”为代表的高能级平台,联合中国中医科学院、北京中医药大学等顶尖机构,突破中药药效物质基础、制剂工艺等关键技术,推动传统中药现代化发展。鲁南制药还与全球百余所高校及科研机构建立深度合作,承担国家级项目20余项,荣获省部级科技奖励30余项,持续夯实创新发展根基。鲁南制药构建以企业为主体的“产学研用”深度融合创新体系。企业现已拥有5个国家级和11个省级科研平台,覆盖化学药、中药、生物药三大领域,每年研发投入占营收10%以上,支撑超过100个新药项目同步推进。汇聚人才力量 激发创新活力“人才是创新的第一资源”,这是出身科研一线的张贵民始终坚持的理念。通过完善的“引育留用”机制,他打造出一支年轻化、高水平的研发团队。目前企业研发人员占比超过15%,平均年龄仅32岁,展现出充沛的创新活力。近年来,鲁南制药引进包括泰山学者在内的领军人才30余名,组建2000余人的研发团队,并与山东大学、中国药科大学等高校共建人才联合培养基地,年投入培训经费超亿元。通过“揭榜挂帅”机制和亿元级创新基金的激励,实现科研人员“名利双收”,极大释放了人才创新潜能。2025年推出的“神农计划”,面向全球提供超过1000个岗位,持续扩充高端人才储备,为企业创新发展注入源源不断的智力支持。推动成果转化 释放创新价值张贵民强调:“创新必须转化为实实在在的产业价值。”在他的主导下,鲁南制药建立起“研发—中试—生产—销售”全链条转化机制,推动科研成果快速落地。企业在上游突破培养基等技术垄断;在中游开发平价高效的生物类似药;在下游带动区域1000余家药企协同发展,形成完整的产业链价值布局。通过数字化、智能化改造,企业将经典名方“荆防败毒散”升级为现代中药“启达力®荆防颗粒”,成为广受欢迎的抗疫产品;自主研发的CD20单抗利妥昔注射液成功上市。今年以来,25个产品国内获批,16个产品走向国际,40余个一类新药进入临床,“创新药+仿制药+中药”三驾马车并驱的产品矩阵日益成熟。目前企业成果转化率超90%,拥有专利2800余件,国际收入占比20%,真正实现了“创新走出去、品牌叫得响”。四年蝉联山东民营企业创新100强榜首,是鲁南制药创新实力的体现。在第二届临商大会开幕式上,张贵民作为“新临商”代表,系统阐述了企业以创新为核心驱动的发展战略,展现出这家扎根临沂57年的民族药企正全力冲向“千亿级”目标的决心与实力。在创新驱动中践行沂蒙精神,张贵民带领鲁南制药集团以敢为人先的锐气和永不服输的韧劲,持续推动医药产业高质量发展,展现出“新临商”的责任与担当。未来,在“健康中国”战略引领下,鲁南制药将继续以创新为引擎,向世界一流医药企业目标迈进,为全球健康贡献更多“中国方案”。
2026-05-19
2025年8月26日,由中华中医药学会遴选的2024年度中医药十大学术进展以题为 “Top 10 academic advances in traditional Chinese medicine in 2024” 在 World Journal of Integrated traditional and western Medicine(WJIM)发表。
通讯作者: 张伯礼院士、陈凯先院士、朱立国院士
合作作者: 张霄潇、郭继华、龚述坤、魏玮、荣培晶、刘凤斌、王卫庆、王计秋、贾振华、王伟、王勇、高月、黄璐琦、袁媛、郭娟、陈士林、徐安龙、王遥、徐志超、李梢、程翼宇、范骁辉、王毅、肖小河、陈香美、王伽伯、戴爱国、张俊华、张彤、高昊
单位:中华中医药学会、成都中医药大学药学院、中国中医科学院望京医院、中国中医科学院中医临床基础医学研究所、广州中医药大学第一附属医院、上海交通大学医学院附属瑞金医院、河北以岭医院、广州中医药大学、北京中医药大学东直门医院、中国人民解放军军事科学院军事医学研究院、中国中医科学院、成都中医药大学、北京中医药大学、东北林业大学、清华大学、浙江大学药学院、中国人民解放军总医院、首都医科大学、湖南中医药大学、天津中医药大学、暨南大学、上海中医药大学
张伯礼院士致辞
陈凯先院士致辞
朱立国院士致辞
摘要
发挥中医药原创资源优势,推动中医药研究实现创新突破是中医药传承创新发展战略的重要内核。当前中医药学术进展、科研成果已经成为引领中医药行业发展、推动中医药传承创新的重要支撑。为持续强化学术引领作用,中华中医药学会高度重视中医药领域的最新研究方向与学术成果,聚焦中医药学术发展的亮点与趋势,自2020年起开展“年度中医药十大学术进展”遴选和发布工作。本年度中医药十大学术进展以满足临床需求、回答科学问题、推动产业发展为目标,突出探索性与前瞻性、创新性与突破性的研究成果,经征集、整理和专家评审,遴选出10项本年度具有创新性和突破性的中医药学术进展,即“脑肠”跨器官策略防治消化系统疾病取得新进展,中医药防治心血管事件链获得高水平证据支撑,急性放射损伤中药防治研究取得重要突破,引种驯化导致西洋参“优形优质”变异的分子机制被阐释,中药组分苄基异喹啉类生物碱等微量活性成分生物合成,人工智能技术、中药盲区成分研究技术、新型递药系统在中医药领域示范应用,中药有效性评价整合证据链方法体系的创建与初步应用。
前言
为贯彻落实《中共中央 国务院关于促进中医药传承创新发展的意见》和全国中医药大会精神,定期梳理总结中医药领域研究成果,动态呈现中医药学术研究、创新发展的趋势,充分发挥科技社团的学术引领作用,中华中医药学会组织开展了2024年度中医药十大学术进展遴选工作。本年度遴选工作坚持“四个面向”,突出探索性与前瞻性、创新性与突破性,凸显在中医药临床研究、基础研究和应用研究领域取得的满足临床需求、回答科学问题、推动产业发展的新发现、新规律、新观点、新方法、新技术、新产品等,以此体现本年度研究成果在完善中医药传承创新发展机制,推动中医药事业和产业高质量发展的价值。经动态收集、初审、终审等工作程序,确定2024年度中医药十大学术进展。
2024年度中医药十大学术进展呈现出三个方面的特征:一是聚焦临床问题,体现中医药理论指导下,在重大疾病和难治性疾病防治方面的临床价值;二是阐明作用机制,为“说明白、讲清楚”中药治疗疾病的机理提供研究范式;三是鼓励学科交叉融合,推动新技术与新方法在中医药现代化研究中的应用。
结果
1.“脑肠”跨器官策略防治消化系统疾病取得新进展
消化系统难治性疾病以发病机制复杂、医治难度大、致死率及致残率高、严重影响工作生活为特征,以功能性胃肠病和消化道炎-癌转化疾病最为常见。其难治的关键点在于跨器官、跨脏腑,此类疾病的靶器官不仅在消化系统,且和中枢神经系统急、慢性应激损伤密切相关,是医学研究的难题。
1.1 消化系统难治性疾病“脑肠同调”中医药治疗新策略
1.1.1 研究背景
