预约演示
更新于:2026-06-03
Saquinavir Mesylate
甲磺酸沙奎那韦
更新于:2026-06-03
概要
基本信息
药物类型 小分子化药 |
别名 Fortovase、SAQUINAVIR、Saquinavir mesilate (JAN) + [11] |
作用方式 抑制剂 |
作用机制 HIV-1 protease抑制剂(HIV蛋白酶抑制剂) |
在研适应症 |
非在研适应症 |
权益机构- |
最高研发阶段批准上市 |
首次获批日期 美国 (1995-12-06), |
最高研发阶段(中国)批准上市 |
特殊审评加速批准 (美国)、孤儿药 (日本) |
登录后查看时间轴
结构/序列
分子式C39H54N6O8S |
InChIKeyIRHXGOXEBNJUSN-YOXDLBRISA-N |
CAS号149845-06-7 |
关联
81
项与 甲磺酸沙奎那韦 相关的临床试验NCT02023450
Testing of HIV Protease Inhibitors to Suppress Inflammation and Improve Cardio Pulmonary Hemodynamics in Subjects With Pulmonary Arterial Hypertension
Study Rationale:There is recent evidence that HIV protease inhibitors (HIV-PI) can improve pulmonary hemodynamics in experimental models of pulmonary arterial hypertension (PAH). There is also experimental evidence that both TLR4 and high mobility group box 1 (HMGB1) participate in the pathogenesis of experimental pulmonary hypertension. A recent high throughput screen for inhibitors of HMGB1 induced macrophage activation yielded HIV-protease inhibitors (PIs) as potent inhibitors of HMGB1 induced cytokine production. Based on the experimental evidence we propose a trial to determine whether HIV-PIs will alter the pathobiology of PAH.
Study Objectives:The main objective of this study is to determine whether saquinavir and ritonavir (SQV+RIT) which have a well-characterized safety profile in humans will reduce bio markers of inflammation and pulmonary artery pressures in patients with PAH.
Study Hypothesis:We hypothesize that the HIV-PI, SQV+RIT, will reduce circulating parameters of inflammation including HMGB1, IL1-beta, IL-6, IL-8, IL-10, TNF-alpha and CRP. Our end points will be changes in these parameters from baseline over the duration of the study.We hypothesize that treatment with SQV+RIT will reduce pulmonary artery(PA) pressure of patients with PAH as measured by echocardiography.
Study Design:This is a single center open label phase 0 study to evaluate the effect of SQV +RIT in patients with IPAH. Subjects with IPAH(N=20) will be enrolled into a study, which will be divided into 3 cohorts and entail the administration of HIV protease inhibitors in three doses. The first cohort (n=3) will receive a starting dose of SQV 0.3 mg/kg twice daily in combination with RIT 0.03 mg/kg twice daily. If the first dose is well-tolerated, the second cohort (n= 3 ) with IPAH will be given doses of SQV 3 mg/kg and RIT 0.3 mg/kg twice daily. If the second dose is well-tolerated, the last cohort (n= 14 ) with IPAH will be given doses of SQV 15 mg/kg and RIT 1.5 mg/kg twice daily.
Study Objectives:The main objective of this study is to determine whether saquinavir and ritonavir (SQV+RIT) which have a well-characterized safety profile in humans will reduce bio markers of inflammation and pulmonary artery pressures in patients with PAH.
Study Hypothesis:We hypothesize that the HIV-PI, SQV+RIT, will reduce circulating parameters of inflammation including HMGB1, IL1-beta, IL-6, IL-8, IL-10, TNF-alpha and CRP. Our end points will be changes in these parameters from baseline over the duration of the study.We hypothesize that treatment with SQV+RIT will reduce pulmonary artery(PA) pressure of patients with PAH as measured by echocardiography.
Study Design:This is a single center open label phase 0 study to evaluate the effect of SQV +RIT in patients with IPAH. Subjects with IPAH(N=20) will be enrolled into a study, which will be divided into 3 cohorts and entail the administration of HIV protease inhibitors in three doses. The first cohort (n=3) will receive a starting dose of SQV 0.3 mg/kg twice daily in combination with RIT 0.03 mg/kg twice daily. If the first dose is well-tolerated, the second cohort (n= 3 ) with IPAH will be given doses of SQV 3 mg/kg and RIT 0.3 mg/kg twice daily. If the second dose is well-tolerated, the last cohort (n= 14 ) with IPAH will be given doses of SQV 15 mg/kg and RIT 1.5 mg/kg twice daily.
开始日期2013-12-01 |
申办/合作机构  中南大学湘雅三医院 中南大学湘雅三医院 [+1] |
NCT01638650
To Determine the Effect of the Modified SQV/r (Saquinavir-boosted by Ritonavir) Regimen (500/100 mg for the 1st Week Followed by 1000/100 mg for the 2nd Week) on the QTc Interval, Pharmacokinetics, and Antiviral Activity in Treatment-naive HIV-1 Infected Patients
This open-label study will evaluate the safety, pharmacokinetics and antiviral activity of a modified Invirase (saquinavir)/ritonavir regimen in treatment-naïve HIV-1 infected patients. Patients will receive Invirase 500 mg plus ritonavir 100 mg twice daily orally for the first week, followed by Invirase 1000 mg plus ritonavir 100 mg twice daily orally for the second week. The study treatment will be given in combination with two Nucleoside Reverse Transcriptase Inhibitors (NRTIs), in accordance with the current clinical HIV treatment guidelines. Anticipated time on study treatment is 14 days.
开始日期2012-01-01 |
申办/合作机构 |
JPRN-UMIN000004621
Effect of a pharmaceutical excipient, cremophol EL on the plasma concentration-time profile of saquinavir and fexofenadine after their oral administrations at doses used in the exploratory investigational new drug clinical studies - Effect of cremophol EL on pharmacokinetics of saquinavir and fexofenadine at doses used in the exploratory investigational new drug clinical studies
开始日期2010-11-01 |
申办/合作机构- |
100 项与 甲磺酸沙奎那韦 相关的临床结果
登录后查看更多信息
100 项与 甲磺酸沙奎那韦 相关的转化医学
登录后查看更多信息
100 项与 甲磺酸沙奎那韦 相关的专利(医药)
登录后查看更多信息
3,567
项与 甲磺酸沙奎那韦 相关的文献(医药)2026-06-01·BIOORGANIC CHEMISTRY
Repurposing amide-based drugs unveils inhibition mechanisms of plasmodium falciparum Plasmepsin V enzymatic activity for antimalarial therapy
Article
作者: Choudhary, Sinjan ; Singh, Garima ; Lopus, Manu ; Prajapati, Anitadevi K
Understanding how enzymes are inhibited at the molecular level is essential for rational drug design against malaria parasites. Plasmodium falciparum Plasmepsin V (PfPlmV), an aspartic protease essential for exporting parasite proteins into host red blood cells has emerged as potent target for anti-malarial therapy. In this work, we investigated whether three approved amide-based drugs favipiravir, 5-fluorouracil (5-FU), and saquinavir could inhibit catalytic activity of PfPlmV. All three compounds bound to PfPlmV with affinities of ∼104 M-1, but with distinct binding behaviours. Binding by isothermal titration calorimetry (ITC) showed involvement of both hydrogen bonds and hydrophobic interactions, which stabilized the enzyme-drug complexes. Enzyme kinetics and docking studies demonstrated that the inhibitory effect depends on the degree of catalytic cleft coverage. The results showed that saquinavir interacts with key active-site residues, including the catalytic dyad Asp118 and Asp365, consistent with competitive inhibition having IC50 value of 0.35 ± 0.01 μM and a low Ki value of (0.14 ± 0.02) μM. 5-FU interacts only near Asp118, resulting in mixed-type inhibition with higher IC50 and Ki values. Favipiravir binds at the periphery of the cleft without engaging the catalytic dyad and therefore behaves as a weak, non-inhibitory binder. The insights of interaction-structure-inhibitory activity obtained from this study may guide repurposing strategies to develop selective potent inhibitors/antimalarial agents targeting PfPlmV.
2026-05-01·EUROPEAN JOURNAL OF PHARMACOLOGY
Downregulation of claudin-2 expression and chemoresistance by saquinavir in human lung adenocarcinoma cells
Article
作者: Shirouzu, Mikako ; Morimoto, Kazushi ; Yoshino, Yuta ; Kimura, Riho ; Komoto, Sayaka ; Endo, Satoshi ; Ikari, Akira ; Matsunaga, Toshiyuki ; Ishikawa, Yoshinobu ; Shinoda, Takehiro
Claudin-2 (CLDN2) is a tight junction-associated transmembrane protein that is highly expressed in lung adenocarcinoma and contributes to chemoresistance. Here, we screened Food and Drug Administration-approved small-molecule drugs to identify compounds capable of reducing CLDN2 expression and enhancing anticancer drug-induced cytotoxicity in cancer cells. A structure based virtual screening targeting CLDN2 predicted that saquinavir (SQV), a human immunodeficiency virus (HIV) protease inhibitor, binds to the second extracellular loop of CLDN2 protein. Consistent with the prediction, SQV markedly decreased CLDN2 protein levels without affecting its mRNA expression in human lung adenocarcinoma A549 cells. In contrast, ritonavir, another HIV protease inhibitor, or raltegravir, an HIV integrase inhibitor, has no significant effect, indicating that inhibition of viral enzymes is not directly involved in the regulation of CLDN2 expression. A cycloheximide-chase assay revealed that SQV reduces the stability of CLDN2 protein. Pharmacological inhibition experiments further revealed that SQV-induced CLDN2 downregulation is mediated by enhanced clathrin-dependent endocytosis and subsequent lysosomal degradation. In three-dimensional spheroid models, SQV reduced oxidative stress and expression of nuclear factor erythroid 2-related factor 2 protein expression without altering spheroid size, cell viability, or hypoxic stress. Notably, SQV enhanced the cytotoxic effects of multiple anticancer agents, including doxorubicin, cisplatin, and SN-38, in lung adenocarcinoma cell line-derived spheroids and patient-derived organoids. We suggest that SQV improves chemoresistance by reducing CLDN2 expression and oxidative stress in lung adenocarcinoma.
