预约演示
更新于:2026-03-08

Carmot Therapeutics, Inc.
更新于:2026-03-08
概览
标签
内分泌与代谢疾病
其他疾病
消化系统疾病
小分子化药
合成多肽
放射与诊断药物
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 小分子化药 | 5 |
| 放射与诊断药物 | 2 |
| 合成多肽 | 2 |
关联
9
项与 Carmot Therapeutics, Inc. 相关的药物作用机制 GIPR激动剂 [+1] |
非在研适应症 |
最高研发阶段临床3期 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 GLP-1R激动剂 |
在研机构 |
非在研适应症- |
最高研发阶段临床2期 |
首次获批国家/地区- |
首次获批日期- |
作用机制 GIPR激动剂 [+1] |
最高研发阶段临床2期 |
首次获批国家/地区- |
首次获批日期- |
10
项与 Carmot Therapeutics, Inc. 相关的临床试验NCT06628362
A Randomized, Double-Blind, Placebo-Controlled, Parallel-Group, Multi-Center Phase 2 Study to Evaluate the Efficacy, Safety, and Tolerability of Once-Weekly CT-388 Administered Subcutaneously for 48 Weeks to Participants Who Are Overweight or Obese With Type 2 Diabetes Mellitus
This is a multi-center, randomized, double-blind, placebo-controlled, parallel group dose-finding study to evaluate the efficacy and safety of CT-388 at low, middle, and high doses in participants who are overweight or obese with Type 2 diabetes mellitus (T2DM).
开始日期2024-11-21 |
申办/合作机构 |
ISRCTN90651766
Phase I trial: Quotient code QSC301813
开始日期2024-09-16 |
申办/合作机构 |
NCT06525935
A Randomized, Double-Blind, Placebo-Controlled, Parallel Group, Multi-Center Phase 2 Study to Evaluate the Efficacy, Safety, and Tolerability of Once-Weekly CT-388 Administered for 48 Weeks to Participants With Obesity or Overweight With at Least One Weight-Related Comorbidity
This is a multi-center, randomized, double-blind, placebo-controlled, parallel group dose-finding study to evaluate the efficacy and safety of CT-388 at low, middle, and high doses in participants with obesity or who are overweight with at least one weight-related comorbidity.
开始日期2024-08-16 |
申办/合作机构 |
100 项与 Carmot Therapeutics, Inc. 相关的临床结果
登录后查看更多信息
0 项与 Carmot Therapeutics, Inc. 相关的专利(医药)
登录后查看更多信息
4
项与 Carmot Therapeutics, Inc. 相关的文献(医药)2019-01-01·Cell chemical biology1区 · 生物学
Fragment-Based Drug Discovery: Advancing Fragments in the Absence of Crystal Structures
1区 · 生物学
Review
作者: Erlanson, Daniel A ; Jahnke, Wolfgang ; Davis, Ben J
Fragment-based drug discovery typically requires an interplay between screening methods, structural methods, and medicinal chemistry. X-ray crystallography is generally the method of choice to obtain three-dimensional structures of the bound ligand/protein complex, but this can sometimes be difficult, particularly for early, low-affinity fragment hits. In this Perspective, we discuss strategies to advance and evolve fragments in the absence of crystal structures of protein-fragment complexes, although the structure of the unliganded protein may be available. The strategies can involve other structural techniques, such as NMR spectroscopy, molecular modeling, or a variety of chemical approaches. Often, these strategies are aimed at guiding evolution of initial fragment hits to a stage where crystal structures can be obtained for further structure-based optimization.
2018-01-01·The Journal of biological chemistry2区 · 生物学
Co(II) and Ni(II) binding of the Escherichia coli transcriptional repressor RcnR orders its N terminus, alters helix dynamics, and reduces DNA affinity
2区 · 生物学
Article
作者: Bobst, Cedric E ; Maroney, Michael J ; Iwig, Jeffrey S ; Huang, Hsin-Ting ; Kaltashov, Igor A ; Chivers, Peter T
RcnR, a transcriptional regulator in Escherichia coli, derepresses the expression of the export proteins RcnAB upon binding Ni(II) or Co(II). Lack of structural information has precluded elucidation of the allosteric basis for the decreased DNA affinity in RcnR's metal-bound states. Here, using hydrogen-deuterium exchange coupled with MS (HDX-MS), we probed the RcnR structure in the presence of DNA, the cognate metal ions Ni(II) and Co(II), or the noncognate metal ion Zn(II). We found that cognate metal binding altered flexibility from the N terminus through helix 1 and modulated the RcnR-DNA interaction. Apo-RcnR and RcnR-DNA complexes and the Zn(II)-RcnR complex exhibited similar 2H uptake kinetics, with fast-exchanging segments located in the N terminus, in helix 1 (residues 14-24), and at the C terminus. The largest difference in 2H incorporation between apo- and Ni(II)- and Co(II)-bound RcnR was observed in helix 1, which contains the N terminus and His-3, and has been associated with cognate metal binding. 2H uptake in helix 1 was suppressed in the Ni(II)- and Co(II)-bound RcnR complexes, in particular in the peptide corresponding to residues 14-24, containing Arg-14 and Lys-17. Substitution of these two residues drastically affected DNA-binding affinity, resulting in rcnA expression in the absence of metal. Our results suggest that cognate metal binding to RcnR orders its N terminus, decreases helix 1 flexibility, and induces conformational changes that restrict DNA interactions with the positively charged residues Arg-14 and Lys-17. These metal-induced alterations decrease RcnR-DNA binding affinity, leading to rcnAB expression.