功能性胃肠病是一组由生理、精神心理及社会因素所致的以胃肠道症状为主的消化系统和(或)中枢系统功能紊乱疾病,以功能性消化不良和肠易激综合征最常见。脑-肠轴紊乱造成的消化系统和中枢系统共病是其共同病生理机制,临床实践和科学研究大多围绕单一疾病或系统展开,但以单一疾病为中心的研究范式正面临巨大挑战,不能满足此类疾病跨器官、跨脏腑的实际需求,需要寻求更为全面和整合的治疗策略。中国中医科学院望京医院魏玮教授、中国中医科学院中医临床基础医学研究所荣培晶研究员、广州中医药大学第一附属医院刘凤斌教授,在传承国医大师路志正先生“持中央、怡情志”理论的基础上,针对本类疾病共同病生理学机制“脑肠互动异常”,提出核心病机为“神明之枢失衡(脑)”、“胃肠腑气不通(肠)”,创新了脑肠同调治则治法,创建了“脑-肠”多维度疗效评价新体系。
1.1.2 研究方法与结果
在脑肠同调治则指导下,证实脑肠同调治疗功能性消化不良、腹泻型肠易激综合征临床有效。使用辛开苦降调枢法能够通过辛开苦降、疏肝解郁来治疗功能性消化不良(FD),经998例临床研究显示可同时改善患者精神心理状态及腹部不适;使用温肾健脾调枢法对692例焦虑抑郁患者的临床研究证实可同时改善患者焦虑抑郁状态及腹部不适、腹泻症状,降低复发率;对于症状明显情绪相关患者,采用经皮耳迷走神经刺激及针刺外治法治疗,188例临床研究显示脑肠同调外治法可增强迷走神经活性,以缓解患者消化不良症状、焦虑抑状态,并可改善患者脑区功能。
揭示脑肠同调治则治法多靶点调节脑肠轴的疗效机制。应用蛋白组学、肠道微生态检测等多组学技术结合,发现辛开苦降调枢法可改善肠道菌群、胆汁酸代谢及胃泌素、胃动素、胰血糖素样肽、5-羟色胺等脑肠肽水平,温肾健脾调枢方可调节下丘脑及结肠肾上腺激素释放因子系统的失衡状态,针刺通过迷走神经-胆碱能抗炎通路调节脑默认网络功能状态,产生改善内脏高敏和抗抑郁的双重效应,共同阐释了脑肠同调治疗策略的科学内涵。
创建了功能性胃肠病“脑-肠”多维度疗效评价新方法。首次基于“概念-领域-方面-条目”四级结构,创制了脑肠中西医诊疗评价量表体系;建立了“中华生存质量量表”、“中医健康状况量表”、功能性胃肠病、肠易激综合征医生报告的症状量化推荐表单,完整构建了功能性胃肠病医生、患者报告结局复合评价体系,经3475例临床研究显示本结局复合评价体系具有良好的信度、效度。
1.1.3 研究价值
该研究为消化系统难治性疾病提供了新治疗策略、理论基础与科学证据支持,发挥了中医药整体调节特色优势,提高了消化系统难治性疾病诊疗水平。在中医整体观指导下,提出的“脑肠”跨器官治疗消化系统难治性疾病的新策略,也为重大慢性疾病的诊疗理念提供了新思路。
1.2 葛根素靶向“脑肠轴”减肥机制被揭示
1.2.1 研究背景
全球范围内,高脂肪和高热量食物的消费增加显著促进了肥胖和代谢性疾病的流行。随着生活改善,我国居民饮食中脂肪占比已达32.9%,高能量饮食会增加肠道能量吸收以及脂质蓄积,导致脂代谢异常,最终引起肥胖。同时因肥胖导致的死亡率也在逐年增加,因此肥胖已成为世界范围内一个主要的公共健康问题。但目前的主流观点认为肠道脂质的吸收主要受肠道吸收表面积、膜两侧浓度差等因素的影响,是器官自主的过程,此前尚未有确凿证据表明肠道吸收受到中枢神经系统的调控,其是否受其他途径调控长期被忽略。
1.2.2 研究方法与结果
发现小肠油脂吸收是受到大脑的直接调控。上海交通大学医学院附属瑞金医院研究团队整合化学遗传学、神经电生理、分子探针等研究策略,揭示小肠油脂吸收的脑-肠轴调控机制,发现迷走神经背侧核(DMV)在肠道脂肪吸收中起到关键作用,即DMV的失活能缩短绒毛长度,减少肠道脂肪吸收。通过灭活小鼠投射到十二指肠或回肠的DMV神经元(膈下迷走神经切断术),发现其能够起到减轻体重、减少脂肪吸收、降低血浆甘油三酯(TG)水平和增加粪便脂肪排泄的作用。这些结果表明,投射到空肠的DMV神经元可以直接调节脂肪吸收和体重增加。
发现葛根素具有抑制DMV活性的作用,导致空肠脂肪吸收的减少。研究者对药物分子进行筛选,以求寻找到可以特异性抑制DMV活性的药物分子,将药物治疗和电生理相结合,对小鼠的脑干切片进行了研究。中药单体葛根素在总共22个检测到的神经元中,有16个(72.7%)被抑制活性,随后在动物实验中腹腔注射葛根素之后,与DMV失活后的表型结果一致。结果表明葛根素可以模拟“DMV-迷走神经-肠上皮细胞-脂类吸收”的抑制效果,减少吸收表面积、抑制小肠对油脂的吸收。
证实了葛根素的作用机制。研究团队通过脑片电生理筛选小分子化合物,借助化学蛋白组学探针、结构生物学等策略并结合动物模型,明确葛根素作用在DMV的分子靶点——GABA受体。借助迷走神经背核Gabra1DMV敲除的动物模型,进一步明确了Gabra1DMV对糖脂代谢的调控作用,为代谢性疾病的干预治疗提供了新靶点。通过获得葛根素与GABAAR受体互作复合物的2.4Å超高分辨率的结构图像,解析了葛根素与GABAAR结合的关键位点,精确观察到葛根素对受体通道通道孔径的效应,结合电生理结果共同揭示葛根素抑制DMV神经元的机制。为进一步确认GABAAR(含α1亚基)介导了葛根素的排油减肥功能,构建了迷走神经特异性敲除α1亚基(Phox2b-cre;Gabra1f/f)的动物模型,发现敲除小鼠在高脂情况下体重明显增加,且对葛根素无反应(排油减肥功效消失),从而充分论证了葛根素通过GABAAR受体发挥排油减肥作用。通过大量的形态学分析,发现在DMV抑制或葛根素干预后,空肠微绒毛长度会明显缩短,是导致小肠油脂吸收能力降低的主要原因。
1.2.3 研究价值
该研究将既往认为小肠油脂吸收是“自主发生的被动扩散过程”更新为“受迷走神经精密调控”,完善了小肠营养吸收的知识体系,为肥胖症的干预治疗提供了脑-肠轴新途径。该工作跨越神经、消化、代谢等领域,涵盖药物遗传、神经电生理、光交联反应、冷冻电镜及模式动物等多种前沿技术,发现了一个新颖的脑-肠轴通路,可调节肠道油脂吸收,这为未来的诸多研究开辟了新方向。
上下滑动,查看更多
2.中医药防治心血管事件链获得高水平证据支撑
心血管事件链指以动脉粥样硬化为基础的心血管疾病由高危因素聚集导致动脉硬化,易损斑块破裂引起心肌梗死,出现心律失常、心力衰竭直至死亡的全过程,呈现多因素、多环节作用下因果相连、递进发展、事件突发、后果严重的病变特点,反映了心血管疾病整体性与复杂性的系统特征,带来防治理念由单因素、单环节、单靶点干预,向整体、连续、动态、全程干预的系统思维转变。目前单靶点、单环节药物无法满足临床需求,缺乏针对心血管事件链因果相连、递进发展病变特点的系统干预措施,如何突破这一瓶颈难题是实现心血管事件链系统干预的重大课题。
2.1 心血管事件链系统干预新策略构建及应用
2.1.1 研究背景