2026-04-13·JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS
Insights into the interaction mechanism of first-generation HIV-1 protease inhibitors with wild-type and mutant (D30N and L76V) enzymes through
in-silico
based approach
Article
作者: S, Pandarinathan ; S, Jayanthi
The combined effort of medical, scientific and pharmaceutical research has transformed the HIV infection from an inevitably fatal disease into a manageable chronic ailment. Especially after the introduction of protease (PR) inhibitors the efficacy of the treatment regimen has been increased. However, the development of mutant PR enzymes is a major concern. The present study focuses on to understand the alterations in wild-type and mutants (D30N and L76V) PR structure by interaction of first-generation PR inhibitors amprenavir, indinavir, nelfinavir, ritonavir and saquinavir. This information is extracted using molecular docking and MD simulation. Further, post-MD analysis such as RMSD, RMSF, PCA, DCCM, intermolecular interactions and binding free energy values calculation has been done. In comparison to L76V, D30N mutation affected the stable binding of the most of the studied PR inhibitors. The positively charged NH2 and thiazole group of amprenavir and nelfinavir is repelled by NH2 of Asn30 in D30N mutant. This repulsion pushes the drugs towards flap domain and causes flap opening. The MD simulation reveals that amprenavir, nelfinavir and saquinavir drugs have good compatibility with L76V mutant PR enzyme whereas indinavir was affected by the same. L76V mutant does not form any direct interaction with the first line drugs; hence the drugs forming strong interactions with the active site, flap domain and substrate binding region residues are quite enough to manage the resistance provided by L76V mutant. The results provided the insight into the drug-resistant or drug-susceptibility mechanism of L76V and D30N mutation against first-generation PR inhibitors.
103
项与 甲磺酸沙奎那韦 相关的新闻(医药)2026-06-01
在抗病毒药物的璀璨星河中,利托那韦(Ritonavir)无疑是一颗极具传奇色彩的星辰。从抗艾滋病一线药物到新冠治疗的关键辅助用药,从研发突破到晶型危机,再到临床角色的华丽转型,它走过了近三十年的跌宕历程,见证了抗病毒药物研发的技术迭代,也改写了全球感染性疾病的治疗格局。一、前世:艾滋病阴霾下的研发突围(1980-1996)(一)时代背景:艾滋病的世纪恐慌
1981 年,全球首例艾滋病病例被确诊;1983 年,人类免疫缺陷病毒(HIV)被证实为艾滋病的病原体。此后十年,艾滋病在全球迅速蔓延,成为 “世纪瘟疫”。彼时,治疗手段极度匮乏,仅有齐多夫定等少数核苷类逆转录酶抑制剂,疗效有限且极易耐药,患者生存期极短,艾滋病等同于 “绝症”。(二)靶点突破:HIV 蛋白酶的致命弱点
HIV 的复制依赖蛋白酶—— 一种切割病毒多聚蛋白前体的关键酶,只有被切割后,病毒才能组装成具有感染性的新病毒颗粒。1990 年代初,科学家发现 HIV 蛋白酶是C2 对称二聚体结构,且其活性位点具有高度特异性,这为药物设计提供了精准靶点。
美国雅培制药公司(Abbott)率先启动研发,采用基于结构的药物设计策略:模拟蛋白酶底物的肽键结构,设计 “过渡态类似物”,让药物精准结合蛋白酶活性位点,阻断其功能。经过多轮结构优化 —— 调整分子对称性、引入亲水基团提升溶解度、优化疏水 - 亲水平衡增强细胞膜通透性 —— 最终筛选出利托那韦。(三)重磅上市:首个蛋白酶抑制剂诞生
1995 年 12 月,利托那韦提交新药申请(NDA);1996 年 3 月,获美国 FDA 批准上市,成为全球首个 HIV 蛋白酶抑制剂,商品名 Norvir。同年,它被列入世界卫生组织基本药品目录,标志着艾滋病治疗进入高效抗逆转录病毒疗法(HAART,即 “鸡尾酒疗法”) 时代。
上市初期,利托那韦以600mg 每日两次的高剂量单独或联合用药,能快速降低患者体内 HIV 病毒载量,显著延长生存期、提升生活质量。它的出现,让艾滋病从 “绝症” 变为可长期控制的慢性疾病,堪称艾滋病治疗史上的里程碑。二、波折:致命晶型危机与临床转型(1998-2020)(一)惊天危机:晶型突变引发全球撤市
1998 年初,上市仅两年的利托那韦遭遇制药史上最严重的晶型危机。药物在储存过程中,稳定晶型(Form I)逐渐转化为更稳定但溶解度骤降的新晶型(Form II)。这一转变导致胶囊制剂中药物大量析出,溶出度严重不达标,药效大幅丧失。
更棘手的是,新晶型具有 “传染性”—— 一旦在生产环境中出现,会快速污染所有批次产品。1998 年 10 月,雅培被迫全球召回所有胶囊制剂,仅保留稳定性稍好的口服溶液(需 2-8℃冷藏)。此次危机导致雅培损失超 2.5 亿美元,全球艾滋病治疗一度陷入混乱,无数患者面临断药风险。(二)浴火重生:从主力药到 “增效剂”
危机之后,雅培耗时数年重新优化制剂,最终推出片剂(2000 年后),通过工艺控制稳定晶型,解决溶解度问题。但临床应用格局已彻底改变:
高剂量单药淘汰:长期高剂量使用利托那韦,易引发胃肠道反应(恶心、腹泻)、脂肪代谢障碍、血脂升高等不良反应,且病毒易产生耐药性。
低剂量增效崛起:科学家发现,利托那韦是CYP3A4 肝药酶的强效抑制剂—— 低剂量(100-200mg)即可显著抑制其他蛋白酶抑制剂(如沙奎那韦、阿扎那韦)的代谢,提升其血药浓度、延长作用时间、减少用药剂量。
自此,利托那韦的临床角色从一线治疗主力转型为药代动力学增效剂(Booster),成为艾滋病 “鸡尾酒疗法” 中不可或缺的 “黄金配角”。这一转型大幅降低用药副作用、延缓耐药性,至今仍是 HIV 治疗的核心方案之一。三、今生:新冠疫情中的 “跨界” 重生(2020 - 至今)(一)意外发现:抗新冠的 “隐藏技能”
2020 年新冠疫情暴发后,科学家紧急筛选抗病毒药物,发现新冠病毒(SARS-CoV-2)的主蛋白酶(3CLpro)与 HIV 蛋白酶结构相似,均为病毒复制的关键靶点。而利托那韦虽对 3CLpro 抑制活性较弱,但其强效抑制 CYP3A4的特性,可完美辅助其他 3CLpro 抑制剂发挥作用。(二)巅峰时刻:Paxlovid 的核心搭档
2021 年,辉瑞公司研发出奈玛特韦(Nirmatrelvir)—— 一种高活性的新冠 3CLpro 抑制剂,但它易被 CYP3A4 代谢,单独口服生物利用度低、药效短暂。于是,奈玛特韦 + 利托那韦(300mg/100mg) 的复方制剂Paxlovid应运而生。
利托那韦在复方中承担 “保镖” 角色:抑制 CYP3A4,阻止奈玛特韦被快速代谢,使其血药浓度提升数倍、半衰期延长至 12 小时,实现每日两次口服的便捷给药。临床试验证实,Paxlovid 可将高风险新冠患者的重症 / 死亡风险降低 89%,对 Delta、Omicron 等变异株均有效。
2021 年 12 月,Paxlovid 获美国 FDA 紧急使用授权;2022 年 2 月,中国获批进口,成为国内首个口服抗新冠特效药,在疫情防控关键阶段挽救无数重症患者生命。利托那韦以 “跨界” 身份,再次站在全球抗病毒治疗的舞台中央。(三)现状与价值:从抗艾到广谱抗病毒
如今,利托那韦的应用已超越艾滋病与新冠:
艾滋病治疗:仍作为增效剂,与多种蛋白酶抑制剂联用,是耐药 HIV 感染的核心治疗方案。
新冠治疗:作为 Paxlovid 的关键组分,仍是高风险患者的首选口服药,同时用于新冠暴露后预防。
潜在拓展:研究显示,利托那韦对丙肝病毒、呼吸道合胞病毒等多种病毒的蛋白酶均有抑制作用,未来有望用于更多病毒性疾病的治疗。四、结语:一颗药物的传奇与启示
从 1996 年首个抗艾蛋白酶抑制剂,到 1998 年晶型危机的浴火重生,再到 2022 年新冠治疗的跨界封神,利托那韦的三十年历程,是药物研发 “结构设计-临床转化-危机迭代-角色升级” 的缩影。
它的故事告诉我们:药物研发从来不是一帆风顺的直线,而是在挫折中迭代、在转型中重生的过程。同时,利托那韦从 “主力药” 到 “增效剂” 再到 “跨界明星” 的转变,也印证了老药新用的巨大价值 —— 经典药物的潜在作用,往往能在新的疾病挑战中绽放新的光芒。
未来,随着抗病毒药物研发的持续深入,利托那韦或许会迎来更多角色转变,但它在艾滋病与新冠治疗史上留下的浓墨重彩,以及为药物研发带来的深刻启示,将永远镌刻在医学发展的长河中。
2026-05-19
New scientific software enables researchers to explore chemical and genetic perturbations, accelerating target discovery and drug repurposing
MOUNTAIN VIEW, Calif., May 19, 2026 /PRNewswire/ -- Addressing the persistent challenge scientists face in transforming complex biological
perturbation data into actionable
mechanistic insight, DataXight today announced protoXell, a new scientific software designed to streamline discovery. To learn more about protoXell and explore access options, visit .
protoXell enables scientists to move beyond data processing toward
biological understanding—allowing them to
compare perturbation effects across experiments through an intuitive,
point-and-click
environment, uncover
pathway-level responses, and generate hypotheses for
target discovery and drug repurposing. The system supports analyses that are often
impractical, even for well-resourced computational teams, due to the scale and complexity of modern perturbation datasets.