2011-01-01·Topics in current chemistry
Introduction to Fragment-Based Drug Discovery
Review
作者: Erlanson, Daniel A
Fragment-based drug discovery (FBDD) has emerged in the past decade as a powerful tool for discovering drug leads. The approach first identifies starting points: very small molecules (fragments) that are about half the size of typical drugs. These fragments are then expanded or linked together to generate drug leads. Although the origins of the technique date back some 30 years, it was only in the mid-1990s that experimental techniques became sufficiently sensitive and rapid for the concept to be become practical. Since that time, the field has exploded: FBDD has played a role in discovery of at least 18 drugs that have entered the clinic, and practitioners of FBDD can be found throughout the world in both academia and industry. Literally dozens of reviews have been published on various aspects of FBDD or on the field as a whole, as have three books (Jahnke and Erlanson, Fragment-based approaches in drug discovery, 2006; Zartler and Shapiro, Fragment-based drug discovery: a practical approach, 2008; Kuo, Fragment based drug design: tools, practical approaches, and examples, 2011). However, this chapter will assume that the reader is approaching the field with little prior knowledge. It will introduce some of the key concepts, set the stage for the chapters to follow, and demonstrate how X-ray crystallography plays a central role in fragment identification and advancement.
297
项与 Carmot Therapeutics, Inc. 相关的新闻(医药)2026-03-06
Copenhagen-based Zealand Pharma announced positive petrelintide Phase II results from the ZUPREME-1 dose-finding trial, reporting up to 10.7% mean body weight reduction at 42 weeks in people with overweight and obesity.
Petrelintide is a long-acting amylin analog designed for once-weekly subcutaneous administration. The drug is being developed through an exclusive collaboration between Zealand Pharma and Roche signed in 2025, which grants Roche co-development and co-commercialization rights. The Phase II data for petrelintide produced weight loss below the levels reported for leading GLP-1–based therapies, with semaglutide demonstrating roughly 13%-15% reductions and tirzepatide around 20% in longer-duration Phase III trials. While this suggests more modest efficacy as a monotherapy, petrelintide showed a placebo-like safety profile. Roche has indicated that the drug may ultimately be positioned as part of combination regimens, potentially alongside its GLP-1 pipeline candidate CT-388, which the company acquired through its purchase of Carmot Therapeutics.
ZUPREME-1 (NCT06662539) is a randomized, double-blind, placebo-controlled, parallel-group, multinational Phase II trial conducted across 33 sites in the United States, Poland, and Romania. The study enrolled 493 adults with obesity or overweight with weight-related comorbidities randomized across five petrelintide dose arms and placebo. Dosing involved once-weekly subcutaneous petrelintide with dose escalation every four weeks over a 16-week period, followed by maintenance through week 42 and a nine-week safety follow-up to week 51.
The primary endpoint, percentage change in body weight from baseline to week 28, was met across all five petrelintide arms with statistical significance versus placebo. Weight reduction continued through week 42, reaching up to 10.7% (efficacy estimand) compared to 1.7% with placebo (p<0.001). In the maximally effective arm, 98% of participants escalated to the targeted maintenance dose.
The tolerability profile was the trial’s distinguishing feature. Treatment discontinuation due to adverse events was 4.8% in the maximally effective petrelintide arm versus 4.9% with placebo. There were no cases of vomiting and no GI-related discontinuations at the maximally effective dose. Nausea rates were lower than in the prior 16-week Phase Ib trial, which used faster dose escalation, and nearly disappeared after participants reached maintenance dosing. No unexpected safety signals emerged, including for alopecia, fatigue, or neuropsychiatric events. Overall trial withdrawal was 8.4% across petrelintide arms versus 13.6% with placebo.