基于中医营卫理论“营卫不通、血凝不流”,“血脉相传、壅塞不通”之“凝”→“壅”→“塞”→“不通”血脉传变规律,与心血管事件链关键病理环节相关性,提出“治本病、防未病”心血管事件链系统干预新观点—防上游因素、治当前病变、控下游传变。基于古今医案多维数据挖掘,传承创新营卫理论揭示事件链“凝”→“壅”→“塞”→“不通”传变病机特点,总结“调营卫津血”用药规律,创新诠释《难经》“损其心者,调其营卫”治则,形成心血管事件链系统干预临床用药方案。基于心血管事件链关键病种构建多源医案数据库,优化多维数据挖掘方法,基于“症状集合形成证候、证候聚类反应病机、病机演变揭示传变、用药规律加以反证”思路,揭示系统干预证治规律。建立高质量循证医学证据,结合基础实验研究及网络整合分析,阐明“治本病、防未病”科学内涵。
2.1.2 研究方法与结果
高质量循证研究取得中医药系统干预心血管事件链重大突破。河北以岭医院贾振华教授联合仝小林院士、杨跃进教授、李新立教授、黄鹤教授、卜培莉教授团队开展中医药防治心血管事件链的系统研究。津力达预防2型糖尿病取得中医药突破性进展,对代谢综合征糖耐量异常889例临床研究显示,津力达显著降低糖尿病发生风险41%,系统调节多代谢异常保护血管,延缓动脉硬化;参松养心解决房颤导管消融术后复发难题,优化了药物治疗方案,参松养心对导管消融房颤患者复发影响920例临床研究显示,参松养心组降低消融术后1年内房颤复发风险,减轻房颤负荷,提高生活质量;芪苈强心改善心衰预后取得中西药联合治疗新突破,芪苈强心对慢性心衰复合终点事件影响3119例临床研究显示,芪苈强心改善心衰患者预后,降低心血管死亡和心衰恶化再住院发生风险。
基础研究通过整体、器官、组织、细胞、分子揭示系统干预心血管事件链疗效机制,网络整合分析揭示其关键环节、机制、通路、靶点,阐明“治本病、防未病”系统干预科学内涵。津力达干预“凝”—高危因素:防上游因素—激活FGF21/AMPK通路,促进脂肪利用;激活PPARγ、PGC-1α促进脂肪产热,调节脂肪功能;治当前病变—降低肥胖小鼠体重,改善胰岛素抵抗,调节糖脂代谢;控下游传变—改善肥胖小鼠内皮功能,促进巨噬细胞M1型向M2型极化,抑制动脉硬化。参松养心干预“不通”—心律失常:防上游因素—调节PI3K/Akt等通路,改善代谢异常、缺血对房颤影响;治当前病变—延长有效不应期,降低房颤易感性,逆转电重构;控下游传变—抑制心房纤维化,逆转结构重构,改善心功能。芪苈强心干预“不通”—心衰:防上游因素—激活PPARγ、PGC-1α,抑制心梗后心肌细胞凋亡,减轻病理性心肌肥大;治当前病变—改善苯肾上腺素/异丙肾上腺素诱导及心梗后心肌纤维化和心室重构;控下游传变—提高射血分数,降低ANP、BNP,改善心功能。
2.1.3 研究价值
首次提出“治本病、防未病”事件链系统干预新观点—防上游因素、治当前病变、控下游传变,传承创新营卫理论揭示事件链“凝→壅→塞→不通”中医传变病机特点,总结“调营卫津血”用药规律,奠定系统干预理论基础。该研究阐释了中医药在防治心血管事件链的独特优势和疗效,为中医药防治重大疾病研究提供了研究范式。
2.2 证候理论指导下的心衰新药发现与创新机制研究
2.2.1 研究背景
心力衰竭(心衰)是心血管疾病的终末阶段,死亡率和再住院率居高不下,临床仍缺乏高效治疗手段。近年来,多项随机对照试验表明,中药复方可显著改善心衰症状且安全性良好,中医药可能成为突破心衰治疗困境的新途径。然而,传统中医证候体系难以全面涵盖现代心衰患者的复杂特征,例如慢性心衰反复发作、迁延难愈的“瘀久化毒”证候是否成立?同时,现代医学研究发现,线粒体能量代谢关键调控蛋白——类视黄醇X受体α(RXRα)是改善心功能的潜在靶点,但缺乏高选择性激动剂。因此,该研究旨在创新心衰中医证候体系,明确“毒邪”的临床与生物学基础并从复方芪参颗粒中筛选靶向RXRα的活性成分,探索心衰治疗新策略。
2.2.2 研究方法与结果
总结心衰“毒邪”证候。广州中医药大学王伟教授、北京中医药大学东直门医院王勇教授携团队总结了心衰气虚血瘀证瘀久化毒导致的瘀毒证候,采用真实世界研究与前瞻性研究结合的策略,在益气温阳活血治法中融入心衰解毒新治法,研发出创新中药芪参颗粒。确立了心衰“毒邪”诊断标准,推动了中医证候现代化发展;并研发的芪参颗粒被证实可显著改善心功能、能量代谢及心肌纤维化,目前已获得新药临床批件并完成技术转让。
揭示黄芪甲苷Ⅲ(AS-Ⅲ)靶向RXRα,保护线粒体功能,发挥心脏保护作用。黄芪甲苷是中药黄芪中的主要有效成分,是一类对心血管疾病具有治疗和药理作用的化合物。据报道,几种黄芪甲苷可以改善线粒体功能,包括清除自由基和增强ATP含量。其中,黄芪甲苷IV已被观察到调节腹膜纤维化中的RXRα,表明黄芪甲苷家族的化合物具有激活RXRα并改善线粒体功能的潜力。通过首创“四位一体”斑马鱼胚胎模型药物筛选体系,结合表面等离子共振(SPR)、微量热泳动(MST)技术,筛选出芪参颗粒中的关键活性成AS-III是黄芪甲苷家族中与RXRα亲和力最高的化合物。体内外实验均证实,AS-Ⅲ通过一种独特的机制与RXRα结合,其中AS-Ⅲ停靠在同源二聚体的界面上,随后引起RXRα同源二聚体的反式激活,有助于改善线粒体损伤和心功能障碍。
首次报道RXRα转录调控NADH脱氢酶(泛醌)铁硫蛋白4(Ndufs4),依赖于RXRα同二聚体及其配体激活。该研究进一步揭示了RXRα依赖于配体激活并以其同源二聚体形式直接调节Ndufs4的转录,即AS-Ⅲ激活RXRα同源二聚体,随后激活Ndufs4转录,从而促进线粒体功能和ATP的产生。体外内实验均证实,AS-Ⅲ被证实为RXRα的配体,可以激活RXRα同型二聚体,进而上调Ndufs4,促进心脏线粒体功能,发挥心脏保护作用。
2.2.3 研究价值
该研究创新性地提出“毒邪”概念,丰富了心衰中医诊疗体系,并开创性地将“益气温阳活血”与“解毒”相结合形成新治法,有望成为心衰治疗的新途径,为临床实践提供了全新思路。基于“斑马鱼胚胎-斑马鱼成鱼-大小鼠-小型猪”的“四位一体”心衰在体药物筛选系统,为心衰药物发现提供了新的研究策略与平台。
上下滑动,查看更多
3.急性放射损伤中药防治研究取得重要突破
3.1 研究背景
近年来,民用核技术的广泛应用使民众辐射暴露风险增加,加速急性放射损伤防治药物研究对提升核应急救治能力、完善国家核安全战略体系意义重大。然而,现有急性放射损伤防治药物多为化学药物,疗效有限且副作用明显,存在极大应用限制。中药在辐射防护领域逐渐受到关注,在急性放射损伤防治药物开发中呈现巨大潜力。军事科学院军事医学研究院高月研究员团队基于放射损伤动物模型与体外类器官、细胞培养体系,对骨髓型、肠型与皮肤型放射损伤防治中药及其机制进行了探究。
3.2 研究方法与结果
针对骨髓放射损伤,首次发现火麻仁油可大幅提升致死剂量辐照动物30天生存率。选定其单体成分大麻二酚作为研究对象,基于生存率、造血细胞数量、造血干细胞功能与损伤指标,评估了其急性放射损伤防治功效;利用单细胞RNA-seq解析了大麻二酚致造血干细胞功能变化的分子基础;对重要靶标分子进行了预测,并基于体外培养体系进行了功能验证。研究发现:火麻仁油可大幅提升致死剂量辐照动物生存率,并对造血与免疫细胞具有保护功效;其所含成分大麻二酚可提高急性放射后个体生存率、短期内降低造血干细胞氧化应激水平、促进造血干细胞功能恢复;大麻二酚显著恢复了放射损伤造血干细胞Atf2的表达,其下游调控Lrp6等基因表达,从而促进造血干细胞干性恢复;大麻二酚在体外培养过程中亦可促进造血干细胞扩增与功能维持。