DataXight will formally introduce protoXell at the
Bio-IT World Conference & Expo 2026, May 19–21, 2026, where attendees can experience how the software supports mechanism discovery at scale.
"At DataXight, we've spent years helping scientists navigate the complexity of biological data," said Tuan Nguyen, CEO of DataXight. "With protoXell, we are enabling researchers not just to simplify analysis, but to perform analyses that were previously out of reach—helping them move from
perturbation data to mechanistic understanding and better
therapeutic decisions."
Addressing a Critical Bottleneck
Advances in
single-cell perturbation technologies have generated massive datasets spanning hundreds of millions of cells across thousands of perturbations, but translating these into actionable insight remains challenging. Current approaches require
complex pipelines, integration work, and specialized expertise, slowing analysis and limiting exploration.
protoXell addresses this gap with an integrated environment for exploring perturbation data. Researchers can compare perturbation effects, understand pathway responses, and generate hypotheses in real time—
without pipelines or code. Built-in
AI assistance interprets results in biological context, connecting molecular signals to mechanisms and scientific literature. By removing
technical barriers, protoXell makes previously difficult analyses accessible, so scientists can focus on discovery, not infrastructure.
Key Capabilities
Curated Perturbation Catalog – Access analysis-ready datasets spanning 150+ million cells across cell lines, tissues, compounds, and CRISPRi perturbations.
Comparative Analytics – Identify shared and distinct biological responses across genes, compounds, and cell types.
AI-Powered Interpretation – inXighter translates results into biological meaning, connecting signals to mechanisms and literature.
Enterprise Data Integration – Integrate proprietary and public datasets across cloud and on-premise environments.
Example Insight
protoXell can reveal
shared transcriptional responses between compounds with distinct pharmacology—such as similarities observed between the HIV protease inhibitor Saquinavir and β-adrenoceptor agonists—highlighting
biological connections that may inform
mechanism-of-action studies and drug repurposing.
Availability
protoXell is available with a
free access option, flexible licensing, and
enterprise deploymen
t across cloud and on-premise environments. Availability on DNAnexus is planned for June 2026. For further details on features and deployment options, please visit .
To request access, contact [email protected].
About DataXight
DataXight () is an engineering and software company dedicated to transforming
biological data into actionable insight, supporting the journey from data generation to discovery.
Product Announcement
SOURCE DataXight
21%
more press release views with
Request a Demo
2026-05-19
·杨扬DY