Zealand plans to initiate Phase III development later in 2026, with the ZUPREME-1 data informing trial design. Final results including the follow-up period are expected at a scientific conference in 2026. Topline data from ZUPREME-2, evaluating petrelintide in people with overweight or obesity and type 2 diabetes, are anticipated in the second half of 2026. A Phase II combination trial with CT-388 is planned for the first half of 2026.
Petrelintide mimics endogenous amylin, a peptide co-secreted with insulin from pancreatic beta cells in response to nutrient intake. Amylin receptor activation reduces body weight by restoring sensitivity to the satiety hormone leptin and inducing earlier satiation. This mechanism is distinct from GLP-1 receptor agonism, the pathway exploited by semaglutide and tirzepatide. The tolerability data from ZUPREME-1 are notable because GI side effects, particularly nausea and vomiting, remain the primary cause of treatment discontinuation with GLP-1-based obesity therapies, limiting long-term adherence.
The amylin analog weight loss space has drawn considerable investment. Key competitors include:
Eli Lilly’s eloralintide, a selective amylin receptor agonist being readied for Phase III as a monotherapy and in combinations with other metabolic targets such as GLP-1 or GIP receptor agonists. Novo Nordisk’s amycretin, a dual GLP-1 and amylin receptor agonist in oral formulation that has produced Phase Ib/IIa weight loss data and is advancing toward Phase III.
Your email address will not be published. Required fields are marked *
Comment *
Name *
Email *
Website
临床2期临床结果并购临床3期临床1期
2026-03-04
替尔泊肽作为2025年新晋"药王",其原料药市场正从礼来单极主导转向全球多极竞争格局,中国药企凭借规模化产能和工艺创新快速崛起,同时价格战与专利到期压力正重塑行业竞争逻辑。
一、替尔泊肽市场地位与销售情况
1. "药王"地位确立:礼来替尔泊肽2025年全球销售额达365.07亿美元(+122%),以4亿美元微弱优势超越司美格鲁肽(346.06亿美元),正式加冕全球"药王"宝座。其中降糖版Mounjaro销售额229.65亿美元(+99%),减重版Zepbound销售额135.42亿美元(+175%)。
2. 2026年增长预期:礼来预计2026年全年总营收将达到800-830亿美元(较2025年增长25%),替尔泊肽将继续作为核心增长引擎。全球市场对替尔泊肽的需求持续攀升,礼来已投入超180亿美元在全球扩建产能,包括将印第安纳州黎巴嫩基地投资翻倍至90亿美元。
3. 中国市场表现:替尔泊肽在中国区销售额约为7.96亿丹麦克朗(约合人民币8.76亿元),同比增长314%,显示出强劲的市场渗透力。2026年1月1日,替尔泊肽用于成人2型糖尿病治疗正式纳入中国医保目录,进一步提升市场可及性。
推荐阅读:
1.替尔泊肽原料开发方法及策略
2.国内替尔泊肽原料药竞争格局
3.替尔泊肽公斤级GMP合成工艺
4.替尔泊肽合成工艺过程
二、替尔泊肽原料药全球竞争格局
1. 礼来主导的原研格局:礼来作为替尔泊肽原研企业,拥有完整的知识产权和生产工艺,通过与CDMO企业合作满足全球需求,并通过严格的供应链管理和专利保护维持市场主导地位。
药明康德/合全药业承接了礼来替尔泊肽35%的多肽原料药供应及85%的活性成分(API)供应,2025年底多肽固相合成产能达10万升。
2. 中国药企的快速崛起:
产能布局:截至2025年底,中国已有超20家企业明确规划或已具备替尔泊肽原料药生产能力。
代表企业:
药明康德/合全:TIDES业务2025年收入78.4亿元,同比增长121%
诺泰生物:替尔泊肽单批次产量超10kg,新规划基地将新增500kg产能
泰德医药:具备单批30kg以上、年产能超500kg的商业化交付能力
翰宇药业:2026年1月8日完成替尔泊肽原料药国家药监局官方登记
技术优势:中国药企凭借规模化合成与工艺优化,原料成本可达较低水平,同时通过AI制药工具将研发周期缩短45%。
3. 国际药企的BD布局:
阿斯利康引进诚益生物ECC5004进入2b期临床;罗氏以30亿美元收购Carmot Therapeutics获得CT-996;默沙东与翰森制药合作开发HS-10535;辉瑞于2025年12月与复星医药达成合作,最高里程碑付款达19.35亿美元;这些合作主要聚焦于口服小分子GLP-1和多靶点激动剂等创新方向。
长期大量供应各种多肽固相合成树脂:如2-CTC树脂,wang树脂,Rink 酰胺 AM树脂,Sieber 树脂等,同时供应各种多肽、重组蛋白、抗体纯化填料,层析设备,DAC柱及制备色谱装置。联系:15771973009(微信同)。
三、替尔泊肽原料药技术特点与生产难点
1. 分子结构复杂性:替尔泊肽是一个39个氨基酸长的多肽分子,在20位Lys有一个20碳的脂肪酸修饰,分子式为C225H348N48O68,分子量为4813.53Da,这种复杂结构对合成工艺和纯化技术提出了极高要求
2. 生产工艺难点:
合成路线复杂:原研专利CN113330024A采用四个片段合成工艺,粗肽纯度70%,合成总收率44.4%。
纯化难度大:片段拼接后除去副产物需用纳滤技术,对滤膜要求极高(200Da和400Da),国内滤膜难以达到
溶剂用量大:传统工艺TFA裂解液用量高(8-10ml/克树脂),醚类溶剂用量高达80-100ml/克树脂,产生大量废液
沉降困难:裂解液中替尔泊肽浓度低,沉降颗粒细小,难以过滤分离
3. 工艺创新突破:
新型制备方法采用Siber树脂作为起始固相载体,使用3%-5%TFA的DCM溶液进行肽链切割,该工艺可减少70%TFA用量,降低90%醚类溶剂用量,使沉降颗粒增大,便于过滤,中国药企通过工艺优化,已能实现吨级替尔泊肽原料药生产,如诺泰生物单批次产量超10kg
四、替尔泊肽原料药价格趋势与市场竞争
1. 价格战全面爆发:
2025年11月起,诺和诺德与礼来相继下调司美格鲁肽与替尔泊肽价格。替尔泊肽院内市场最低剂量单剂价格降至81.03元,较上市初期降幅达80%。京东买药平台显示,替尔泊肽2.5mg4次规格降至599元,5mg4次降至899元,降幅均超60%
2. 降价驱动因素:
医保谈判压力:2026年1月1日起,替尔泊肽用于成人2型糖尿病治疗纳入中国医保目录
专利到期临近:司美格鲁肽核心专利将于2026年3月20日到期,替尔泊肽面临仿制药竞争压力
市场竞争加剧:GLP-1赛道已从"蓝海市场"步入"群雄逐鹿"的红海竞争新阶段
3. 价格战影响:
礼来表示,部分销量增长被替尔泊肽实际售价下降所抵消。诺和诺德则宣布下调2026年销售额指引,预计全年全球销售额将较2025年下降5%-13%。高盛分析师预计,到2030年全球肥胖症药物销售额将达1050亿美元,低于此前1300亿美元的预测
五、替尔泊肽原料药未来发展趋势
1. 从单一爆款到多维竞争:2026年后GLP-1竞争不再依赖单一爆款,而是围绕给药方式、减重质量与联合疗法建立综合优势,礼来与诺和诺德正从"减重效率"导向转向"减重质量"深耕,聚焦更安全、精准、适配个体需求的差异化路径。
2. 口服GLP-1成为关键突破口:诺和诺德已获批全球首个口服版GLP-1药物Wegovy,64周平均减重16.6%,礼来Orforglipron作为小分子路线领跑者,已完成III期临床,36mg剂量组平均减重8.9kg(减重率9.2%)口服制剂因给药便捷性和。安全性优势,契合BMI介于24-28轻症患者的长期体重管理需求。
3. 多靶点协同治疗成为增效革命核心:
“GLP-1+Amylin":诺和诺德CagriSema(司美格鲁肽+卡格列肽)已提交NDA,依从性人群平均减重22.7%
"GLP-1+FGF21":诺和诺德以最高52亿美元收购Akero Therapeutics,获得III期阶段的Efruxifermin
"减脂增肌"新焦点:礼来主攻ActRII受体抑制剂Bimagrumab,联合替尔泊肽减重22.1%,92.9%来自脂肪
4. 中国药企的差异化战略:"全剂型+低成本+快进度":翰宇药业布局日制剂、周制剂、超长效月制剂、口服制剂,满足不同患者需求
多靶点创新:华东医药DR10624(GLP1R/GCGR/FGF21三激动剂)进入II期临床国际化布局:中国药企依托90多个国家和地区的业务布局,加速全球化销售渗透。
当前,替尔泊肽原料药市场正处于从原研主导向多极竞争的关键转型期,价格战与专利到期压力正加速行业洗牌。未来,具备全产业链能力、研发创新积累与国际化品质体系的企业将更具竞争优势,而口服制剂、多靶点激动剂和减脂增肌等创新方向将成为行业发展的新引擎。中国药企凭借规模化产能和工艺创新正快速崛起,有望在全球替尔泊肽原料药市场中占据重要地位。
免责声明:本微信文章中的信息仅供一般参考之用,部分内容摘自网络及AI生成,如有侵权联系删除。此公众号任何时候都不保证文中描述内容的可靠性,不可直接作为决策内容,生物制药合伙人不对任何主体因使用本文内容而导致的任何结果承担责任。
公众号已建立“生物药工艺开发技术交流”微信群,添加小编微信514099167 加入,加微信时备注姓名-单位-岗位,非生物制药行业人员勿扰。
生物制药合伙人
扫码关注我们
ID:zpt_Partner
W:514099167
分享,点赞,在看!为生物制药人三连击!