针对肠放射损伤,凉血固元益肾汤干预能够促进肠损伤修复,改善辐照动物生存率。利用16s RNA测序与组化方法,检测了高剂量辐射导致的肠道微生物组时序变化、肠屏障损伤等指标;挖掘了AKK菌作为影响损伤响应关键菌群,其丰度在凉血固元益肾汤干预后得到恢复;基于菌体外培养与移植方法,评估了AKK菌改善肠型放射损伤与生存时间的作用;分析了凉血固元益肾汤入血成分、AKK菌产生的短链脂肪酸对肠类器官生长的促进作用。研究发现:凉血固元益肾汤有效增加辐照动物肠道内AKK菌丰度、提高肠道异丁酸水平,促进肠损伤修复,改善辐照后生存率,靶向AKK菌和肠上皮保护,筛选相关活性成分,发现在中、重度肠型急性放射病救治方面疗效显著。
针对皮肤放射损伤,阿魏酸水凝胶显著降低皮肤炎性与氧化应激水平、减少血流,从而促进辐射所致皮肤损伤的恢复。通过卡波姆940交联法研制了阿魏酸水凝胶,通过Calcein-AM/PI染色、CCK8染色、溶血试验等验证了其生物相容性;基于SS-OCT技术评估了水凝胶对皮肤急性放射损伤的治疗效果;结合分子实验对水凝胶抗炎功效展开机制解析。研究发现:在良好稳定性与生物相容性基础上,不同浓度的阿魏酸水凝胶通过抑制NLRP3炎性小体活性,显著降低皮肤炎性与氧化应激水平、促进血管生成,从而促进皮肤放射损伤的恢复。
3.3 研究价值
围绕核辐射所致关键组织急性放射损伤,基于中药发掘了系列防治候选药物,极大丰富了核应急药物储备,有望突破中、重度急性放射病无好药困境,开拓了中医药治疗新领域,可为国家核安全应急响应体系建设提供“中医药”选项,助力维护国家安全。
上下滑动,查看更多
4.引种驯化导致西洋参“优形优质”变异的分子机制被阐释
4.1 研究背景
目前已有超过100种大宗中药材实现了从野生到人工栽培的转变,然而与野生药材相比,栽培药材在性状和功效成分上可能因驯化变异而发生变化,进而影响药材的质量。因此如何破解中药材引种驯化的生物学本质,进而评价中药材质量,培育高品质中药材对中药产业高质量发展至关重要。西洋参(Panacis Quinquefolii Radix)为五加科人参属植物西洋参(Panax quinquefolium L.)的干燥根,具有补气养阴、清热生津之功效,原产北美,我国于20世纪70年代引种成功,目前已成为全球西洋参三大主产国之一,与野生西洋参相比引种驯化导致西洋参的侧根数减少,人参皂苷Rg1含量降低。
4.2 研究方法与结果
发现西洋参根系形态建成的关键因子。中国中医科学院黄璐琦院士院士和袁媛研究员团队提出道地药材应具有“三优”特征,拓宽和深化了传统药材“辨状论质”理论。在此基础上,研究团队利用PacBio三代测序数据完成野生西洋参基因组组装,并通过比较基因组学分析,鉴定西洋参中的扩张和收缩的基因家族,发现与三七相比,西洋参和人参中1,3-β-葡聚糖合酶(1,3-β-glucan synthase)、纤维素合酶(cellulose synthase)、莽草酸羟基肉桂酰转移酶(shikimate O-hydroxycinnamoyl transferase)和细胞膨胀蛋白(expansin)等与细胞壁发育相关的基因家族显著扩张,推测细胞壁发育是调控西洋参根系形态建成的关键因子。通过收集26株野生西洋参和30株栽培西洋参进行全基因组重测序,分析野生和栽培西洋参群体的遗传结构差异,并采用选择清除分析筛选野生和栽培西洋参群体中的受选择的与植物细胞壁发育相关的Expansin-like细胞膨胀蛋白基因(Pq17G51776)和β-半乳糖苷酶基因(Pq06G18190)。
验证西洋参“优形优质”特征变异的关键基因。为进一步探究Expansin-like亚家族基因调控西洋参根系生长发育的功能,分别通过西洋参expansin-like基因的过表达(overexpression,OE)和RNA干扰(RNA interference,RNAi),利用基因枪法建立转基因西洋参愈伤组织系。并在西洋参愈伤组织中分别过表达和敲低了Expansin-like A和Expansin-like B亚家族的7个基因,结果显示PqEXLA1可显著提高愈伤组织的生长速率,促进细胞体积变大;PqEXLB1可显著促进西洋参细胞体积变大和细胞壁厚度增加。通过RT-qPCR和UPLC-MS/MS分析,发现PqEXLA1和PqEXLB1的转录水平与人参皂苷Rg1的合成存在拮抗关系,表明PqEXLA1和PqEXLB1作为平衡细胞生长和Rg1合成中的关键枢纽,在西洋参引种驯化过程中通过调节植物生长与防御的平衡,进而导致西洋参“优形优质”特征变异。
4.3 研究价值
该研究提出气候主导型、生产措施主导型、种质主导型等道地药材“优形优质”特征形成模式,建立药用模式植物综合研究平台;提出具“内稳态”特征(稳态的外观形状和代谢谱)的中药材种质创新和栽培策略,为高品质中药材新品种选育和绿色生产提供新理论和新范式。
上下滑动,查看更多
5.应用一体多维数据平台生物合成中药微量活性成分取得新进展
5.1 研究背景
中药活性成分是天然药物挖掘的宝库,但大部分中药活性成分含量低且化学合成困难,限制了其深入挖掘和利用。以合成生物学为代表的生物技术为中药活性成分或微量成分的合成提供了新的机遇和发展空间,为中药活性成分绿色、可持续的新型中药现代化经济体系提供方法,从而实现中药资源的可持续利用。
5.2 研究方法与结果
构建“数据—元件—功能—生产”一体多维平台,开展高药用价值中药活性成分的生物合成学研究。中国中医科学院黄璐琦院士和郭娟研究员团队通过整合中药植物遗传数据资源,搭建了中药多维度组学数据库,构建了库容1000+的中药活性成分生物合成实体元件库,创建了“多组学—多体系”联合分析的基因功能研究技术体系,形成了一体多维、交互融合的中药活性成分生物合成研究平台。利用差异代谢物、差异表达基因、蛋白质与DNA/RNA的结合区域、微生物群落的特异性以及基因本体(GO)相似性分析,在不同实验条件下比较代谢物、基因、基因组区域和微生物类群等数据需求,开发交互式韦恩图、欧拉图、UpSet图、花瓣图和交互式花瓣图等专门的可视化技术平台,允许直观识别共性与差异,清晰地展示多组数据关系,为中药微量活性成分生物合成途径途径解析提供分析方法。
通过多种策略从头合成多种中药微量活性成分。在酿酒酵母、大肠杆菌等微生物中开展丹参酮、雷公藤甲素、冰片、穿心莲内酯等高药用价值中药活性成分的生物合成学研究,构建了高效合成的细胞工厂,实现了中药微量活性成分的生物合成。搭建中药天然产物合成途径元件挖掘和合成生物学利用的合作平台,为中药天然产物尤其是苄基异喹啉类生物碱的可持续生产提供新的方式。