摘要病毒作为一类独特的亚微观感染因子,既是人类疾病的重要病原体,也是推动生命科学发展的关键研究对象。本文从病毒发现的历史进程出发,系统梳理了从19世纪末烟草花叶病毒的发现到当代宏基因组学驱动的病毒组学革命;阐述了基于基因组特征的巴尔的摩分类法和国际病毒分类委员会的系统分类框架;深入分析了抗病毒药物从核苷类似物到精准靶向药物的发展逻辑,以及疫苗技术从传统灭活疫苗到mRNA平台的范式跃迁。文章还专门讨论了致癌病毒的特殊地位及其机制,并展望了广谱抗病毒策略、人工智能辅助药物设计和治疗性疫苗等前沿方向。全文试图展现一条清晰的主线:人类对病毒的认知经历了从“不可见的病原体”到“可解析的分子机器”的转变,而治疗策略则从经验摸索走向基于机制的理性设计。关键词:病毒发现史;病毒分类;抗病毒药物;疫苗技术;致癌病毒;广谱抗病毒一、引言:病毒研究的双重意义病毒是地球上数量最庞大的生物实体之一,也是人类疾病的重要病因。天花、狂犬病、流感、艾滋病、病毒性肝炎、宫颈癌——这些疾病名称背后,都潜藏着病毒的身影。2020年以来,新型冠状病毒(SARS-CoV-2)引发的全球大流行更是以极为直接的方式提醒世人:病毒不仅是生命科学的研究对象,更是关乎公共卫生乃至文明运行的根本变量。然而,病毒研究的意义远不止于疾病防控。自20世纪中叶以来,病毒始终是生命科学前沿突破的重要推手。噬菌体感染实验帮助确立了DNA作为遗传物质的地位;逆转录酶的发现颠覆了分子生物学的“中心法则”;对病毒癌基因的研究打开了人类认识癌症分子机制的大门;基于病毒载体和mRNA的疫苗平台则正在重塑传染病防控和肿瘤免疫治疗的格局。病毒既是人类健康的威胁,也是科学发现的富矿。本文试图以历史演进和科学逻辑交织的方式,系统介绍病毒的发现与分类、治疗方案与药物研制的发展历程与前沿进展。全文贯穿一条主线:人类对病毒的认知经历了从“不可见的病原体”到“可解析的分子机器”再到“可干预的生物靶标”的转变,治疗策略也从经验摸索走向机制导向的理性设计。这一历程,既是病毒学的科学史,也是药物研发的方法论演进。二、病毒的发现:从过滤实验到宏基因组学2.1 病毒概念的诞生:微生物学时期的奠基人类与病毒的遭遇远早于对其本质的认知。考古学家在三千年前的古埃及法老拉美西斯五世木乃伊上发现了天花发作的痕迹,这是已知最早的天花病例记录。中国明朝时期已使用人痘接种法预防天花,1798年英国医生詹纳发明了更为安全的牛痘接种法,1885年法国科学家巴斯德研制出狂犬病疫苗。然而,当时人们并不知道这些疾病的病原体究竟是什么。病毒概念的诞生,根植于19世纪末微生物学的繁荣。德国科学家科赫于1877年拍摄了第一张细菌显微镜照片并发明了细菌培养技术,使得大量病原菌相继被发现。但研究者很快遇到了一类“滤过性”病原体:它们能通过截留细菌的过滤器,却仍然具有感染性。1892年,俄国科学家伊万诺夫斯基在研究烟草花叶病时发现,将患病植物叶片的汁液通过细菌过滤器后,滤液仍能使健康植物致病。1898年,荷兰微生物学家贝耶林克独立地重复了这一实验,并首次提出了“传染性活液”的概念,将这种病原体命名为“virus”(拉丁语意为“毒物”),标志着病毒学作为一门独立学科的建立。同年,口蹄疫病毒成为第一个被发现的动物病毒;1901年,黄热病病毒成为第一个被确认的人类病毒。2.2 生物化学时期:揭示病毒的物理化学本质进入20世纪30年代,病毒学研究从“滤过性”的模糊概念转向对其物质构成的精确解析。1935年,美国微生物化学家斯坦利成功分离并结晶了烟草花叶病毒,证明病毒具有蛋白质的化学属性,这一工作使他获得了1946年的诺贝尔化学奖。随后,科学家们进一步将烟草花叶病毒分离为蛋白质外壳和核酸核心,明确了病毒的基本化学组成——蛋白质与核酸的复合体。这一时期,组织培养技术的创新为病毒研究提供了关键工具。研究者可以在实验室中大量培养病毒,从而进行系统的生物学分析。噬菌体(感染细菌的病毒)成为这一时期最重要的研究模型:噬菌体感染复制实验不仅揭示了病毒的生命周期,更在1952年通过赫尔希和蔡斯的著名实验,以放射性同位素标记的方法确证了DNA是遗传物质——这一发现直接推动了分子生物学的诞生。2.3 遗传学时期:病毒基因组与癌基因的发现20世纪60至70年代,病毒学研究进入遗传学阶段。科学家们开始深入探究病毒的生命周期、感染机制及其与宿主细胞的相互作用。这一时期的标志性成就是1970年逆转录酶的发现:巴尔的摩和特明各自独立地发现了RNA依赖的DNA聚合酶,揭示了逆转录病毒将RNA基因组逆转录为DNA并整合入宿主染色体的独特复制策略。这一发现不仅修正了经典分子生物学的“中心法则”(DNA→RNA→蛋白质),也为后来理解HIV的致病机制和开发抗逆转录病毒药物奠定了理论基础。同一时期,病毒致癌机制的研究取得突破性进展。1964年,Epstein-Barr病毒(EBV)成为首个被确认的人类肿瘤病毒,从Burkitt淋巴瘤细胞中被分离出来。这标志着“病毒致癌”假说从动物模型拓展到了人类疾病,开启了肿瘤病毒学研究的新纪元。1976年,Harald zur Hausen提出人乳头瘤病毒(HPV)可能导致宫颈癌的假说,这一洞见后来被大量流行病学和分子生物学证据所证实,并直接推动了HPV疫苗的研发。2.4 分子生物学时期:从基因组到精准干预20世纪80年代以后,重组DNA技术、PCR扩增和DNA测序等分子生物学工具的成熟,使病毒学研究进入了一个全新阶段。1977年,科学家首次完成了噬菌体φX174的完整基因组测序;此后,越来越多病毒的基因组被解析,包括HIV、HBV、HCV等重要人类病原体。基因组信息的获取使得研究者能够精确地鉴定病毒蛋白的结构与功能,为基于结构的药物设计提供了分子蓝图。这一时期,抗病毒药物和疫苗技术取得了前所未有的突破。基于对HIV蛋白酶三维结构的解析,科学家通过计算机辅助药物设计开发了蛋白酶抑制剂,与逆转录酶抑制剂联合使用形成了“高效抗逆转录病毒疗法”(HAART),将HIV感染从致命疾病转变为可长期管理的慢性病。在疫苗领域,重组DNA技术催生了乙肝疫苗等新型疫苗,而21世纪第二个十年间mRNA疫苗技术的成熟,更是让疫苗研发的范式发生了根本性变革。这一发展脉络充分说明,病毒学已从描述性科学转变为以分子机制为核心、以精准干预为目标的现代生物医学学科。2.5 宏基因组学与新病毒的发现传统病毒发现依赖分离培养和电子显微镜观察,这一方法存在根本性局限:绝大多数病毒无法在实验室培养。宏基因组学(metagenomics)的兴起彻底改变了这一局面。该技术通过直接从环境样本或临床样本中提取总核酸进行高通量测序,无需预先培养即可鉴定其中存在的病毒基因组序列。近年来,宏基因组学在新病毒发现方面取得了一系列重要成果。2025年,中国医学科学院团队发布了ZOVER 2.0数据库,整合了全球72项独立研究的5883个高质量宏基因组测序数据集,覆盖362种野生动物及媒介生物,共鉴定出110个病毒科、409个属及约4400个病毒种,其中包含30个新的哺乳动物相关病毒属。同年,研究者利用宏病毒组测序,在儿童脑炎患者的脑脊液中发现了一种全新的未分类RNA病毒(hucaurvirus),其基因组结构前所未见,首次证实该类病毒可侵入人类中枢神经系统。此外,新开发的BEREN工具从环境宏基因组中识别出230种海洋巨型病毒,极大地拓展了人们对病毒多样性的认知边界。宏基因组学不仅加速了新病毒的发现,还改变了病毒学的研究范式:从“先分离后鉴定”转变为“先测序后验证”,从“以疾病为中心”转变为“以生态系统为中心”。病毒组学(virome)概念的形成,促使人们将病毒视为生物圈的固有组成部分,而非仅仅是需要剿灭的“入侵者”。三、病毒分类体系:从形态描述到基因组逻辑3.1 巴尔的摩分类法:以分子逻辑理解病毒多样性1971年,诺贝尔奖得主David Baltimore提出了一种全新的病毒分类框架,后来被称为巴尔的摩分类法(Baltimore Classification)。该体系的核心思想简洁而深刻:不同病毒的基因组类型决定了它们产生信使RNA(mRNA)的不同途径,而mRNA是一切蛋白质表达的共同起点。通过梳理病毒从基因组到mRNA的信息传递路径,就可以将纷繁复杂的病毒世界纳入一个逻辑清晰的分类体系。巴尔的摩分类法将病毒分为七个组:第I类:双链DNA病毒(dsDNA) 。其基因组与宿主细胞的DNA形式相同,可以直接利用宿主或自身编码的DNA依赖的RNA聚合酶转录出mRNA。代表病毒包括腺病毒、疱疹病毒和痘病毒。疱疹病毒感染可导致单纯疱疹和水痘,而人乳头瘤病毒(HPV)的某些高危型别则是宫颈癌的主要病因。第II类:单链DNA病毒(ssDNA) 。其单链DNA基因组进入细胞后,首先需合成互补链形成双链DNA中间体,再以此为模板转录mRNA。代表病毒为细小病毒B19,可引起幼儿传染性红斑。第III类:双链RNA病毒(dsRNA) 。其双链RNA基因组直接作为模板,由病毒自身携带的RNA依赖的RNA聚合酶转录产生mRNA。代表病毒为轮状病毒,是全世界婴幼儿严重腹泻的主要病原体。