并购专利到期财报
2026-02-28
2026年初的全球医疗市场,罗氏(Roche)正处于其企业历史上最具决定性的转型期之一。当市场为礼来和诺和诺德的代谢双寡头神话狂热时,罗氏的预期市盈率却徘徊在20倍以下。
然而,这份由于入局GLP-1较晚而带来的估值折价,掩盖了罗氏正在执行的一场激进且高风险的战略重组。这份重组的意义不在于一家百年巨头如何艰难防守,而在于它如何果断打破传统的研发孤岛,利用偏向性机制和底层药理学的差异化,向已固化的千亿级赛道发起降维打击。
对于医药行业而言,罗氏的转型提供了一个极为硬核的战略范本:如何在管线挫折后迅速转向,并利用诊断+制药的双重生态建立长效护城河。
强劲的内生增长与被低估的管线期权
最新的财务切片与市场数据显示,罗氏的基础业务极具韧性:
* 增长引擎切换:2025年前九个月,制药部门销售额达356亿瑞士法郎,按固定汇率(CER)强劲增长9%。增长已成功从老一代肿瘤三驾马车切换至Vabysmo、Ocrevus、Hemlibra等新星。
* 眼科的统治力:双特异性抗体Vabysmo年化销售额突破40亿美元,并在美国湿性AMD市场拿下了约30%的份额,主导了市场份额的掠夺。
* 资本市场的认知错位:罗氏当前的估值与礼来(约40-48倍)、诺和诺德(约35倍)存在显著鸿沟。市场尚未将其2028年前后潜在的爆发点充分计入价格。
打破非在此发明的研发孤岛
战略的突围始于文化的重塑。历史上,罗氏旗下的基因泰克(Genentech)以极其强大的内部研发引擎著称,对外部资产往往持审慎甚至排斥的态度(即非在此发明综合症)。但在肺癌(TIGIT抑制剂)和阿尔茨海默病项目连续遭遇重大挫折后,现任CEO Thomas Schinecker果断打破了这一传统。商业模式与资本配置随之发生剧变:
从内部死磕➔外部敏捷并购。
2023至2024年间,罗氏斥资27亿美元收购Carmot Therapeutics切入肥胖领域,随后又以71亿美元收购Telavant大举进军炎症性肠病(IBD)市场。这证明了新管理层对速度的极致追求,并确立了通过外部同类最佳(Best-in-Disease)资产填补空白的务实路线。