5.3 研究价值
该研究构建了中药活性成分生物合成“数据-元件-功能-生产”一体多维平台,将会从基础数据层面促进药用植物分子代谢途径的解析,实现萜类、生物碱类等代表性中药微量活性成分高效异源生产,为中药微量活性成分绿色、可持续的新型中药现代化经济体系提供范式,进而在推动合成生物学的发展、促进药物发现和药物生产的天然来源探索方面发挥重要作用。
上下滑动,查看更多
6.中药组分苄基异喹啉类生物碱生物合成途径与抗病毒机制被解析
6.1 研究背景
我国是呼吸道病毒感染性疾病的高负担国家,急需研发应对新发突发呼吸道病毒性传染病的储备药物。中医药在新冠疫情防控中发挥重要作用,其中千金藤素等苄基异喹啉类生物碱(BIAs)因其显著的抗新冠病毒活性受到国际关注。然而,BIAs化合物具有结构多样、在基原植物中含量低、立体构型复杂、化学合成难度大的特点,严重制约了其药物发现及资源的可持续利用。解析BIAs的生物合成途径,以指导其异源合成和高含量种质的选育,是解决资源瓶颈的重要策略。此外,BIAs抗病毒机制不清,限制了其在中药创新药物研发及精准临床应用中的潜力。
6.2 研究方法与结果
解析双苄基异喹啉生物碱(BBAs)生物合成途径。成都中医药大学陈士林院士联合北京中医药大学徐安龙、王遥教授和东北林业大学徐志超教授团队率先创建本草基因组学学科体系,提出了“千种本草基因组计划”,辅助育种培育优良中草药品种,并利用合成生物学技术生产重要天然药物或新药原料。以富含双苄基异喹啉生物碱(BBAs)的中药千金藤和北豆根为研究对象,完成了中药千金藤和北豆根等基原植物的染色体水平高质量基因组图谱构建,通过代谢组学手段鉴定了一系列BIAs化合物的上下游代谢产物,推测其潜在的生物合成途径。发现防己科物种CYP80基因的显著扩张驱动了BIAs结构多样性的形成。此外,鉴定了一条全新的BIAs生物合成通路,发现高效的氧甲基化、C-C连接酶等,建立“一锅法”烟草底盘合成体系,为BIAs化合物的资源替代及药物开发提供了重要参考。
揭示了千金藤素及其结构类似物的广谱抗冠状病毒潜力。聚焦中医药免疫代谢调控,通过生物化学、分子生物学、遗传学和生物信息学手段,结合体内外实验证明BBAs重塑细胞抗病毒免疫代谢以抵抗多种病毒感染的科学原理。鉴定出北豆根中BBAs抗病毒感染的关键靶点NPC1,证明BBAs抑制NPC1并重塑细胞“免疫代谢”,实现直接干预病毒入侵(基于脂质代谢)和激活抗病毒免疫(宿主免疫因素)抑制病毒的协同作用。揭示细胞核胆汁酸受体FXR是限制胆源中药抗病毒药效的关键免疫代谢因子,阐明胆粉与植物甾醇“配伍”提升抗病毒免疫药效的分子机制。发现了广谱有效、活性明确、安全性高的抗病毒候选药物汉防己甲素、千金藤素北豆根总生物碱、小檗胺等。
6.3 研究价值
该研究破解了千金藤、北豆根等中药的高质量基因组与代谢组,解析复杂BIAs的生物合成途径,为规模化生物制造奠定基础。同时筛选出高效、安全的广谱抗病毒候选药物,阐明其通过调控宿主免疫代谢靶点抑制病毒复制的药理机制,为应对新发突发病毒疫情提供了药源储备与干预新策略,为中医药现代化及抗病毒创新药研发提供了重要范式。
上下滑动,查看更多
7.人工智能技术在中医药领域示范应用取得新进展
随着生物医学研究进入人工智能(AI)时代,如何运用AI前沿技术,科学解读中医药复杂体系原理,发掘中医药在重大疾病防治上的特色理论和实践经验,提升重大疾病的中医药防治水平,既是中医药、人工智能和信息科学共同面临的关键科学问题,也是中医药传承创新发展和“健康中国”重大战略实施的重大挑战。在中医药现代化过程中,囿于研究技术、研究策略的局限,较难从分子水平诠释中药的药效物质及其关键作用靶点与关键细胞群落,亟需技术方法的突破。
7.1 中医药+AI”关键技术及其在胃癌防治的示范应用
7.1.1 研究背景
中国是全世界受肿瘤困扰最严重的国家之一。以胃癌为例,我国每年新发胃癌病例就约占全球的43.9%。早期有效防治对于降低肿瘤发生率至关重要,其核心挑战在于如何精准预警肿瘤发生风险并给予有效干预。中医强调“未病先防、既病防变”,在肿瘤防治上具有整体观、治未病等特色理论与丰富的临床实践经验。随着生物医学研究进入人工智能时代,如何运用AI前沿技术,深入挖掘中医药在肿瘤防治上的特色理论与实践经验,形成中西医融合的肿瘤防治新范式?这既是中西医学面临的共性难题。
7.1.2 研究方法与结果
建立中西医药分子网络导航系统(UNIQ系统),实现中医药复杂体系原理的全景式解析。清华大学教授、欧洲科学与艺术学院李梢院士团队自主研发UNIQ系统,构建网络表征基座模型和智能决策基座模型,实现中西医“表型—细胞—分子—中西药物”多层次网络关联的全景式解析。UNIQ系统具有计算精度高、数据质量优、发现能力强、转化效率高的突出优势基于UNIQ系统的规模化应用,制定网络药理学应用于中药新药研发专家共识总论。其中,网络表征基座模型能够建立中药物质数据、细胞与分子多层次数据、病证临床数据等多模态大数据的网络关联,从而实现中药与病证关系的精准定位与智能导航,从全局上解析中药复杂系统;智能决策基座模型利用基于生物网络的反馈校正学习,将中药研发过程中的实验与临床验证结果作为计算模型的训练知识,实现计算-实验-临床的迭代优化,通过智能决策,显著提升中药新药研发的针对性和精准度。
构建胃癌极早期中西医智能与精准防治体系。基于UNIQ系统平台,发展了中西医多模态多组学AI算法(AI-TWM),系统采集与智能解析50余万例胃炎癌转化中西医临床数据、以及典型序贯病例的多组学数据,构建了胃炎癌转化“表型—细胞—分子—药物”等多层次生物网络。采用网络药理学全面预测和验证,发现靶向胃癌极早期网络的药食同源中药,经RCT临床试验验证能显著阻断胃炎癌转化,并系统解析肿瘤发生相关中西医多模态大数据中的诊疗规律,实现肿瘤发生风险的智能预警以及防治中药的精准发现。率先提出“胃癌极早期”这一表征胃炎癌转化临界状态的全新分期,并发现胃癌极早期的中西医特征、病证标志物和防治中药,示范构建胃癌极早期中西医智能与精准防治体系。
7.1.3 研究价值
该研究构建胃癌极早期中西医智能与精准防治体系,以智能预警、极早诊断、精准早治为特色,破解了胃癌早期防治难题,是中医药防治胃癌的“突破性”发现,显著提升胃癌中医药防治水平。该体系已在中国多个胃癌高发区及其医院推广应用,取得显著的防治效果,有望形成具有中国原创特色的肿瘤防治新模式,为AI时代肿瘤早期中医药防治研究提供了新视角与新思路。
7.2 人工智能新技术助力中药作用机制解析
7.2.1 研究背景
近年来,表型组学、单细胞组学等新兴技术在中医药现代研究中得到广泛应用,大量组学数据的不断累积也对解析中药作用下细胞形态及细胞组成的时—空细微变化提出了更大挑战。中药作用于机体组织中哪些细胞起效?能否基于单细胞空间转录谱、细胞结构的相似性预测中药潜在作用靶标?回答这些科学问题亟需创新单细胞表型组、转录组等多组学数据分析工具。