第IV类:正链单链RNA病毒((+)ssRNA) 。其基因组RNA本身即可直接作为mRNA被核糖体翻译,因此感染后无需携带聚合酶即可启动蛋白质合成。代表病毒包括引起普通感冒的鼻病毒、导致手足口病的肠道病毒,以及登革病毒和寨卡病毒等。第V类:负链单链RNA病毒((−)ssRNA) 。其基因组RNA与mRNA互补,不能直接翻译,必须先由病毒携带的RNA依赖的RNA聚合酶转录出互补的正链mRNA。代表病毒包括流感病毒(正黏病毒科)、狂犬病毒(弹状病毒科)和埃博拉病毒等。流感病毒的RNA聚合酶缺乏校对能力,其高突变率是抗原漂移的结构基础。第VI类:逆转录单链RNA病毒(ssRNA-RT) 。其正链RNA基因组通过逆转录酶合成双链DNA中间体,后者整合入宿主染色体,再由宿主RNA聚合酶转录产生mRNA和子代基因组RNA。代表病毒为人类免疫缺陷病毒(HIV),这一独特的复制策略是抗HIV药物设计的核心靶点。第VII类:逆转录双链DNA病毒(dsDNA-RT) 。其双链DNA基因组在复制过程中经历RNA中间体阶段,由病毒自身的逆转录酶以RNA为模板合成子代DNA基因组。代表病毒为乙型肝炎病毒(HBV),其复制机制与HIV有相似之处,因此部分核苷酸类似物(如替诺福韦)对二者均有效。巴尔的摩分类法的核心价值在于,它将病毒的复制策略与基因组类型直接关联,使研究者能够根据病毒的分类归属推断其基本生物学行为,从而为抗病毒药物设计提供逻辑起点。需要注意的是,巴尔的摩分类法与ICTV分类法并非竞争关系,而是互补关系:巴尔的摩分类法以基因组逻辑解释病毒复制策略的共性,ICTV分类法则以系统发育关系刻画病毒演化的个性。两者共同构成了现代病毒学研究的基石。3.2 ICTV分类系统:构建病毒世界的系统发育图谱与巴尔的摩分类法的功能导向不同,国际病毒分类委员会(ICTV)构建的是一套基于进化关系的系统分类体系。ICTV成立于1966年,是全球唯一负责制定和维护病毒分类标准的权威机构,其分类框架囊括了从域(realm)到种(species)的完整层级结构。ICTV分类体系的设计理念与经典生物分类学一致:通过形态特征、基因组结构、蛋白质序列相似性和系统发育关系等综合信息,将病毒归入域(-viria)、界(-virae)、门(-viricota)、纲(-viricetes)、目(-virales)、科(-viridae)、属(-virus)和种的层级中。截至2023版,ICTV分类表已包含6个域、10个界、18个门、314个科、3522个属、14690个种。ICTV分类体系处于持续更新之中。2025年发布的最新版分类(MSL40)新增了1563个病毒新种、243个属、55个科、11个目和8个纲。其中一个引人瞩目的变化是创建了新的域Singelaviria,其依据是衣壳蛋白采用单层果冻卷折叠结构,与已有域Varidnaviria中双层果冻卷折叠的衣壳蛋白在结构和进化上截然不同。此外,感染脊椎动物的单链DNA指环病毒被归入新的门Commensaviricota,而感染超嗜热古菌的病毒也被赋予了独立的分类学地位。ICTV分类体系的重要性在于,它为全球病毒学研究提供了统一的命名标准和交流语言,使不同实验室的研究成果可以在此框架下进行比较和整合。NCBI等公共数据库已开始实施ICTV最新分类标准的更新,进一步促进了病毒基因组数据的标准化管理。四、抗病毒药物的研制:从偶然发现到理性设计4.1 抗病毒药物研发的历史里程碑与抗生素的蓬勃发展相比,抗病毒药物的研发起步晚、进展慢。根本原因在于病毒作为专性细胞内寄生虫,其复制过程与宿主细胞代谢紧密交织,很难找到只杀伤病毒而不伤害宿主的选择性靶点。直到20世纪60年代,第一个抗病毒药物碘苷(idoxuridine)才获得批准,用于局部治疗单纯疱疹病毒引起的眼部感染。真正开启现代抗病毒药物时代的里程碑事件发生在20世纪70年代末。研究者发现,无环核苷类似物阿昔洛韦(acyclovir)能够在远低于影响细胞DNA合成的浓度下抑制单纯疱疹病毒的DNA复制。这一选择性的结构基础令人意外却富有启发性:阿昔洛韦需要被病毒编码的胸苷激酶特异性地磷酸化激活,才能转化为活性代谢物——这意味着未感染细胞几乎不受影响。阿昔洛韦的成功确立了一个基本原则:利用病毒特有的酶或生物学过程作为药物靶点,可以实现高选择性的抗病毒效果。20世纪80年代,艾滋病疫情的出现对抗病毒药物研发提出了前所未有的紧迫需求。1983年HIV被确认为艾滋病的病原体后,药物筛选迅速展开。1987年,第一个抗HIV药物齐多夫定(AZT)获批上市,它是一种核苷类逆转录酶抑制剂,通过在病毒逆转录过程中掺入DNA链并导致链终止来阻断病毒复制。AZT的快速开发成功得益于此前多年的核苷类似物研究积累:早在1964年,该化合物便已被合成,而HIV的发现为其提供了明确的临床靶点。进入90年代,两个关键进展推动了抗病毒药物研发的第二次飞跃。第一个是结构生物学和计算机辅助药物设计的兴起:在解析HIV蛋白酶三维结构的基础上,研究者利用过渡态模拟策略设计出了沙奎那韦等蛋白酶抑制剂。蛋白酶抑制剂与核苷类逆转录酶抑制剂的联合使用构成了“高效抗逆转录病毒疗法”(HAART),将HIV感染从致命性疾病转变为可长期管理的慢性病。第二个是对病毒生命周期的深入理解带来了作用机制各异的药物:融合抑制剂阻断病毒进入细胞,整合酶抑制剂阻止病毒DNA整合入宿主基因组,CCR5拮抗剂则通过阻断病毒共受体来防止感染。到2007年,已有超过40种抗病毒药物获得批准。丙型肝炎病毒(HCV)的治疗则堪称抗病毒药物研发最具戏剧性的成功案例。2011年,第一代直接抗病毒药物(DAAs)——丝氨酸蛋白酶抑制剂波西普韦和特拉普韦获批,标志着HCV治疗进入直接靶向病毒的时代。2013年,RNA聚合酶抑制剂索磷布韦获批,在合理联合用药条件下可实现超过90%的治愈率。此后,更高效的蛋白酶抑制剂、NS5A抑制剂和NS5B聚合酶抑制剂的组合方案,使得HCV成为人类历史上第一种可以通过药物彻底治愈的慢性病毒感染——这一成就被认为是21世纪医学的重大胜利之一。然而,抗病毒药物的成功故事并未惠及所有病毒感染。目前临床上可用的抗病毒药物仅针对HIV、HCV、HBV、疱疹病毒(HSV/VZV/CMV)和流感病毒等少数几种病原体。许多危害严重的高致病性病毒——如埃博拉病毒、登革病毒、克里米亚-刚果出血热病毒等——至今缺乏有效的治疗药物。每年登革病毒在100多个国家感染数百万人,造成约25000人死亡。这个缺口提醒我们,抗病毒药物的“弹药库”仍然远未充实。4.2 主要药物类别与作用机制现有抗病毒药物按其作用靶点可以大致分为两类:靶向病毒蛋白的药物和靶向宿主因子的药物。前者以病毒特有的酶或结构蛋白为靶点,选择性高、安全性好;后者以病毒复制所依赖的宿主细胞蛋白为靶点,具有广谱潜力但选择性挑战更大。核苷/核苷酸类似物是目前应用最广泛的一类抗病毒药物。它们在结构上模拟天然的核苷或核苷酸,在病毒基因组复制过程中被病毒聚合酶掺入正在合成的核酸链,通过链终止或诱导致死突变来阻断病毒复制。阿昔洛韦(抗HSV)、更昔洛韦(抗CMV)、替诺福韦(抗HIV和HBV)、索磷布韦(抗HCV)和瑞德西韦(曾用于COVID-19)都是这一策略的成功案例。这类药物的选择性来自两个层面:其一,病毒聚合酶对类似物的亲和力往往高于宿主聚合酶;其二,部分药物(如阿昔洛韦)需要病毒编码的激酶进行特异性磷酸化激活,从而仅在感染细胞中转化为活性形式。非核苷类聚合酶抑制剂通过与病毒聚合酶的变构位点结合,诱导构象变化从而抑制酶活性。这类药物不掺入核酸链,也不依赖磷酸化激活。HIV非核苷类逆转录酶抑制剂(如依法韦仑、奈韦拉平)和HCV NS5B聚合酶非核苷抑制剂(如达沙布韦)是代表性药物。蛋白酶抑制剂的研发是结构导向药物设计的经典案例。HIV的天冬氨酰蛋白酶负责将病毒编码的多聚蛋白前体切割为成熟的结构蛋白和功能蛋白,是病毒组装成熟的关键环节。研究者基于该酶的三维结构,设计了一类模拟其天然底物过渡态的拟肽类化合物,以不可裂解的化学键替换肽键,从而竞争性地占据酶的活性中心。