从跟随者到规则重塑者
1.代谢领域的偏向性赌局(Biased Agonism)
罗氏重返代谢领域并非简单的“Me-Too”复制。其通过Carmot获得的核心资产(CT-388和CT-996)在分子设计上采用了突破性的偏向性信号传导机制。
* 解决耐药痛点:现有的GLP-1药物会同时激活G蛋白通路和导致受体内吞的β-arrestin通路。而罗氏的资产能选择性激活有益的G蛋白通路,几乎不招募β-arrestin。这有望解决患者在用药60-70周后出现的减重平台期问题,维持更长久的疗效。
* 临床数据印证:1b期试验中,CT-388展现了极快的起效速度,24周平均减重达18.8%。同时,口服小分子CT-996凭借极低的生产成本优势,旨在未来颠覆巨大的维持治疗市场。
2. IBD领域的精准医疗降维打击
在IBD领域,罗氏通过71亿美元豪赌TL1A抗体RVT-3101。
区别于现有的抗TNF疗法,TL1A不仅能驱动炎症,还能直接激活成纤维细胞导致纤维化。RVT-3101有望逆转或预防纤维化并发症。
更关键的是,罗氏正利用其强大的伴随诊断能力,在3期试验中筛选高TL1A表达患者,试图将IBD治疗带入精准医疗时代。
3. 眼科双靶点的防守反击
Vabysmo通过同时靶向VEGF-A和Ang-2两条通路,在临床上实现了更强的视网膜积液干燥能力。这种双重机制成功将给药间隔延长至16周,直接促成了约60%的新患者从竞争对手(Eylea)阵营中转换而来,稳固了罗氏在眼科高端市场的压倒性优势。
生态融合与地缘对冲
未来制药巨头的竞争早已超越了单纯的分子发现。罗氏的终极护城河正在向基础设施层面转移:
* 诊断与AI的生态绑定:罗氏推出了革命性的cobas Mass Spec流水线,将原本极其复杂的液相色谱-质谱(LC-MS)分析全自动化,极大拓宽了质谱检测的普及面。同时,Navify数字病理系统通过AI算法深度绑定了医院客户,提高了转换成本。