7.2.2 研究方法与结果
构建了生成式AI驱动的组学数据模拟框架scCube,实现了中药活性成分的效应细胞类型和潜在靶点的发现。浙江大学药学院程翼宇教授、范骁辉教授、王毅教授研究团队将AI技术引入中药科学原理的系统解析研究,提出组学数据模拟框架scCube,针对真实组学数据存在批次效应和标注偏差等瓶颈,利用变分自编码器深度学习模型建模真实数据中的多种变异性,并生成无偏模拟数据,为相关多组学数据分析方法的开发提供准确且多样化的基准测试数据支持。基于scCube生成的模拟基准测试数据创建并优化了scRank算法,利用靶点扰动的基因调控网络(tpGRN)整合生物网络和药物靶点信息对药物扰动进行建模,实现了从单细胞转录组数据中推断出中药活性成分的效应细胞类型和潜在靶点,成功解析了活血中药丹参中丹参酮IIA作用于缺血心肌组织的巨噬细胞亚群MΦ5,而CTSB是其潜在作用靶点。
构建全球首个心肌细胞线粒体表型的高内涵动态分析方法,实现了细胞筛选模型智能辨识药靶的技术突破。基于单细胞表型组数据的靶点预测算法MitoReID,为捕获中药作用下细胞形态的时—空细微变化并识别相应效应分子靶点,建立了心肌细胞线粒体表型的高内涵动态分析方法,大规模采集已上市小分子化学药作用下的线粒体表型时序图像,构建了涵盖1068种FDA批准小分子药物、包含超过57万张单细胞图像的数据集。随后,基于scCube生成的模拟基准测试数据创建并优化了MitoReID深度学习模型,利用信任重识别框架提取待测样本作用的线粒体形态、膜电位等时序图像特征,并通过匹配查询中药活性成分与已上市小分子药物的线粒体表型特征来推断黄芩汤等方剂中主要成分的作用机制,发现表儿茶素直接结合COX-2抑制其活性发挥抗炎作用。通过构建全球首个药物作用下的细胞线粒体动态表型大规模数据集(逾57万张单细胞形态图像),研究发现FDA批准已上市药物的靶点与线粒体表型具有相关性,由此可依据药物高内涵筛选图像推测出其作用靶标,实现了细胞筛选模型智能辨识药靶的技术突破,进而发明了一种基于线粒体表型的新药AI筛选技术。
7.2.3 研究价值
该研究突破了传统组学技术的效率低和分辨率不足等缺陷,在中药药效物质发现、作用网络构建和整合调节机制阐明等方面有着巨大的应用前景,为“说清楚、讲明白”中药临床功效的科学内涵提供了新的技术工具和研究示范。该方法整合多模态表型组数据为药物成分发现与深入解析中药效应细胞与作用靶标提供了新方法,不仅为老药新用提供了重要的技术支持,也为应用AI技术加速科学发现提供了新范例。
上下滑动,查看更多
8.中药有效性评价整合证据链方法体系的创建与初步应用
中医药为人类的健康事业做出了巨大贡献,但其疗效在国际范围内却未得到广泛认可。究其原因,是在以RCT为唯一“金标准”的循证医学体系下,中医药难以获得高质量证据支持,致使一大批疗效卓著的中医疗法和方药的有效性证据难以真实地反映其客观疗效和临床价值,亟待建立一套符合中医药特点且被国际认可的有效性评价方法。
8.1中药有效性评价新策略新方法:整合证据链法
8.1.1 研究背景
为了突破长期制约中医药事业发展的临床疗效评价关键技术这一“卡脖子”难题,解放军总医院第五医学中心肖小河教授领衔团队近年来在系统分析和反思当今中医药研究特别是中药有效性评价现状和问题的基础上,提出应从底层逻辑出发,深度体悟中西医药的认知差异,特别是中西医从物质组成内涵、生产研发路径、临床诊疗理念到临床疗效评价不同层面存在的倾向性乃至本质性差异,多方位思考由中医药特质性带来的疗效评价方法学挑战与对策,探索建立一套真正符合中医药特点并融合现代循证医学思想的中药有效性评价策略和方法体系,从而为促进中医药传承创新与高质量发展提供具有战略性和全局性的科技支撑。
8.1.2 研究方法与结果
首次提出并建立中药有效性评价新策略和方法体系——整合证据链法(iEC-Eff)。研究团队在底层逻辑认知创新的基础上,结合和发挥中医药“临床—实验室—临床”双向转化医学研究的特色优势,创制了中药有效性评价新策略和方法体系——“整合证据链法(integrated evidence chain-based effectiveness evaluation of traditional Chinese medicine,iEC-Eff)”。iEC-Eff法整合了临床经验(初证)、实验研究(佐证)和临床试验(确证)等方面的证据要素,同时注重吸纳生物信息学、人工智能、大数据分析等研究手段,原创性地建立了多环节一体化的综合评价与确证模式和方法体系。具体来说,针对某一方药,其疗效证据链由3个环节构成:既往应用经验可以提供初步的疗效证据(初证),多模型多团队实验研究可为其提供佐证性的疗效证据(佐证),临床试验可提供确证性的疗效证据(确证)。有效性证据链越完整,每个链条上的证据级别越高,则支撑其有效性的证据力越强,其临床疗效的可信度越高。iEC-Eff法不仅充分体现了中医药双向转化医学优势,而且符合人们认识事物本质的“实践-认识-再实践-再认识”基本规律,能够更客观全面地反映中药的临床价值和地位,有望发展成为具有突破性和引领性的中药有效性评价新方法和新策略。
“整合证据链法”的示范性研究与转化应用。首先对iEC-Eff的可行性和评价效能进行了示范性应用研究,通过分别采用iEC-Eff法和目前国际主流的GRADE法,对六味地黄丸改善2型糖尿病症状、藿香正气方治疗胃肠型感冒、安宫牛黄丸改善急性脑缺血卒中、通心络改善心肌梗死等相关症状的有效性进行了评价。结果显示,上述药物GRADE评价证据等级均为“低级”或“极低级”,而iEC-Eff法评价证据等级为“高级”或“高级+”。结果表明,iEC-Eff法可以更全面地反映了中药有效性和临床价值,使中药特别是经典名方的有效性评价证据等级得到科学合理提升。进一步针对肝纤维化和早期肝硬化这一国际治疗难题,且中医药有潜在治疗优势的病种,采用iEC-Eff法,对大黄䗪虫丸、鳖甲煎丸、复方鳖甲软肝片、扶正化瘀胶囊、安络化纤丸和和络舒肝胶囊等临床常用的6种抗肝纤维化中成药进行了有效性评价。结果显示,整合证据链法能使治疗肝纤维化类中药的有效性证据级别得到合理的提升,鳖甲煎丸、大黄䗪虫丸等经典方药均从GRADE评价的D(极低级)提升到了iEC-Eff评价的AAB(高级),复方鳖甲软肝片、扶正化瘀胶囊从GRADE评价的B(中级)提升到了iEC-Eff评价的AAA(高级)。iEC-Eff法评价结果与行业规范性文件的认可度吻合性好,反映了iEC-Eff法具有良好的评价效能。
8.1.3 研究价值
该方法突破了中药有效性评价以RCT为唯一金标准的局面及局限,为破解中医药有效性评价难题提供了具有原创性的研究思路和可操性的工具方法,科学提升了一大批中医药经典方药的有效性评价证据等级,使其更客观地反映其临床价值和地位。