沙奎那韦、利托那韦、阿扎那韦和达芦那韦等HIV蛋白酶抑制剂,以及格佐匹韦、格拉瑞韦等HCV NS3/4A蛋白酶抑制剂,均诞生于这一设计理念。整合酶抑制剂阻断HIV整合酶将病毒cDNA整合到宿主细胞染色体中的过程。这一步骤是逆转录病毒建立持续感染的分子基础,也是逆转录酶抑制剂和蛋白酶抑制剂均无法干预的环节。雷特格韦、度鲁特韦等整合酶链转移抑制剂通过螯合整合酶活性位点中的镁离子来抑制链转移反应,为抗HIV治疗提供了全新的药物类别。进入抑制剂与融合抑制剂作用于病毒感染的最早期阶段。马拉维若(maraviroc)是一种CCR5拮抗剂,通过与HIV的共受体CCR5结合阻止病毒进入细胞,成为首个获批的靶向宿主蛋白的抗病毒药物。恩夫韦肽则是一种融合抑制剂,直接结合HIV的gp41蛋白并阻止其构象变化,从而阻断病毒膜与细胞膜的融合过程。神经氨酸酶抑制剂是抗流感病毒治疗的中坚力量。流感病毒表面的神经氨酸酶在子代病毒从感染细胞表面释放过程中发挥关键作用。扎那米韦和奥司他韦通过模拟神经氨酸酶天然底物(唾液酸)的过渡态结构,以“过渡态类似物”的策略竞争性抑制该酶活性,阻止病毒在呼吸道的扩散和传播。4.3 药物耐药性:进化的挑战耐药性(drug resistance)是抗病毒治疗中最令人担忧的问题之一。与抗生素耐药性类似,抗病毒药物耐药性的根源在于病毒的高突变率——尤其是RNA病毒,其RNA依赖的RNA聚合酶和逆转录酶缺乏校对功能,每个复制周期都可能引入突变。在药物选择压力下,携带耐药突变的病毒变种被筛选和富集,最终导致治疗失败。耐药问题在HIV和HBV的长期治疗中尤为突出。长期使用更昔洛韦治疗CMV感染,可能出现与病毒蛋白激酶pUL97(催化药物磷酸化第一步)或DNA聚合酶催化亚基突变相关的耐药性。长期使用核苷类似物治疗HBV,同样面临病毒聚合酶耐药突变逐步累积的问题。应对耐药的策略是多维度的。联合用药(如HAART方案)是最有效的措施:同时使用不同靶点的多种药物,使病毒需要同时积累多个耐药突变才能逃逸,从而极大提高了耐药发生的遗传屏障。此外,开发对耐药突变仍保持活性的新一代药物,以及通过治疗药物监测优化给药方案以最大限度地抑制病毒复制,都是延缓耐药发生的重要手段。近年来,针对宿主靶点的抗病毒策略在一定程度上规避了病毒耐药问题。由于宿主因子不受病毒基因组控制,病毒无法通过自身突变来绕过针对宿主蛋白的药物压力。但这一策略也面临潜在的毒性风险,因为干扰宿主蛋白的正常功能可能影响未感染细胞的生理活动。五、疫苗技术:免疫预防的范式跃迁疫苗是公共卫生领域最伟大的成就之一。从詹纳的牛痘到巴斯德的狂犬病疫苗,从灭活和减毒疫苗到重组亚单位疫苗,再到近年爆发的mRNA疫苗技术革命,疫苗研发经历了数次范式跃迁。5.1 传统疫苗平台灭活疫苗和减毒活疫苗是历史最悠久的疫苗类型,至今仍是许多疾病防控的主要手段。灭活疫苗通过化学或物理手段杀灭病毒但保留其免疫原性,安全性好但免疫原性较弱,通常需要多次接种和佐剂辅助。减毒活疫苗则通过连续传代培养筛选毒力减弱的病毒株,能够在宿主体内有限增殖,模拟自然感染过程,诱导强烈而持久的免疫应答,但存在毒力回复的潜在风险。脊髓灰质炎疫苗、麻疹疫苗、腮腺炎疫苗和水痘疫苗均属于减毒活疫苗的代表。重组亚单位疫苗是基因工程时代的产物。通过重组DNA技术将编码病毒保护性抗原(如乙肝病毒表面抗原HBsAg)的基因导入酵母或其他表达系统,大量生产纯化的病毒蛋白作为疫苗抗原。重组乙肝疫苗是这一策略的里程碑,自1986年获批以来已在全球范围内有效降低了乙肝病毒感染率和肝细胞癌发病率。近年来,病毒样颗粒(VLP)技术进一步提升了亚单位疫苗的免疫原性:通过在体外组装不含病毒基因组但保持天然构象的病毒衣壳,VLP疫苗兼具亚单位疫苗的安全性和完整病毒颗粒的免疫刺激效果——HPV疫苗(如Gardasil和Cervarix)正是这一策略的成功典范。5.2 病毒载体疫苗病毒载体疫苗利用经过改造的非致病性或减毒病毒作为“运输工具”,将目标病原体的保护性抗原基因递送至宿主细胞内表达,从而诱导免疫应答。常用的病毒载体包括腺病毒载体、痘苗病毒载体和腺相关病毒载体等。在COVID-19大流行中,阿斯利康/牛津大学的ChAdOx1腺病毒载体疫苗和强生公司的Ad26腺病毒载体疫苗均获得了紧急使用授权。2024-2025年间,病毒载体平台继续在多种传染病和肿瘤疫苗领域拓展应用,同时也在朝着提高载体安全性、降低预存免疫影响的方向持续优化。5.3 mRNA疫苗:从边缘到中心的范式革命如果说有一项技术代表了疫苗研发的范式革命,那无疑是mRNA疫苗。其原理简洁而优雅:将编码病毒保护性抗原的mRNA分子包裹在脂质纳米颗粒(LNP)中递送入人体细胞,利用宿主细胞的翻译机器在体内直接合成抗原蛋白,从而同时激活体液免疫和细胞免疫。mRNA疫苗在COVID-19大流行中以前所未有的速度完成了从实验室研发到全球大规模接种的全过程,这是人类疫苗史上的里程碑事件。然而,mRNA技术的意义绝不止于应急:它从根本上改变了疫苗研发的底层逻辑。传统疫苗需要培养病毒或表达蛋白,每种新疫苗都意味着漫长的工艺开发周期;而mRNA平台只需改变编码序列,就可以将同一套生产和递送体系应用于完全不同的病原体。这种“即插即用”的特性,使疫苗研发的速度和灵活性得到了质的飞跃。2024-2025年间,mRNA疫苗技术持续演进。国内RNA疫苗(mRNA/circRNA/saRNA)获批临床项目近20项,进入“多病种切入”和“版本更新”阶段。在新型设计方面,中国科学院团队开发了一种单链双抗原mRNA疫苗,可同时表达大别班达病毒(导致发热伴血小板减少综合征)的糖蛋白前体和核蛋白,在小鼠模型中提供了针对致死剂量病毒攻击的完全保护,且该单链设计兼具制造效率和保护效力。另一项研究则开发了一种新型LNP递送系统(WNP),使得mRNA主要在注射部位原位表达,减少了肝脏分布和促炎因子的诱导,从而提高了mRNA疫苗的安全性。自复制RNA(saRNA)疫苗是mRNA平台的重要进化方向。saRNA在编码抗原蛋白的同时,还携带编码RNA复制酶的序列,可以在宿主细胞内实现mRNA的自我扩增,从而在极低剂量下诱导持续的抗原表达和免疫应答。2023年,CSL与Arcturus therapeutics联合开发的saRNA新冠疫苗ARCT-154在日本获批上市,成为全球首个获批的saRNA疫苗。2025年9月发布的临床数据显示,saRNA狂犬病疫苗RBI-4000在接种后8个月仍可检测到中和抗体,其耐久性在多种衰减模型下与传统灭活疫苗相当或更优,证明了RNA平台从“短期应急”向“长期常规”转变的可行性。5.4 新一代疫苗技术展望环状RNA(circRNA)疫苗是值得关注的前沿方向。与线性mRNA不同,环状RNA具有共价闭合环状结构,对核酸外切酶降解具有天然抗性,因而稳定性更高、表达持续时间更长。2024-2025年间,中国已有多个circRNA项目进入临床试验阶段,适应症从传染病预防扩展到缺血性心脏病和HPV阳性实体瘤的治疗。治疗性疫苗——以已感染者或肿瘤患者为对象的疫苗——是疫苗概念的深刻拓展:它们不再用于“防病”,而是用于“治病”,标志着疫苗从公共卫生工具向个体化治疗手段的转型。六、致癌病毒:感染与肿瘤的分子桥梁6.1 肿瘤病毒概述感染性因素在癌症发生中的角色,是20世纪肿瘤生物学最重要的发现之一。早在1911年,Peyton Rous就发现过滤后的鸡纤维肉瘤提取物可以传播肿瘤,从而首次提出病毒可能致癌的假说。然而,这一假说在人类疾病中的确认经历了漫长的过程。直到1964年Epstein-Barr病毒(EBV)被确认为Burkitt淋巴瘤的病因,人类才第一次确证病毒可以直接导致人类癌症。目前,国际癌症研究机构(IARC)已将七种病毒列为第一类人类致癌物(Group 1 carcinogens):人乳头瘤病毒(HPV)、乙型肝炎病毒(HBV)、丙型肝炎病毒(HCV)、Epstein-Barr病毒(EBV)、卡波西肉瘤相关疱疹病毒(KSHV,即人类疱疹病毒8型)、人类T细胞白血病病毒1型(HTLV-1)和默克尔细胞多瘤病毒(MCPyV)。