* 在美国,为美国的重资产对冲:面对全球日益加剧的贸易保护主义和潜在关税威胁,罗氏制定了防御性的资本布局,承诺未来五年内向美国基础设施和研发中心投资500亿美元。这种前瞻性布局确保了其最大单一市场的供应链绝对安全。
【MedVerse Pro 观察】
抛开市盈率的表象,2026年的罗氏正在执行一场教科书级别的大象转身。对于中国本土的生物医药企业和营销操盘手而言,罗氏的战略选择极具启发性:
迟到者的破局点在于底层重构。
在高度拥挤的赛道(如GLP-1、自免),如果仅仅是效仿头部企业,只会被价格战吞噬。后发企业必须像罗氏寻找偏向性激动剂一样,从更深的药理学机制(耐药性、给药途径)寻找突破口,实现Best-in-Disease的价值主张。
诊疗一体化将决定商业化上限。
随着靶向药物的竞争白热化,如何证明你的药物比同类更好?答案在于患者筛选。利用伴随诊断实现精准用药(如罗氏在IBD领域的TL1A生物标志物策略),将是未来获取医保支付支持、提高临床响应率的核心营销抓手。
当资本市场还在为即时的减肥数字买单时,罗氏已经将重金压在了更为底层的分子机制与全自动生态上。这场关于深度与速度的战役,才刚刚开始。
内容声明: “MedVerse Pro”持续跟踪全球医药产业动态,旨在传递前沿医药资讯与全域营销智库洞察。本文所有内容仅限定向供医疗卫生专业人士(HCP)及医药行业从业者进行学术探讨与商业逻辑交流,绝不构成任何临床诊疗建议、用药指导或对特定处方药物的商业推广。文中引用的财务数据、企业管线及获批信息均源自各国监管机构通报(如NMPA、FDA)或公开核实渠道。如存在数据疏漏或探讨之处,欢迎联系我们修正交流。
TeamMedVerse
100 项与 Carmot Therapeutics, Inc. 相关的药物交易
登录后查看更多信息
100 项与 Carmot Therapeutics, Inc. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2026年05月07日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