8.2 构建中医药疗效证据链与评价方法
8.2.1 研究背景
随着中医药现代化研究的发展,有关中医药临床疗效及作用机制的研究证据快速增长。然而,由于缺乏相关的方法学指引,中医药临床与基础研究脱节问题突显,难以形成支持“说明白,讲清楚”中医药科学价值的证据链。因此,需要研制以临床价值为导向的临床与基础研究证据关联转化方法,形成可靠的中医药证据链,以进一步发现规律、确证疗效、彰显优势。
8.2.2 研究方法与结果
天津中医药大学张俊华教授带领团队围绕构建符合中医药疗效特点的评价方法开展了系列研究,针对临床与基础证据脱节问题,率先提出中医药疗效证据链概念,完成了证据链构建与评价方法体系构建,并完成实践验证。该研究以突出中医药临床价值为基点,结合循证医学证据分类分级与评价基本方法,对证据链构建的维度、层次、结点等方面进行分析和专家论证;按照“实践-理论-再实践”的研究路径,以“三药三方”证据转化为载体,在实践中总结、梳理、形成了证据链理论与方法体系,最终通过专家共识形成技术规范。研究过程综合采用了核心指标集、系统评价、Meta分析、Delphi调查、半结构化访谈、专家共识等具体方法。
形成了中医药疗效证据链理论体系及具体的构建与评价方法,共包括:证据链框架,明确了证据链包括临床、药效、机制三个环节,涵盖临床、动物、细胞、分子四个层次的研究证据;构建方法包括成立专家组、临床价值定位、证据收集、质量评价、数据整合、证据链接、证据链可视化等步骤;评价方法包括重要性、真实性、系统性、稳健性四个维度;分类分级方法,根据证据链功能将其分为Ⅰ、Ⅱ、Ⅲ类,基于证据链质量将其分为Ia-Ⅲc级。此外,在具体实践中完成了“三药三方”疗效证据链及中医药治疗呼吸道传染病证据链。
8.2.3 研究价值
中医药疗效证据链实现了临床与基础研究证据的有机整合。将低级别、分散、割裂的证据整合形成体系化证据集,能够促进基础研究成果与临床研究的对接,提高中医药证据的论证强度、可靠性和认可度,进而更好地体现中医药的优势与价值。
上下滑动,查看更多
9.中药盲区成分研究技术的突破驱动常用中药药效物质的揭示
9.1 研究背景
中医药的传承创新发展是国家战略,用现代科学解读中医药学原理是中医药传承创新发展的科学之基,中药药效物质揭示是解读中医药学原理的基础。目前,中药中尚存在大量未被认知的盲区成分,即使已有大量研究的常用中药,受限于技术瓶颈,一些难分离、难鉴定的化学成分仍有待阐明,药效物质更无从揭示。如何突破常用中药盲区成分的瓶颈,揭示其药效物质和科学内涵已成为推动中药学科和产业高质量发展的关键。
9.2 研究方法与结果
依托全国重点实验室、NSFC创新研究群体、教育部国联实验室、教育部双一流学科、国家中医药局重点学科等优势平台,暨南大学中药及天然药物研究所高昊课题组通过构建中药化学成分“分析分离-结构鉴定-化学/生物合成”关键技术体系,发展表观等价非对映异构判别方法、生源规律约束的理论化学空间映射策略等中药药效物质研究创新技术,突破中药盲区成分研究瓶颈,成功应用于以枸杞子、当归为代表的常用中药,取得重要创新成果和进展。
以技术突破为驱动,揭示了以枸杞子、当归为代表的常用中药盲区成分,通过对化学成分的全面揭示和功效关联的活性评价,阐明了药效物质基础,并开展重要活性分子的仿生合成/生物合成研究。从当归中发现了一类新结构物质群,命名为法卡林苯酞(falcarinphthalide),是当归中继多糖、苯丙素、苯酞单倍体、苯酞多聚体、聚炔后一类新的成分类群,即苯酞杂聚体。法卡林苯酞具有抗骨质疏松等活性,是当归药效物质基础的重要组成。法卡林苯酞结构复杂、生源独特,发表后引发关注,被Natural Product Reports 选评为“国际热点分子”(Hot off the press),目前已完成其仿生合成,为法卡林苯酞新药研制提供了物质来源保障。从枸杞子中发现了中一类主含、特征性的新成分二苯丙酰亚精胺,命名为枸杞亚精胺(lycibarbarspermidine),是枸杞子中继多糖、一般酚性成分、苯丙酰芳香胺、枸杞红色素后一类新的成分类群,属于苯丙酰脂肪胺。枸杞亚精胺在神经系统、代谢系统等具有多样的生物活性,是枸杞子药效物质基础的重要组成,目前已纳入国家标准物质研制。枸杞亚精胺的生物合成途径已被成功解析,糖基转移酶催化机制也得以阐明,异源表达的实现为枸杞亚精胺新药创制提供了药源保障。
9.3 研究价值
表观等价非对映异构判别原则和方法纳入医学规划教材《天然药物化学》第三版(科学出版社),新成分类群相关研究发表在ACS Central Science、Nature Communications等期刊。中药药效物质的底层创新发现为中药道地性解析、优良品种选育、作用机制阐明、质量标准提升、加工生产指导、创新药物研制、健康产品研发等提出了新的科学问题和要求,更提供了解决这些问题的手段和途径。药效成分生物合成途径和关键酶催化机制的解析,不但为创新药物药源保障提供绿色制造的解决路径,实现“造物致用”之目标,还有助于深刻理解药效成分生成的生物学机制,为中药道地性成因的解析、品质评价体系建立等提供科学依据,是中药药效物质研究从“格物致知”到“建物致知”的升华。
上下滑动,查看更多
10.新型递药系统助力抗肿瘤类中药活性成分靶向释放
10.1 研究背景
恶性肿瘤是严重威胁人类健康的重大疑难疾病,中药在提高恶性肿瘤临床疗效、减轻不良反应方面具有独特优势,但中药活性成分溶解性差、生物利用度低、靶向性差等缺陷限制了抗肿瘤中药的疗效发挥。中医药在提高恶性肿瘤治疗疗效、减轻不良反应方面具有独特优势,基于肿瘤“正虚伏毒”的病机,中医始终坚持“扶正祛邪”的主要治法,但作用机制尚未阐明。
10.2 研究方法与结果
人参皂苷Rg3仿生靶向脂质体载藤黄酸“扶正祛邪”抗三阴性乳腺癌。三阴性乳腺癌(TNBC)复杂的肿瘤微环境,尤其是血管生成和免疫抑制特性,使临床化疗治疗受限,急需开发一种安全、有效的治疗手段抗TNBC及其转移,以延长患者生存期。以中医“扶正祛邪”理论为指导,联合应用扶正固本的Rg3和以毒攻毒的GA,从抑制肿瘤细胞增殖迁移、改善肿瘤微环境和增强机体抗肿瘤免疫等方面发挥协同治疗TNBC作用。基于“药辅合一”的中药制剂策略,设计、构建了包覆肿瘤细胞膜的人参皂苷Rg3仿生脂质体共载藤黄酸GA用于TNBC治疗。体内外实验证明可降低免疫原性,避免被免疫系统识别清除,延长药物体内的循环时间,靶向递送药物至肿瘤组织及肺转移灶,降低GA对其他正常组织的毒副作用。在肿瘤部位长效释药,提高GA的细胞毒性,有效杀伤肿瘤细胞,有效抑制TNBC生长及肺转移。