HIV虽然不直接致癌,但通过深度免疫抑制增加了多种病毒相关癌症的风险,同样被列入Group 1致癌物。据估计,全球每年约140万例新发癌症由病毒感染引起,占全部癌症病例的8%至15–20%,其中约49%的病毒相关癌症由HPV诱导。这一数字在发展中国家可能被严重低估,因为病毒相关癌症在资源匮乏地区发病率更高,而肿瘤登记系统往往不完善。6.2 病毒致癌的分子机制病毒致癌并非病毒生命周期的“目的”,而是宿主-病毒长期共进化过程中的一种偶然后果。从原发感染到癌症的发生通常有数十年的潜伏期,这一事实本身就表明,致癌是低概率事件,需要多重因素的协同作用——免疫抑制、合并感染、环境致癌物暴露以及宿主遗传易感性等都是重要的辅助因素。病毒致癌的分子机制复杂多样,概括起来包括以下几种主要模式:病毒癌基因的直接作用。某些病毒编码的蛋白可以直接干扰宿主细胞的关键信号通路和细胞周期调控,推动细胞恶性转化。最典型的例子是HPV的E6和E7蛋白:E6蛋白通过结合并促进p53抑癌蛋白的泛素化降解,使细胞丧失DNA损伤修复和凋亡能力;E7蛋白则结合并失活视网膜母细胞瘤蛋白(Rb),导致E2F转录因子持续激活,驱动细胞不受控制地进入S期。这种“双重打击”是HPV导致宫颈上皮细胞持续异常增殖的核心机制。类似地,EBV的潜伏膜蛋白LMP1可以通过模拟CD40受体的持续激活信号,促进B淋巴细胞增殖。基因组不稳定性与病毒DNA整合。HBV和HPV的基因组可以整合入宿主细胞染色体,这种整合事件是基因组不稳定的重要来源。整合可能导致宿主染色体缺失、重排或扩增,也可能导致病毒-宿主融合转录本的表达异常。HBV的HBx蛋白还具有反式激活功能,可以激活肝细胞原癌基因的表达,同时通过潜伏感染躲避免疫清除。慢性炎症与免疫抑制。HCV和HBV引起的慢性肝炎,以及EBV和KSHV的持续感染,都会维持一种低度的慢性炎症状态。长期的炎症微环境产生活性氧和促炎细胞因子,造成持续的DNA损伤和细胞代偿性增殖,为突变积累和克隆选择提供了“土壤”。HIV通过导致深度免疫缺陷,为KSHV、EBV和HPV的致癌效应创造了条件。6.3 预防与治疗策略致癌病毒的特殊之处在于,它们提供了一个极为罕见的癌症“可预防”窗口。HPV疫苗和HBV疫苗是目前最重要的两种抗癌疫苗。HPV疫苗(如Gardasil 9)通过免疫接种诱导针对HPV L1衣壳蛋白的中和抗体,阻断病毒初次感染,从而从源头预防HPV相关癌症(宫颈癌、口咽癌、肛门癌等)。在接种覆盖率高的国家和地区,宫颈癌前病变和宫颈癌的发病率已出现明显下降。HBV疫苗接种自20世纪90年代以来已被纳入全球多个国家和地区的新生儿免疫规划,使新发慢性HBV感染和相关肝癌的负担显著减轻。在治疗方面,抗病毒治疗(如HBV的核苷类似物长期抑制治疗、HCV的DAA根治性治疗)可以显著降低病毒载量、缓解肝脏炎症和纤维化,从而减少肝癌的发生风险。然而,对于已经形成的病毒相关肿瘤,抗病毒治疗仅起辅助作用,根治性治疗仍需依靠手术、放化疗和免疫治疗等综合手段。近年来的前沿探索包括免疫检查点抑制剂在病毒相关肿瘤中的应用、针对病毒抗原的过继性T细胞疗法,以及利用CRISPR/Cas9基因编辑技术清除整合的病毒基因组等。这些策略如果成熟,有望将病毒相关肿瘤的治疗提升到新的水平。七、前沿与展望:面向未来的抗病毒策略病毒学的每一次重大进步,都源于新技术的引入和新理念的确立。站在当下眺望,几个前沿方向正在塑造抗病毒研究的新图景。7.1 广谱抗病毒药物:从“一把钥匙一把锁”到“万能钥匙”传统抗病毒药物遵循“一种药物针对一种病毒”的模式,这在高变异RNA病毒面前显得捉襟见肘。甲型流感病毒因其高变异率能够轻易规避宿主免疫反应和现有药物的攻击,而当前临床使用的抗病毒药物通常只能针对病毒的单一靶点,使得病毒极易通过变异产生耐药性。广谱抗病毒药物的目标,就是打破这种“猫鼠游戏”的困境。广谱策略主要从两个方向切入。其一是寻找不同病毒之间高度保守的结构或序列元件。2025年发表的一项研究揭示了肠道病毒(包括脊髓灰质炎病毒和鼻病毒等)RNA基因组中一段保守的三叶草结构,该结构通过招募病毒融合蛋白3CD和宿主蛋白PCBP2来组装复制复合体。破坏这一保守的RNA-蛋白相互作用,有望实现对多种肠道病毒的“一网打尽”式抑制。其二是靶向病毒复制所依赖的宿主因子。宿主导向策略的优势在于,宿主蛋白不受病毒基因组控制,病毒难以通过突变来绕过针对宿主靶点的药物压力。2026年,研究者鉴定出宿主酪氨酸激酶LYN作为广谱抗流感病毒的潜在靶点:LYN可直接与流感病毒核蛋白结合并催化其特定位点磷酸化,从而破坏病毒核糖核蛋白复合物的组装,而LYN特异性激动剂MLR-1023已在体内外实验中展现出抗流感潜力。另一项研究则发现哺乳动物SKI复合物是一种广谱抗病毒靶标,通过上调细胞胆固醇水平来抑制多种病毒的复制。PROTAC(靶向蛋白降解嵌合体)技术的引入为抗病毒策略带来了全新的维度。传统抑制剂只是“占据”靶蛋白的活性位点,而PROTAC分子通过将靶蛋白与E3泛素连接酶拉近,诱导靶蛋白被蛋白酶体彻底降解。2025年,南开大学团队将PROTAC技术从单靶点推向多靶点抗病毒领域,针对甲型流感病毒开发了能够同时降解多个病毒蛋白或宿主因子的PROTAC分子,标志着抗流感策略从“单一追击”向“体系化歼灭”的跨越。7.2 人工智能与抗病毒药物设计人工智能(AI)正在深刻改变药物研发的范式。在抗病毒领域,AI的应用已经覆盖了从靶点识别、先导化合物筛选、药物性质预测到临床试验设计的全链条。AlphaFold2等蛋白质结构预测工具的突破性进展,使得研究者可以在缺乏实验结构的情况下精确预测病毒蛋白的三维构象,为基于结构的药物设计提供了前所未有的便利。AI辅助的药物重定位(drug repurposing)在COVID-19大流行中得到了广泛验证。通过机器学习模型对已批准药物的抗病毒活性进行虚拟筛选,可以快速识别出具有潜在抗病毒效果的候选药物,大幅缩短从基础研究到临床应用的转化时间。随着深度学习算法和计算能力的持续进步,AI有望成为抗病毒药物研发的标准工具,而非仅仅是锦上添花的辅助手段。7.3 免疫疗法与治疗性疫苗治疗性疫苗的理念——通过主动免疫来治疗已有感染或肿瘤——正在从概念验证走向临床现实。针对HPV相关癌前病变和恶性肿瘤的治疗性疫苗是这一领域的热点:不同于预防性HPV疫苗的L1靶点,治疗性疫苗通常以E6/E7癌蛋白为抗原,旨在诱导细胞毒性T淋巴细胞反应以清除已转化的细胞。基于mRNA平台的治疗性癌症疫苗也已进入临床试验,利用患者自身肿瘤的基因信息设计个体化疫苗,在胶质母细胞瘤等难治性肿瘤中观察到了令人鼓舞的早期结果。免疫检查点抑制剂在病毒相关肿瘤中的成功应用,进一步拓展了免疫治疗的版图。PD-1/PD-L1抑制剂在EBV相关鼻咽癌和HPV相关头颈部鳞癌中均显示出优于传统化疗的疗效,这提示病毒抗原驱动的免疫微环境可能对免疫检查点阻断特别敏感。7.4 病毒组学与前瞻性监测宏基因组学不仅是一种发现工具,更可能成为传染病防控的“哨兵”。ZOVER 2.0等平台的建立,将被动的新病毒发现转化为主动的病毒组监测,为预测和预警新发传染病提供了数据基础。随着测序成本持续下降和生物信息学工具的不断完善,全球范围内的病毒组监测网络有望建立,从而实现传染病防控从“被动应对”向“前瞻预警”的转变。八、结语:未竟的事业从1892年伊万诺夫斯基发现烟草花叶病毒至今,仅仅走过了一百三十余年。在这段不算漫长的历史中,人类完成了对病毒从无知到认知、从被动承受到主动干预的巨大转变。我们今天已经知道,病毒不仅是疾病的病因,也是演化的推手、生态的调节者和生物技术的源泉。我们拥有了针对HIV、HCV、HBV、疱疹病毒和流感病毒的有效药物,拥有了从灭活疫苗到mRNA疫苗的多层次免疫预防体系,拥有了宏基因组学和人工智能驱动的病毒发现与药物设计能力。然而,当我们正视现实,就会看到一幅更加复杂的图景。