4
1
临床前
临床1期
2
2
临床2期
其他
3
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
CT-388 ( GIPR x GLP-1R ) | 肥胖 更多 | 临床2期 |
CT-868 ( GIPR x GLP-1R ) | 肥胖 更多 | 临床2期 |
CT-996 ( GLP-1R ) | 2型糖尿病 更多 | 临床1期 |
CT-966 ( GLP-1R ) | 肥胖 更多 | 临床1期 |
CT-859 ( GIPR x GLP-1R ) | 肝细胞癌 更多 | 临床前 |
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
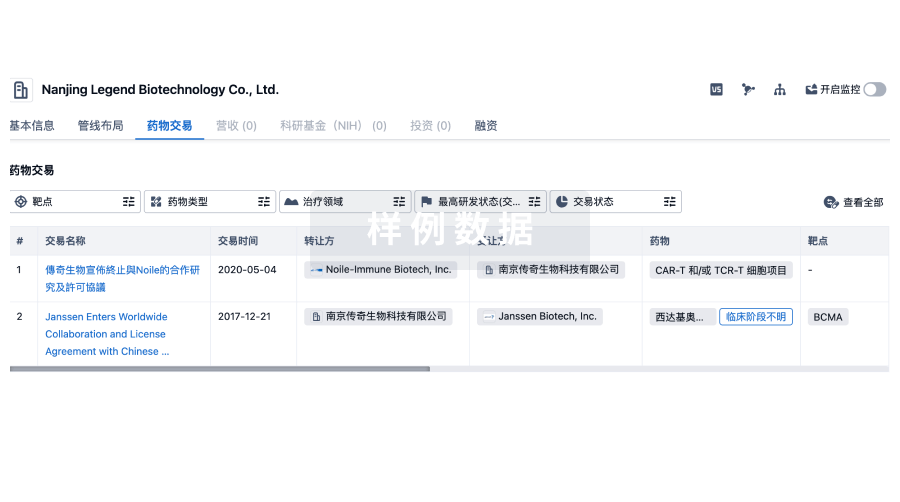
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
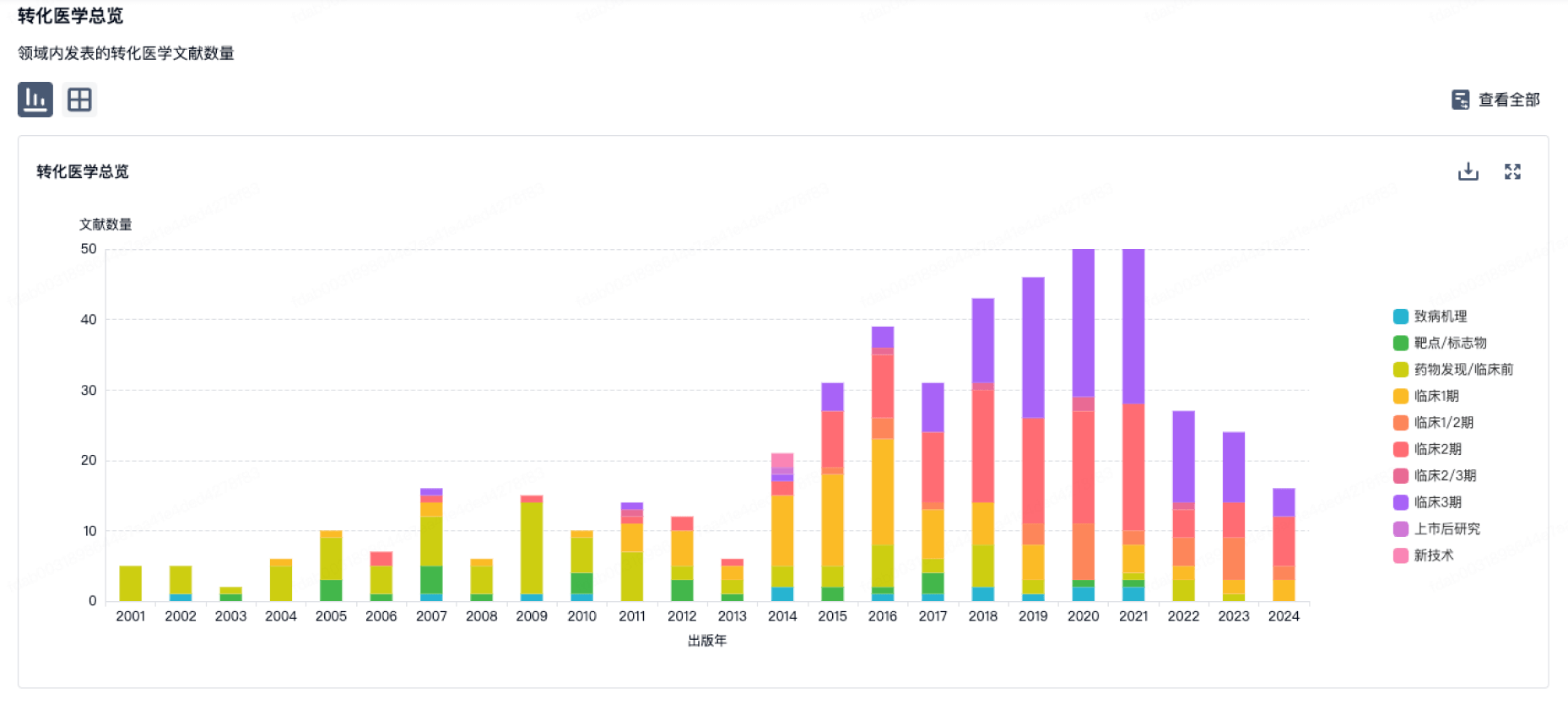
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
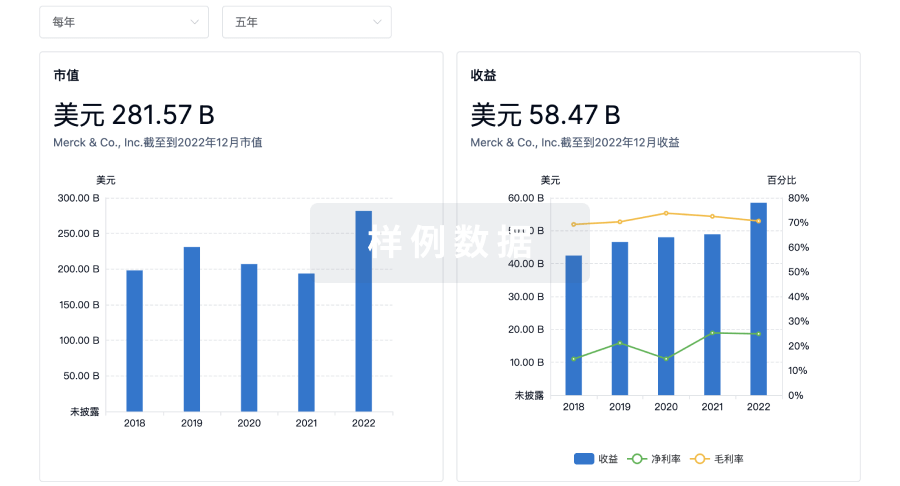
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
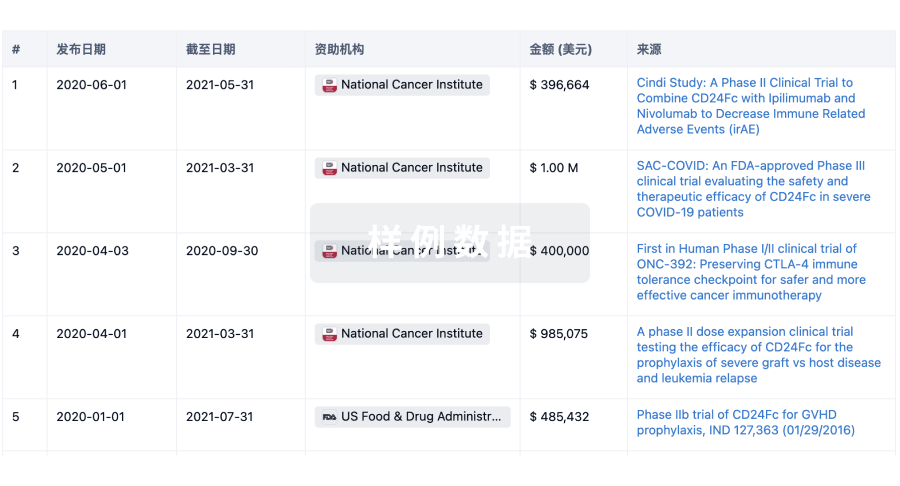
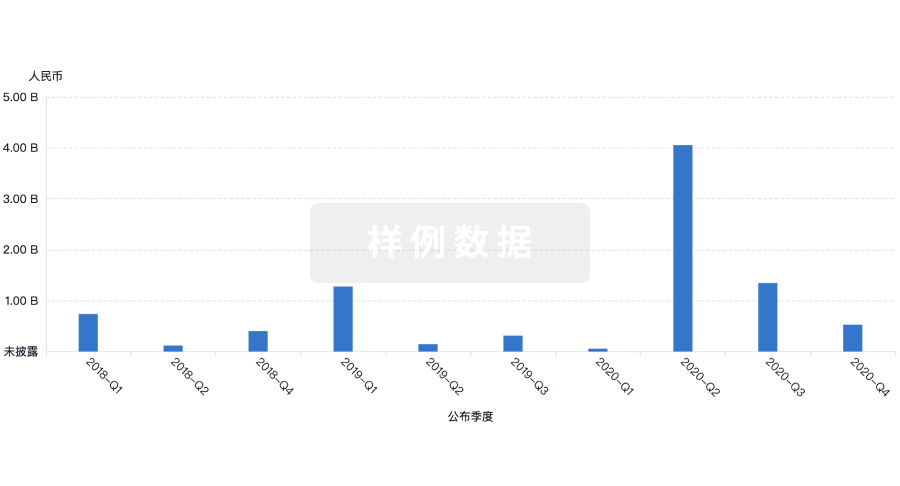
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
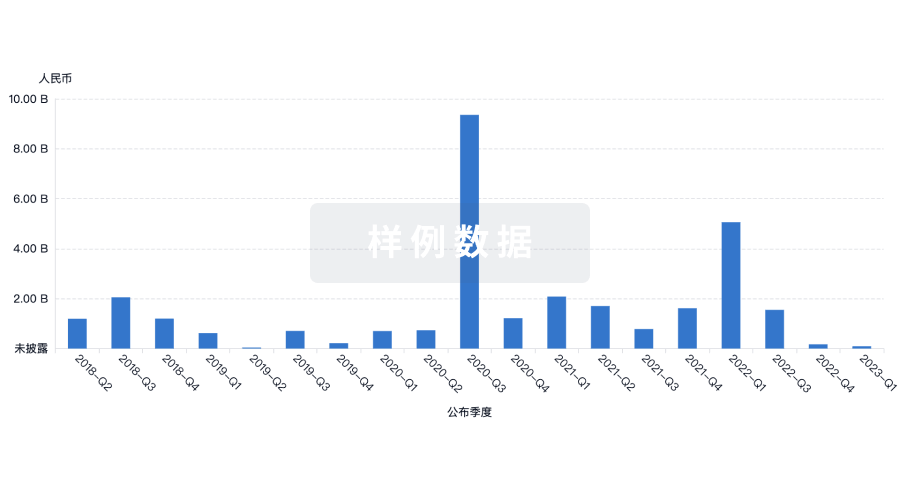
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