构建载藤黄酸仿生纳米递药系统及其协同anti-PD-L1抗乳腺癌。免疫疗法是乳腺癌及其转移的有效治疗手段,但目前临床肿瘤免疫治疗响应效率低,其主要原因是T细胞浸润不足以及效应T细胞分化效率低。针对该难题,利用4T1细胞膜的同源靶向性,制备了包覆4T1细胞膜的仿生纳米粒Tm@PDA-GA NPs。制剂中GA通过下调HSP90不仅增强PTT实现协同抑制肿瘤增殖,而且引起ICD,促进DC细胞成熟;GA抗血管生成改善肿瘤血管功能并增强肿瘤免疫浸润。Tm@PDA-GA NPs协同免疫检查点抑制剂anti-PD-L1,能够进一步增强肿瘤免疫治疗效应,有效抑制TNBC生长及肺转移。
蟾毒灵纳米递药系统增敏索拉非尼抗肝癌。索拉非尼(Sorafenib,SFN)作为美国食品药品监督管理局最早批准的晚期肝癌小分子靶向治疗药物,在中晚期肝癌的治疗中取得了较好的效果,但长期治疗会导致SFN的应答敏感性降低,使得其临床应用受到限制。蟾毒灵(Buf)作为临床肝癌辅助治疗药物华蟾素胶囊的主要成分之一,可作为HIF-1α靶点抑制剂,恢复肝癌细胞对SFN的敏感性,体现Buf“祛邪”优势。Buf和SFN联用不仅可实现协同促进肝癌细胞凋亡,而且还可协同抑制血管内皮生长因子VEGF生成,增强抗肿瘤血管新生能力,进一步发挥联合治疗肝癌的作用。为了实现SFN和Buf在血管和肿瘤部位的靶向递送,并实现药物在肿瘤部位的响应性释放,团队开发了一种肿瘤-血管双重靶向纳米递送系统S/B@FA/cRGD-LB-ZIF-8,将SFN和Buf高效递送到血管和肿瘤部位,可以改善肿瘤乏氧微环境、抑制血管生长和诱导肿瘤细胞凋亡,增强SFN与Buf的协同抗肝癌作用。
10.3 研究价值
该研究将中医药理论与现代纳米递药系统相结合,充分发挥Rg3、GA和Buf的“扶正祛邪”优势作用,为“扶正祛邪”抗肿瘤中药临床优势阐明及二次开发提供参考。针对源于临床抗肿瘤中药的活性成分靶向性差、水溶性低、毒副作用大、分布广、消除速度快等问题,搭建中药纳米递送技术平台,探索中西医结合治疗抗肿瘤的临床优势,亦为中药创新药物研究和临床用药提供支撑。
上下滑动,查看更多
总结
2024年度中医药十大学术进展遴选发布工作集中展现了中医药领域的前沿成果和蓬勃发展态势。新技术新方法的融合推动了中医原创理论、中药疗效作用机制的科学解读;高质量临床研究,彰显了中医药在防治重大慢病和难治性疾病方面的独特优势;中药资源保障、新药研发、新型递药系统等领域的研究,促进了中药全链条创新能力的提升;中药有效性评价方法体系的构建,则为中药临床应用和产业化发展提供了重要的保障。
虽然近年来中医药学术研究已取得了很大的进步,但距离党中央、国务院的要求以及人民对健康的迫切需求还有很大的提升空间。原创优势发挥不足、中医诊疗优势未充分显现、重大研究成果较少、研究深度不够等挑战依然存在。未来,中医药学术发展需要重点关注以下几个方面:一是发挥中医药原创理论优势,加强人工智能、系统生物学等多学科前沿技术与中医药的深度交叉融合,推动中医药原理解读研究范式转变与方法学创新,推动中医药理论取得新突破;二是发挥中医药防病治病独特优势,优先选择中西医具有共识、临床需求未被满足、中医药或中西医结合诊疗具有优势、中医理论具有原创性的疾病,推动中西医结合的临床研究,形成符合中医药自身规律的中西医结合临床诊疗新模式,提升中医药在健康中国建设中的贡献度;三是加强中药全链条创新能力的基础研究,从资源保障、新药研发、工业制造、合理应用等方面开展研究,创新适合中药的研究方法,构建符合中药特色的中药全链条创新策略,推动形成复杂疾病的药物研发和应用新理论。
期刊简介
Journal Introduction
World Journal of Integrated traditional and western Medicine(WJIM,CN 10-1354/R,ISSN 2096-0964)是由中国科学技术协会主管、中华中医药学会主办的唯一一本依托“本土一体化平台”的中医药英文期刊,季刊,国内外公开发行。2015年创刊,时任主编中国工程院院士吴以岭,名誉主编国医大师路志正教授,现任主编中华中医药学会青年委员会主任委员张霄潇。
期刊依托国家政策及学会资源支撑,遴选行业全球前10%顶尖学术成果,对标国际顶级期刊,致力打造影响因子>10的行业标杆。
WJIM立足四大核心定位:
学术引领地——凝聚行业智慧,汇聚学术资源,发表年度中医药十大进展及评述、年度中医药重大科学问题、中医药标志性科技成果总结等高质量论文,引领中医药高质量发展。
医学启迪地——报道具有重大科学研究启发价值的临床案例(类似砒霜治疗白血病临床案例启发三氧化二砷治疗M3型白血病的原创机制发现),启迪世界生命科学原始创新。
标签打造地——刊载中医药、中西医结合领域优秀青年学者的原创理论、新技术新方法的开发及应用、临床科学问题的揭示等前沿成果,塑造学术标签,助力青年学者跨越式成长。
信息集散地——统筹报道高端学术活动(如科技项目布局、中医药临床优势病种沙龙、中医药新技术新方法沙龙、求实项目研究成果、岐黄论坛、青年岐黄论坛等内容)的成果,驱动多学科深度交叉。
WJIM官网:
https://www.tcmjc.com/wjim
WJIM投稿网址:
https://cacm.tcmjc.com/WJIM
编排:徐宝南
编辑:刘艳玲
初审:肖云
复审:郭蕾
终审:郭希勇
100 项与 北京中医药大学 相关的药物交易
登录后查看更多信息
100 项与 北京中医药大学 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年06月10日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
2
22
临床前
临床申请批准
1
1
临床2期
临床3期
2
4
其他
登录后查看更多信息
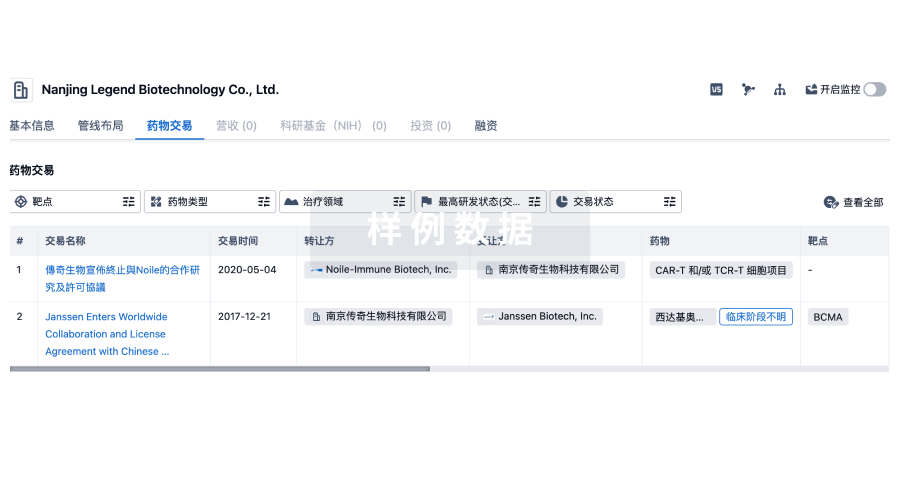
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
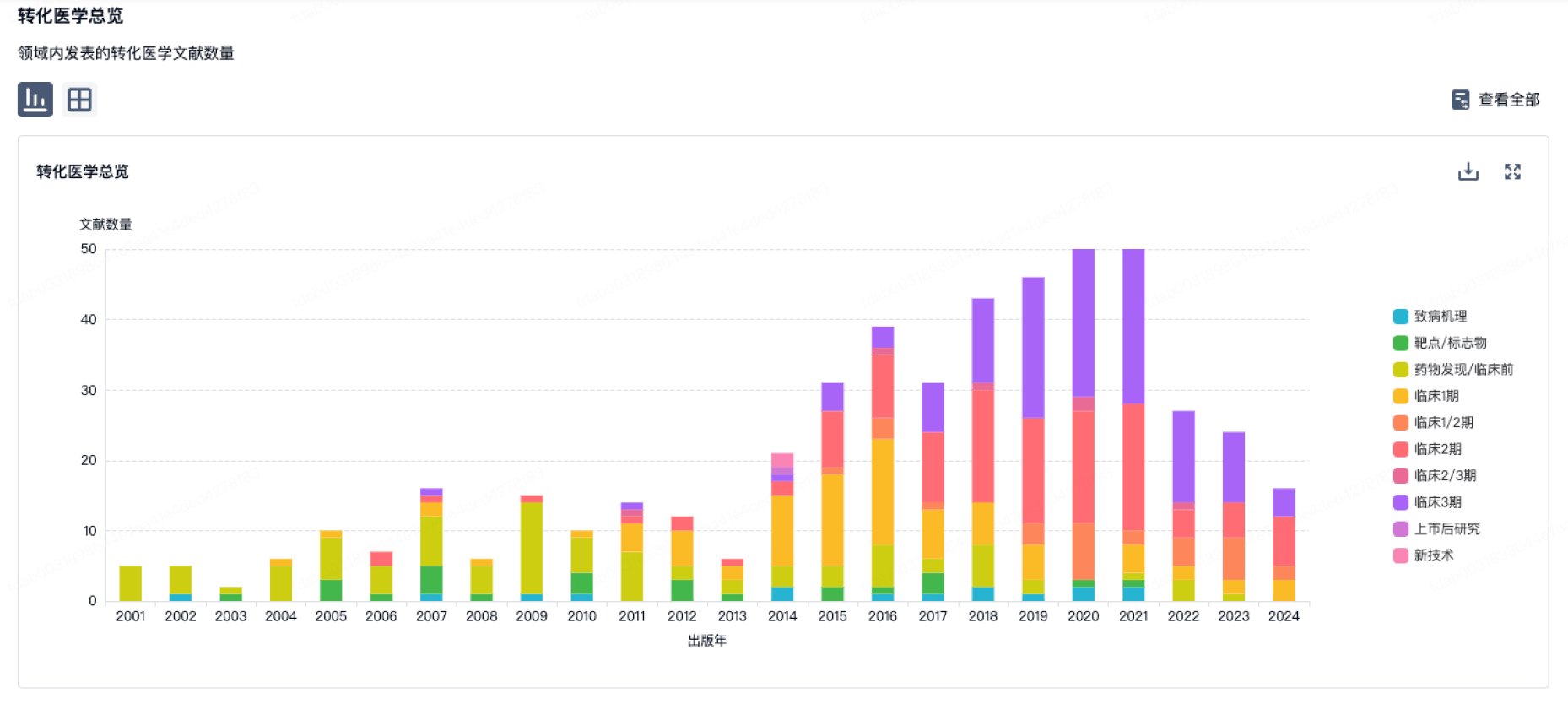
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
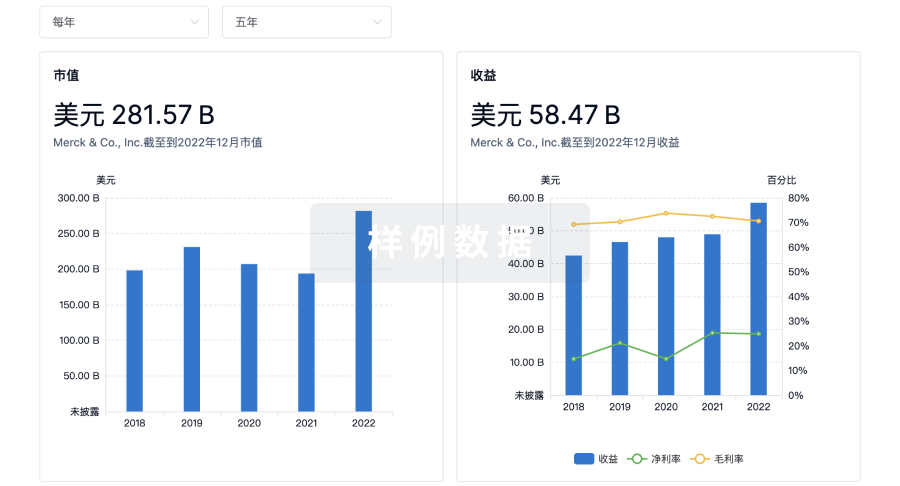
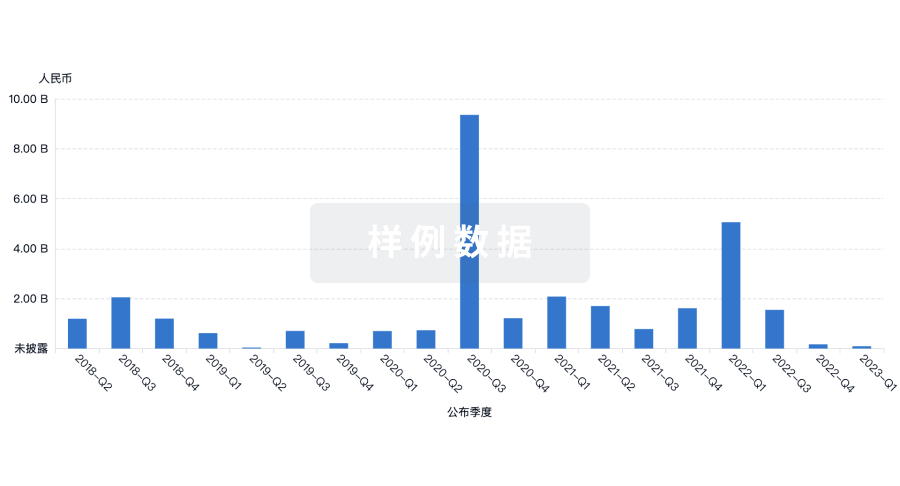
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
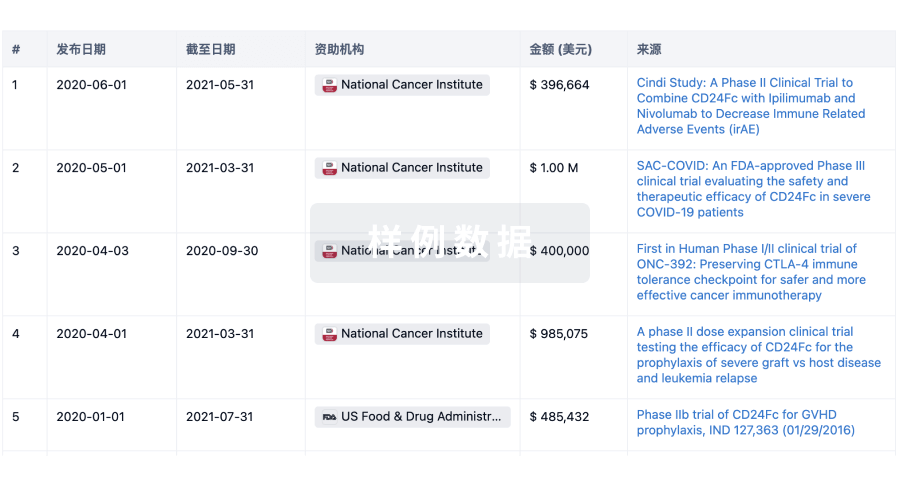
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
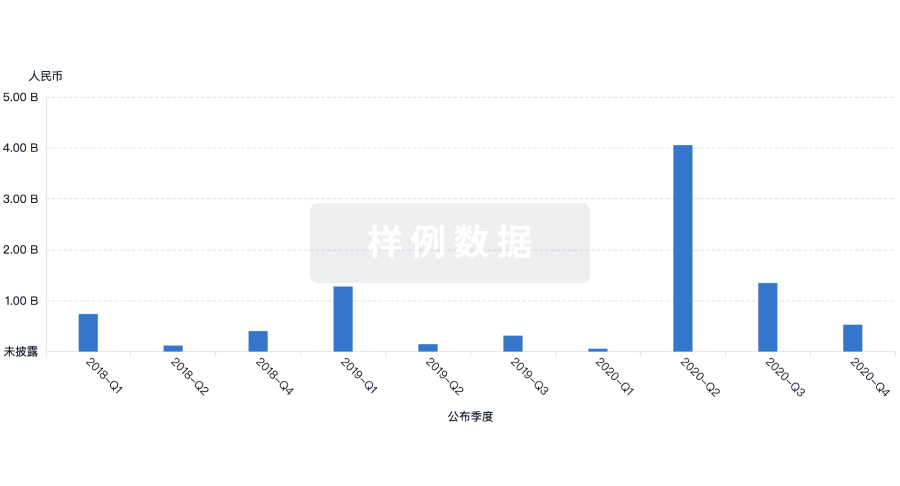
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