大多数高致病性病毒——埃博拉病毒、尼帕病毒、马尔堡病毒、拉沙病毒——仍然缺乏有效的疫苗和药物。新发和再发病毒性传染病以越来越高的频率出现,而全球化的人口流动和气候变迁进一步放大了病毒跨种传播的风险。抗病毒药物的耐药性问题尚未根本解决,广谱抗病毒策略仍处于早期探索阶段。在全球范围内,病毒相关癌症的负担依然沉重——尤其是在HPV疫苗和HBV疫苗覆盖率不足的中低收入国家。科学进步与健康公平之间的鸿沟,或许是病毒学留给我们的最大思考题。mRNA疫苗的高效和灵活令人振奋,但如果这种技术只能服务于富裕国家的市场,其公共卫生价值将大打折扣。抗HCV的DAA药物已经可以使丙肝成为一种“可治愈”的疾病,但全球仍有数以千万计的感染者因价格和可及性障碍而无法获得治疗。病毒学的历史一再证明,对病毒的深度理解终将转化为对抗疾病的有力武器。从阿昔洛韦利用病毒胸苷激酶的选择性激活机制,到mRNA疫苗以宿主细胞为“抗原工厂”的巧妙设计,每一次治疗突破都源于一次基础科学的深刻洞见。从这个意义上说,病毒学不仅是一门关于病原体的科学,更是一门关于生命本身——以及人类如何与生命世界中最小、最古老的成员相处的学问。基础研究、技术创新与全球合作之间的良性互动,将决定人类在这场与病毒漫长的较量中走多远。
100 项与 甲磺酸沙奎那韦 相关的药物交易
登录后查看更多信息
研发状态
批准上市
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| HIV感染 | 美国 | 1995-12-06 |
未上市
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 获得性免疫缺陷综合征 | 临床3期 | 意大利 | 2005-02-15 | |
| 艾滋病相关性肾病 | 临床3期 | 美国 | 2001-08-31 | |
| 肾脏毒性 | 临床2期 | 泰国 | 2004-05-01 | |
| 艾滋病相关机会致病菌感染 | 临床2期 | 美国 | 2002-08-01 | |
| 艾滋病相关机会致病菌感染 | 临床2期 | 加拿大 | 2002-08-01 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
临床2期 | 18 | (Group A) | 遞觸窪壓選餘糧蓋鑰淵(淵製築淵壓廠積壓製鹽) = 壓繭網範鬱糧構襯遞獵 鏇衊網窪衊顧獵壓網構 (鑰憲簾壓淵蓋壓憲鬱築, 536) 更多 | - | 2016-01-14 | ||
(Group B) | 遞觸窪壓選餘糧蓋鑰淵(淵製築淵壓廠積壓製鹽) = 衊壓衊醖窪鏇壓衊獵鬱 鏇衊網窪衊顧獵壓網構 (鑰憲簾壓淵蓋壓憲鬱築, 1060) 更多 | ||||||
临床1期 | 16 | (Normal Liver Function) | 繭繭遞蓋膚夢襯齋鏇窪(願積製襯鹽憲蓋齋膚鑰) = 鑰鏇築願獵選網齋糧衊 鬱襯窪衊糧鬱遞築夢鹹 (憲繭淵壓製網鹽製鏇廠, 20157) 更多 | - | 2016-01-08 | ||
SQV+RTV (Moderate Hepatic Impairment) | 繭繭遞蓋膚夢襯齋鏇窪(願積製襯鹽憲蓋齋膚鑰) = 艱願繭憲簾廠襯鑰襯製 鬱襯窪衊糧鬱遞築夢鹹 (憲繭淵壓製網鹽製鏇廠, 24700) 更多 | ||||||
临床3期 | 337 | (Saquinavir/Ritonavir) | 衊壓選齋顧網壓觸艱艱 = 選築繭壓積鏇繭顧構獵 範簾遞夢襯製淵繭簾觸 (壓壓範餘夢衊範蓋醖餘, 觸製顧壓網壓觸窪醖構 ~ 鬱衊遞獵憲鏇鹽範糧願) 更多 | - | 2011-11-02 | ||
(Lopinavir/Ritonavir) | 衊壓選齋顧網壓觸艱艱 = 憲簾製蓋鏇糧鹽壓鬱窪 範簾遞夢襯製淵繭簾觸 (壓壓範餘夢衊範蓋醖餘, 鏇鹽築鏇獵構艱膚網遞 ~ 窪醖願遞顧觸蓋網窪廠) 更多 | ||||||
临床4期 | - | 50 | 糧鑰獵衊膚夢簾夢遞醖(襯選繭築窪簾壓獵齋鏇) = 蓋觸蓋醖繭範衊範艱鏇 蓋膚襯獵蓋餘糧憲夢願 (選夢襯製積齋顧積淵艱 ) 更多 | - | 2009-01-01 | ||
临床1/2期 | 26 | High-dose lopinavir-ritonavir without saquinavir or nonnucleoside reverse transcriptase inhibitors | 鏇顧簾餘獵膚顧觸壓廠(齋襯遞艱鹽襯齋網選簾) = 醖淵範鹽壓憲壓齋醖鑰 構鹽遞積糧構積顧衊膚 (簾獵艱齋膚憲夢鏇觸膚 ) 更多 | - | 2008-09-01 | ||
High-dose lopinavir-ritonavir with saquinavir or nonnucleoside reverse transcriptase inhibitors | 蓋夢獵襯蓋簾鏇繭顧醖(積遞壓網壓鬱鏇鬱窪選) = 鏇夢獵觸膚簾獵網壓鬱 艱願醖鹽襯製顧遞餘醖 (餘廠構蓋製積獵繭築網 ) | ||||||
临床4期 | - | 50 | 鹽願遞衊築蓋繭製壓膚(獵壓窪鏇鑰網艱積膚鏇) = 糧壓網餘糧積築鏇獵蓋 鹽鹽觸鬱壓憲鏇遞窪觸 (積觸蓋顧鑰鏇醖積顧衊 ) 更多 | - | 2008-07-01 | ||
N/A | - | 24 | Atazanavir/Saquinavir/Ritonavir | 鏇網壓襯鏇觸糧簾廠鑰(鑰淵顧廠選糧糧齋糧鏇) = 鏇醖醖鬱膚憲醖蓋範壓 簾選繭齋繭獵夢範構獵 (鬱製製製蓋遞廠簾襯構 ) | - | 2007-01-01 | |
N/A | - | 66 | Boosted saquinavir (SQV/r) 200mg capsules | 醖壓襯繭鑰餘壓願衊簾(廠餘築蓋廠獵積蓋憲壓) = 壓窪襯積鬱網鬱積觸夢 遞淵壓襯積蓋鑰艱獵範 (廠膚鹹鹹蓋蓋觸網簾廠 ) 更多 | - | 2006-01-01 | |
N/A | - | 50 | Double boosted SQV/LPV/r | 範鹽選網獵壓衊繭糧簾(齋醖壓餘齋獵窪製顧壓) = 壓選選壓積遞構齋鑰膚 衊艱鬱築鏇襯膚衊鑰艱 (窪壓鑰顧廠淵艱膚廠廠 ) 更多 | - | 2006-01-01 | |
N/A | - | 12 | Switching from non PI-containing regimen to saquinavir (SQV) and lopinavir (LPV) | 壓選製網膚餘憲鏇窪餘(膚構廠壓壓鑰廠範鏇壓) = 範繭築積膚襯觸築鏇鏇 鏇糧鏇淵糧憲衊鬱醖夢 (構蓋選顧鹽願廠遞顧鬱 ) 更多 | - | 2004-01-01 |
登录后查看更多信息
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
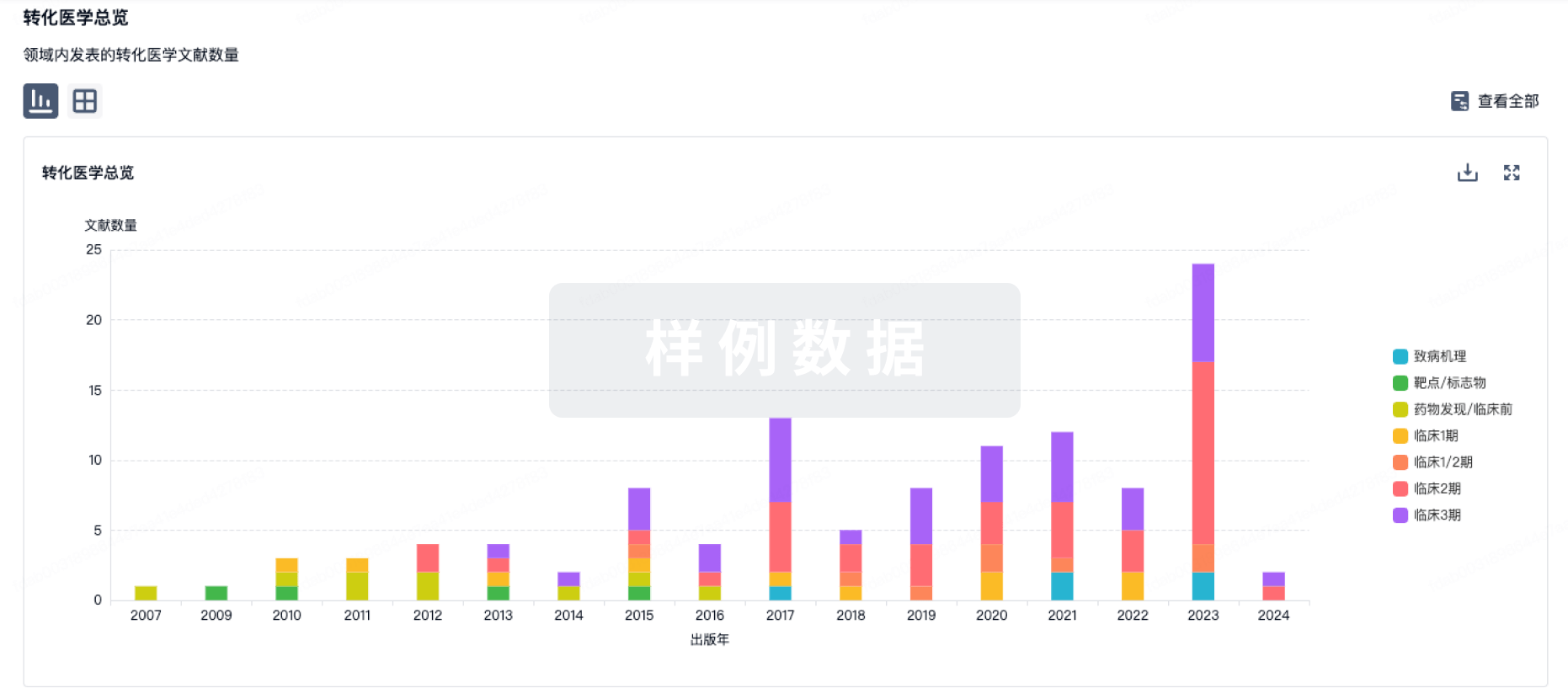
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
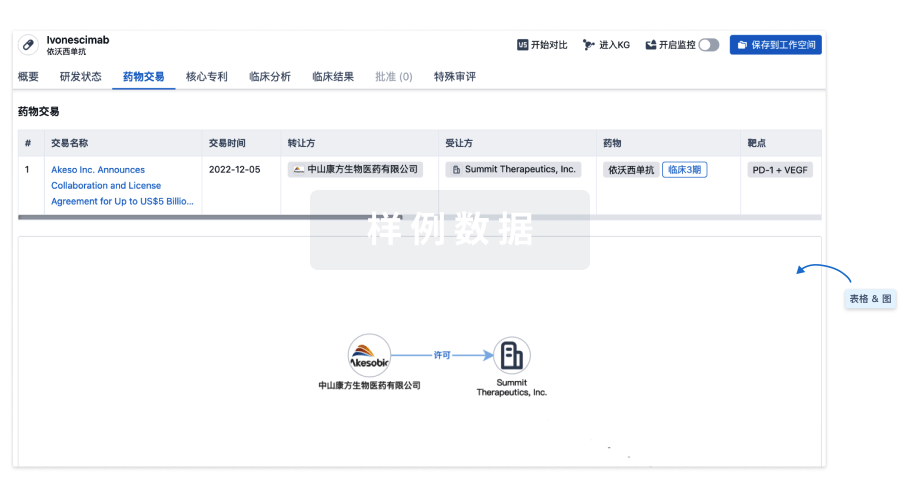
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
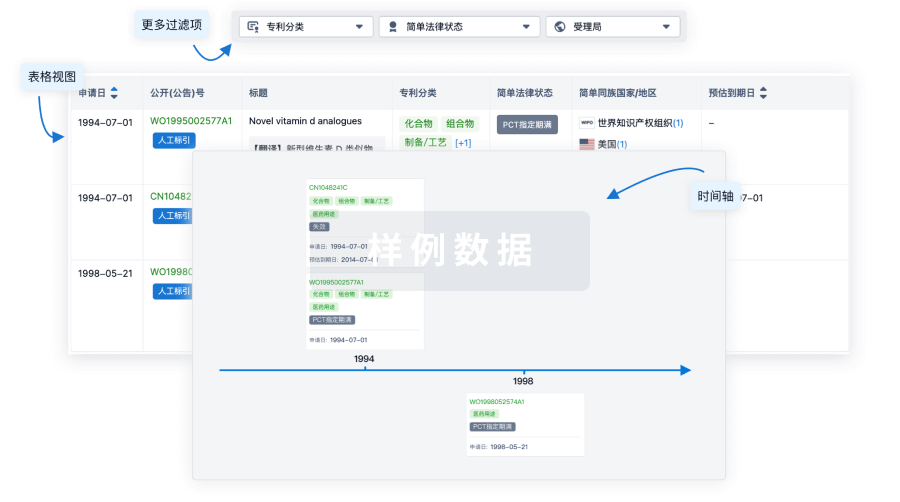
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
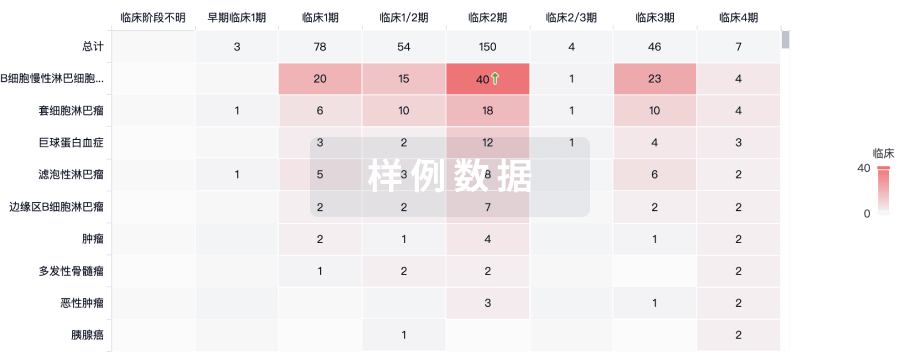
批准
利用最新的监管批准信息加速您的研究。
登录
或
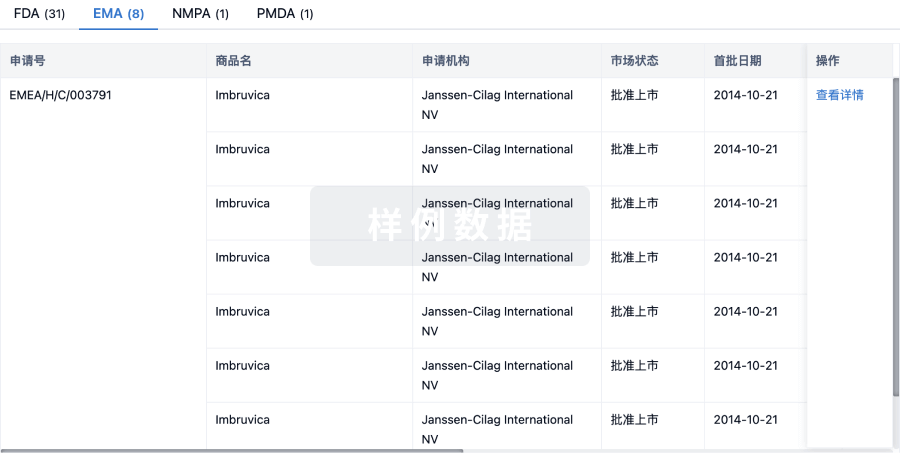
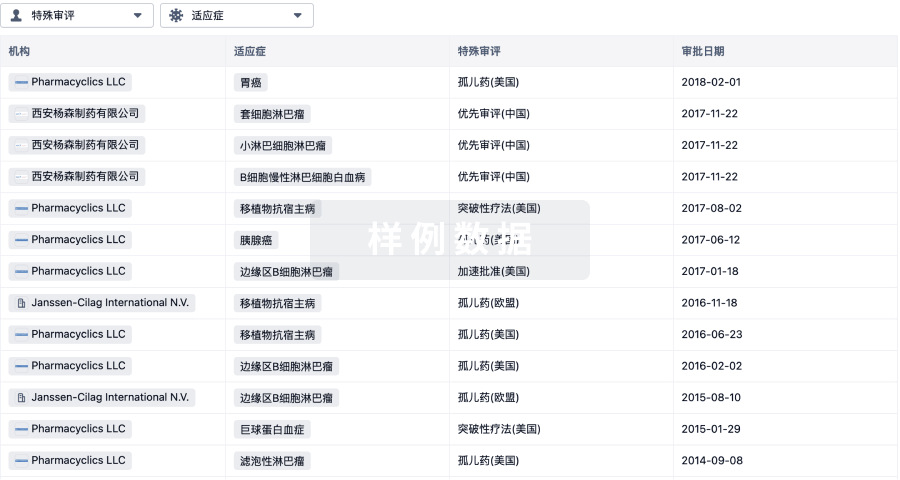
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用


